Chapter 15: Machine learning
Robert Johansson
Source code listings for Numerical Python - Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib (ISBN 979-8-8688-0412-0).
from sklearn import (
cluster,
datasets,
ensemble,
linear_model,
metrics,
model_selection,
neighbors,
svm,
tree,
)%matplotlib inline
import matplotlib.pyplot as pltimport numpy as npimport seaborn as snsimport matplotlib as mpl
mpl.rcParams["mathtext.fontset"] = "stix"
mpl.rcParams["font.family"] = "serif"
mpl.rcParams["font.sans-serif"] = "stix"
sns.set(style="whitegrid")sns.set(style="darkgrid")Built in datasets¶
datasets.load_wine # ()<function sklearn.datasets._base.load_wine(*, return_X_y=False, as_frame=False)>datasets.fetch_california_housing<function sklearn.datasets._california_housing.fetch_california_housing(*, data_home=None, download_if_missing=True, return_X_y=False, as_frame=False, n_retries=3, delay=1.0)>datasets.make_regression<function sklearn.datasets._samples_generator.make_regression(n_samples=100, n_features=100, *, n_informative=10, n_targets=1, bias=0.0, effective_rank=None, tail_strength=0.5, noise=0.0, shuffle=True, coef=False, random_state=None)>Regression¶
np.random.seed(123)X_all, y_all = datasets.make_regression(
n_samples=50, n_features=50, n_informative=10
) # , noise=2.5)X_train, X_test, y_train, y_test = model_selection.train_test_split(
X_all, y_all, train_size=0.5
)X_train.shape, y_train.shape((25, 50), (25,))X_test.shape, y_test.shape((25, 50), (25,))model = linear_model.LinearRegression()model.fit(X_train, y_train)Loading...
def sse(resid):
return sum(resid**2)resid_train = y_train - model.predict(X_train)
sse_train = sse(resid_train)
sse_trainnp.float64(1.5798927556531918e-24)resid_test = y_test - model.predict(X_test)
sse_test = sse(resid_train)
sse_testnp.float64(1.5798927556531918e-24)model.score(X_train, y_train)1.0model.score(X_test, y_test)0.314074006752017def plot_residuals_and_coeff(resid_train, resid_test, coeff):
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
axes[0].bar(np.arange(len(resid_train)), resid_train)
axes[0].set_xlabel("sample number")
axes[0].set_ylabel("residual")
axes[0].set_title("training data")
axes[1].bar(np.arange(len(resid_test)), resid_test)
axes[1].set_xlabel("sample number")
axes[1].set_ylabel("residual")
axes[1].set_title("testing data")
axes[2].bar(np.arange(len(coeff)), coeff)
axes[2].set_xlabel("coefficient number")
axes[2].set_ylabel("coefficient")
fig.tight_layout()
return fig, axesfig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-ols.pdf")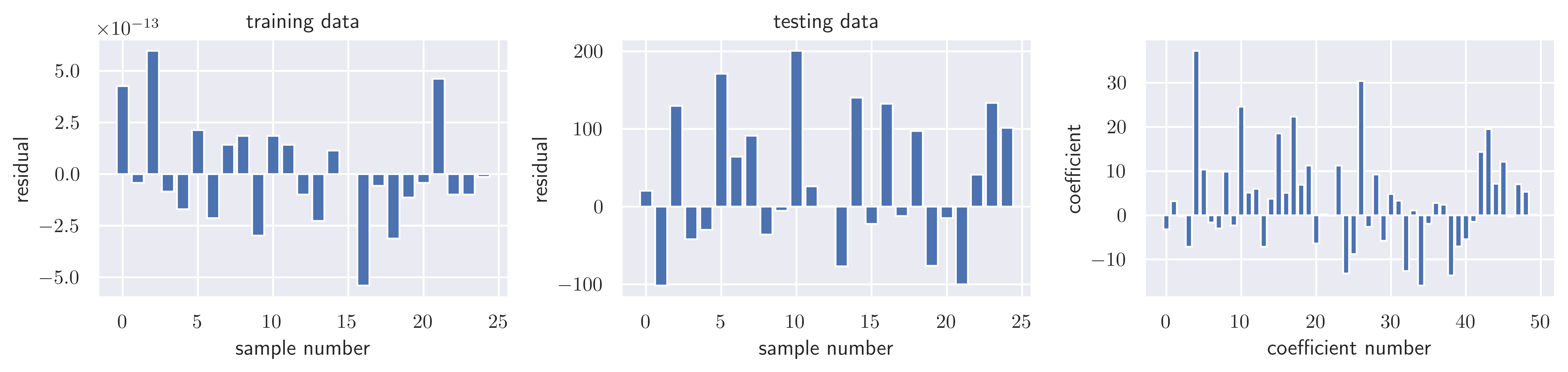
model = linear_model.Ridge() # alpha=2.5)model.fit(X_train, y_train)Loading...
resid_train = y_train - model.predict(X_train)
sse_train = sum(resid_train**2)
sse_trainnp.float64(178.50695164951017)resid_test = y_test - model.predict(X_test)
sse_test = sum(resid_test**2)
sse_testnp.float64(212737.0016010584)model.score(X_train, y_train), model.score(X_test, y_test)(0.9994595515017335, 0.31670332736075446)fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-ridge.pdf")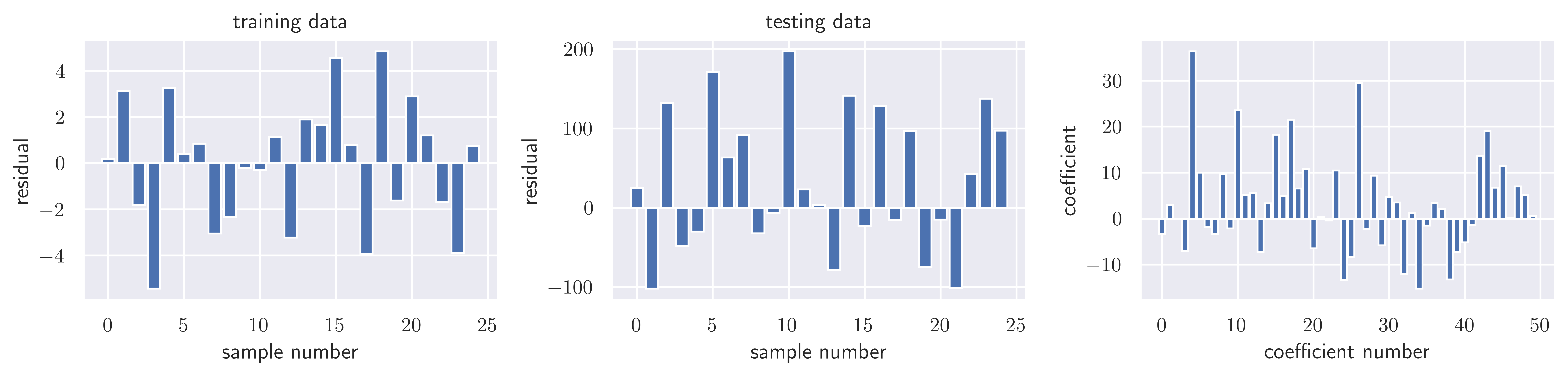
model = linear_model.Lasso(alpha=1.0)model.fit(X_train, y_train)Loading...
resid_train = y_train - model.predict(X_train)
sse_train = sse(resid_train)
sse_trainnp.float64(309.74971389532453)resid_test = y_test - model.predict(X_test)
sse_test = sse(resid_test)
sse_testnp.float64(1489.1176065002896)fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-lasso.pdf")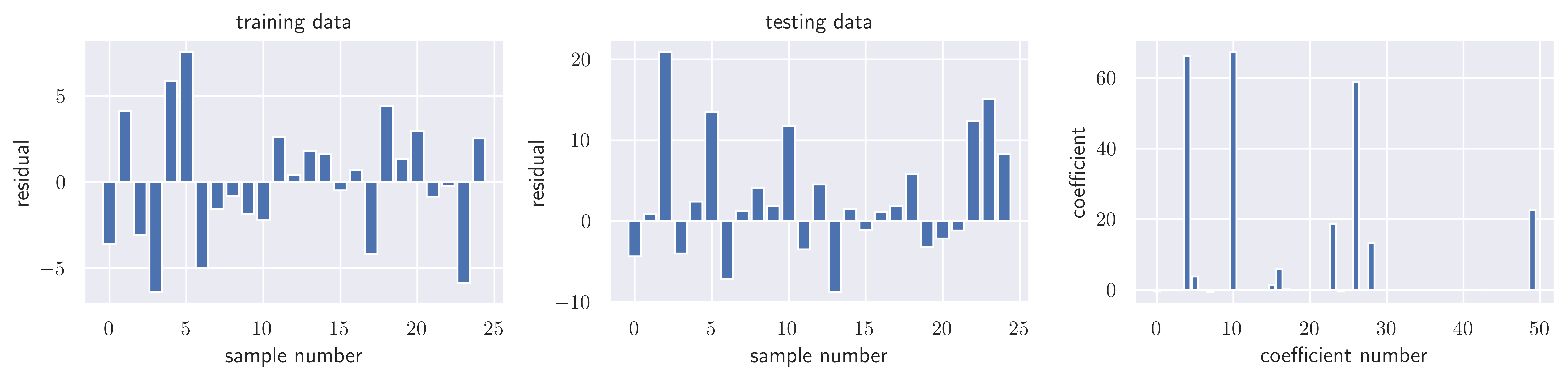
alphas = np.logspace(-4, 2, 100)coeffs = np.zeros((len(alphas), X_train.shape[1]))
sse_train = np.zeros_like(alphas)
sse_test = np.zeros_like(alphas)
for n, alpha in enumerate(alphas):
model = linear_model.Lasso(alpha=alpha)
model.fit(X_train, y_train)
coeffs[n, :] = model.coef_
resid = y_train - model.predict(X_train)
sse_train[n] = sum(resid**2)
resid = y_test - model.predict(X_test)
sse_test[n] = sum(resid**2)/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.311e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.682e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.220e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.515e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.462e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.865e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.408e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.514e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.364e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.928e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.447e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.509e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.718e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.941e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
/usr/lib/python3.14/site-packages/sklearn/linear_model/_coordinate_descent.py:716: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.351e+01, tolerance: 3.303e+01
model = cd_fast.enet_coordinate_descent(
fig, axes = plt.subplots(1, 2, figsize=(12, 4), sharex=True)
for n in range(coeffs.shape[1]):
axes[0].plot(np.log10(alphas), coeffs[:, n], color="k", lw=0.5)
axes[1].semilogy(np.log10(alphas), sse_train, label="train")
axes[1].semilogy(np.log10(alphas), sse_test, label="test")
axes[1].legend(loc=0)
axes[0].set_xlabel(r"${\log_{10}}\alpha$", fontsize=18)
axes[0].set_ylabel(r"coefficients", fontsize=18)
axes[1].set_xlabel(r"${\log_{10}}\alpha$", fontsize=18)
axes[1].set_ylabel(r"sse", fontsize=18)
fig.tight_layout()
fig.savefig("ch15-regression-lasso-vs-alpha.pdf")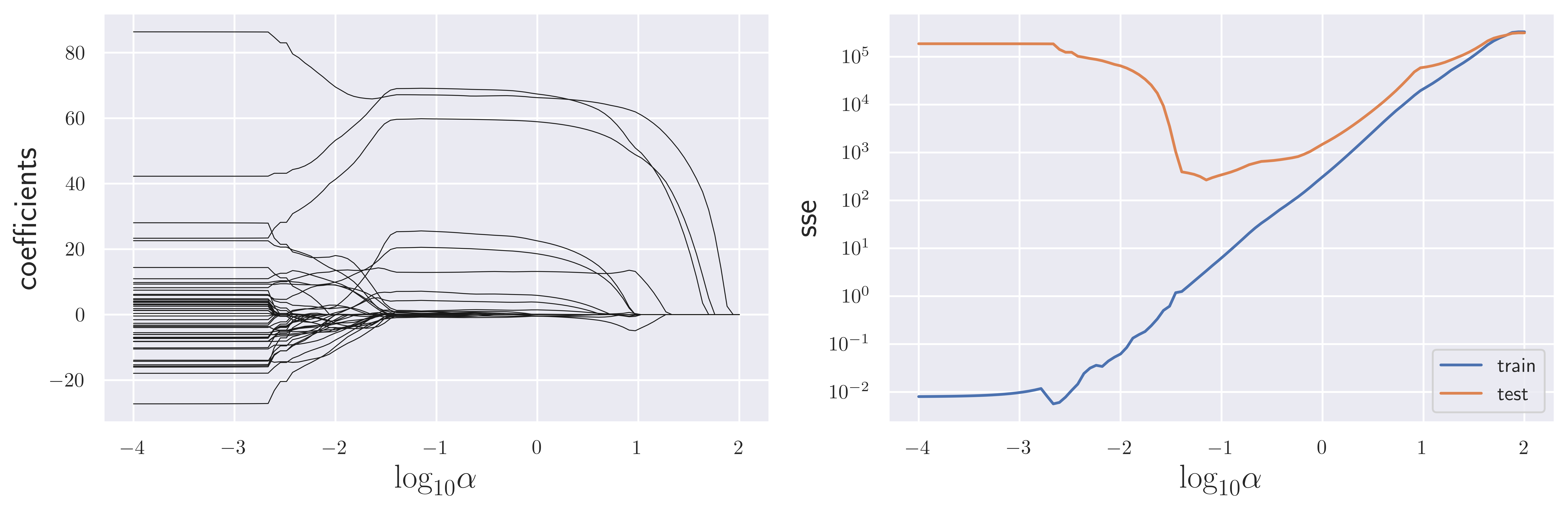
model = linear_model.LassoCV()model.fit(X_all, y_all)Loading...
model.alpha_np.float64(0.10689924823270765)resid_train = y_train - model.predict(X_train)
sse_train = sse(resid_train)
sse_trainnp.float64(4.064731090810533)resid_test = y_test - model.predict(X_test)
sse_test = sse(resid_test)
sse_testnp.float64(4.040111242413343)model.score(X_train, y_train), model.score(X_test, y_test)(0.9999876936007612, 0.9999870234395133)fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-lasso-cv.pdf")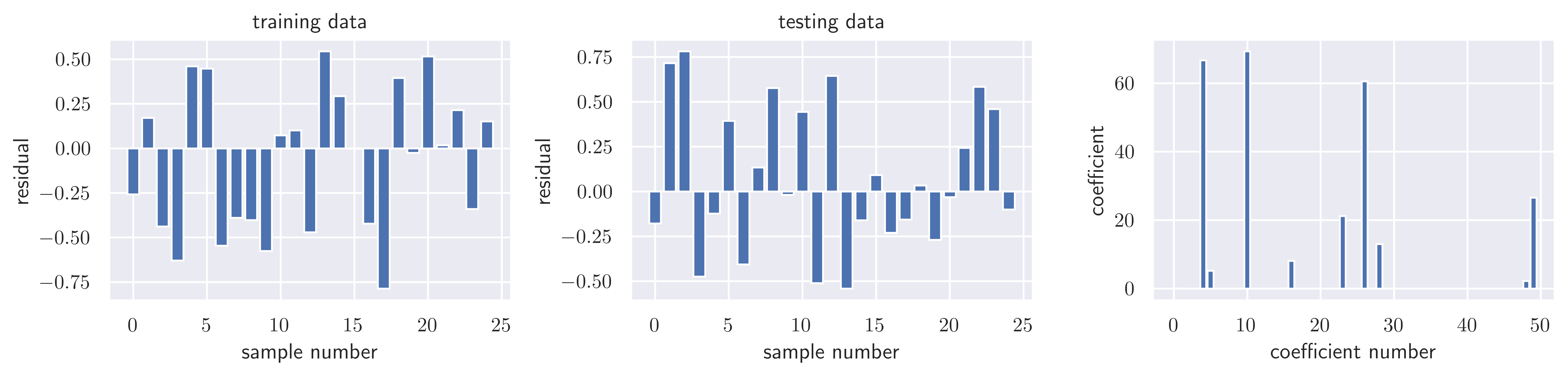
model = linear_model.ElasticNetCV()model.fit(X_all, y_all)Loading...
model.alpha_np.float64(0.13118477495069428)model.l1_ratio0.5resid_train = y_train - model.predict(X_train)
sse_train = sum(resid_train**2)
sse_trainnp.float64(2183.839172939127)resid_test = y_test - model.predict(X_test)
sse_test = sum(resid_test**2)
sse_testnp.float64(2650.050446338245)model.score(X_train, y_train), model.score(X_test, y_test)(0.9933881981034111, 0.9914882195448783)fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-elastic-net-cv.pdf")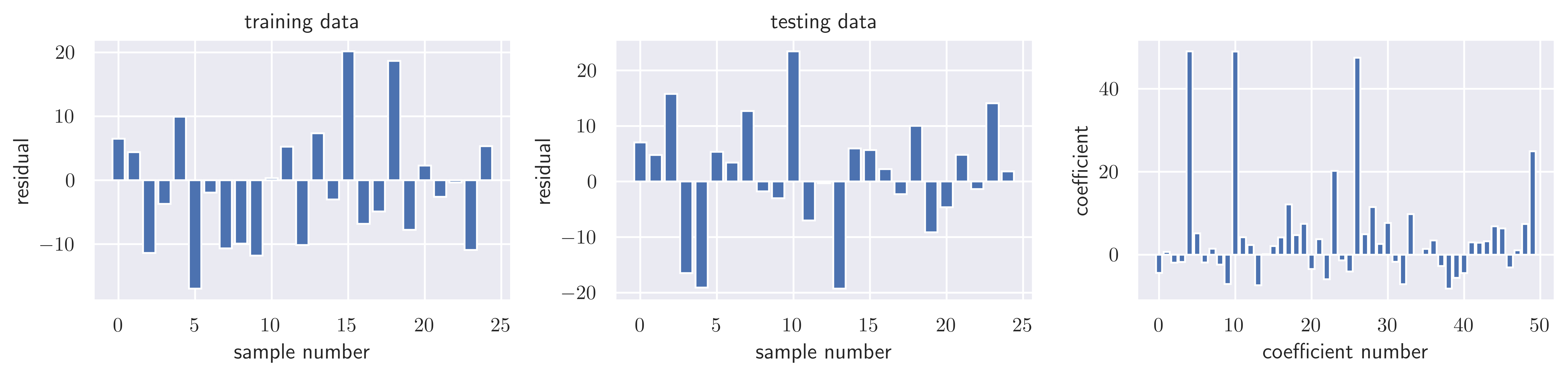
Classification¶
iris = datasets.load_iris()type(iris)sklearn.utils._bunch.Bunchiris.target_namesarray(['setosa', 'versicolor', 'virginica'], dtype='<U10')iris.feature_names['sepal length (cm)',
'sepal width (cm)',
'petal length (cm)',
'petal width (cm)']iris.data.shape(150, 4)iris.target.shape(150,)# print(iris['DESCR'])X_train, X_test, y_train, y_test = model_selection.train_test_split(
iris.data, iris.target, train_size=0.7, random_state=0
)classifier = linear_model.LogisticRegression()classifier.fit(X_train, y_train)Loading...
y_test_pred = classifier.predict(X_test)print(metrics.classification_report(y_test, y_test_pred)) precision recall f1-score support
0 1.00 1.00 1.00 16
1 1.00 0.94 0.97 18
2 0.92 1.00 0.96 11
accuracy 0.98 45
macro avg 0.97 0.98 0.98 45
weighted avg 0.98 0.98 0.98 45
np.bincount(y_test)array([16, 18, 11])metrics.confusion_matrix(y_test, y_test_pred)array([[16, 0, 0],
[ 0, 17, 1],
[ 0, 0, 11]])classifier = tree.DecisionTreeClassifier()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
metrics.confusion_matrix(y_test, y_test_pred)array([[16, 0, 0],
[ 0, 17, 1],
[ 0, 0, 11]])classifier = neighbors.KNeighborsClassifier()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
metrics.confusion_matrix(y_test, y_test_pred)array([[16, 0, 0],
[ 0, 17, 1],
[ 0, 0, 11]])classifier = svm.SVC()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
metrics.confusion_matrix(y_test, y_test_pred)array([[16, 0, 0],
[ 0, 17, 1],
[ 0, 0, 11]])classifier = ensemble.RandomForestClassifier()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
metrics.confusion_matrix(y_test, y_test_pred)array([[16, 0, 0],
[ 0, 17, 1],
[ 0, 0, 11]])train_size_vec = np.linspace(0.1, 0.9, 30)classifiers = [
tree.DecisionTreeClassifier,
neighbors.KNeighborsClassifier,
svm.SVC,
ensemble.RandomForestClassifier,
]cm_diags = np.zeros((3, len(train_size_vec), len(classifiers)), dtype=float)for n, train_size in enumerate(train_size_vec):
X_train, X_test, y_train, y_test = model_selection.train_test_split(
iris.data, iris.target, train_size=train_size
)
for m, Classifier in enumerate(classifiers):
classifier = Classifier()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
cm_diags[:, n, m] = metrics.confusion_matrix(y_test, y_test_pred).diagonal()
cm_diags[:, n, m] /= np.bincount(y_test)fig, axes = plt.subplots(1, len(classifiers), figsize=(12, 3))
for m, Classifier in enumerate(classifiers):
axes[m].plot(train_size_vec, cm_diags[2, :, m], label=iris.target_names[2])
axes[m].plot(train_size_vec, cm_diags[1, :, m], label=iris.target_names[1])
axes[m].plot(train_size_vec, cm_diags[0, :, m], label=iris.target_names[0])
axes[m].set_title(type(Classifier()).__name__)
axes[m].set_ylim(0, 1.1)
axes[m].set_xlim(0.1, 0.9)
axes[m].set_ylabel("classification accuracy")
axes[m].set_xlabel("training size ratio")
axes[m].legend(loc=4)
fig.tight_layout()
fig.savefig("ch15-classification-comparison.pdf")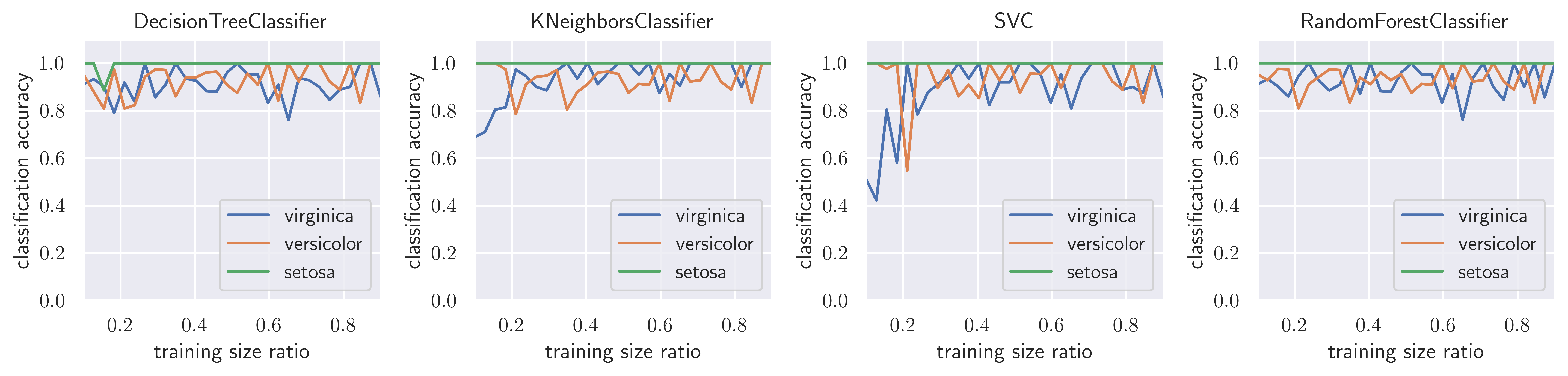
Clustering¶
X, y = iris.data, iris.targetnp.random.seed(123)n_clusters = 3c = cluster.KMeans(n_clusters=n_clusters)c.fit(X)Loading...
y_pred = c.predict(X)y_pred[::8]array([1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 0, 0, 0, 0, 0, 0],
dtype=int32)y[::8]array([0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2])idx_0, idx_1, idx_2 = (np.where(y_pred == n) for n in range(3))y_pred[idx_0], y_pred[idx_1], y_pred[idx_2] = 2, 0, 1y_pred[::8]array([0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2],
dtype=int32)metrics.confusion_matrix(y, y_pred)array([[50, 0, 0],
[ 0, 48, 2],
[ 0, 14, 36]])N = X.shape[1]
fig, axes = plt.subplots(N, N, figsize=(12, 12), sharex=True, sharey=True)
colors = ["coral", "blue", "green"]
markers = ["^", "v", "o"]
for m in range(N):
for n in range(N):
for p in range(n_clusters):
mask = y_pred == p
axes[m, n].scatter(
X[:, m][mask],
X[:, n][mask],
marker=markers[p],
s=30,
color=colors[p],
alpha=0.25,
)
for idx in np.where(y != y_pred):
axes[m, n].scatter(
X[idx, m],
X[idx, n],
marker="s",
s=30,
edgecolor="red",
facecolor=(1, 1, 1, 0),
)
axes[N - 1, m].set_xlabel(iris.feature_names[m], fontsize=16)
axes[m, 0].set_ylabel(iris.feature_names[m], fontsize=16)
fig.tight_layout()
fig.savefig("ch15-clustering.pdf")
fig.savefig("ch15-clustering.png", dpi=600)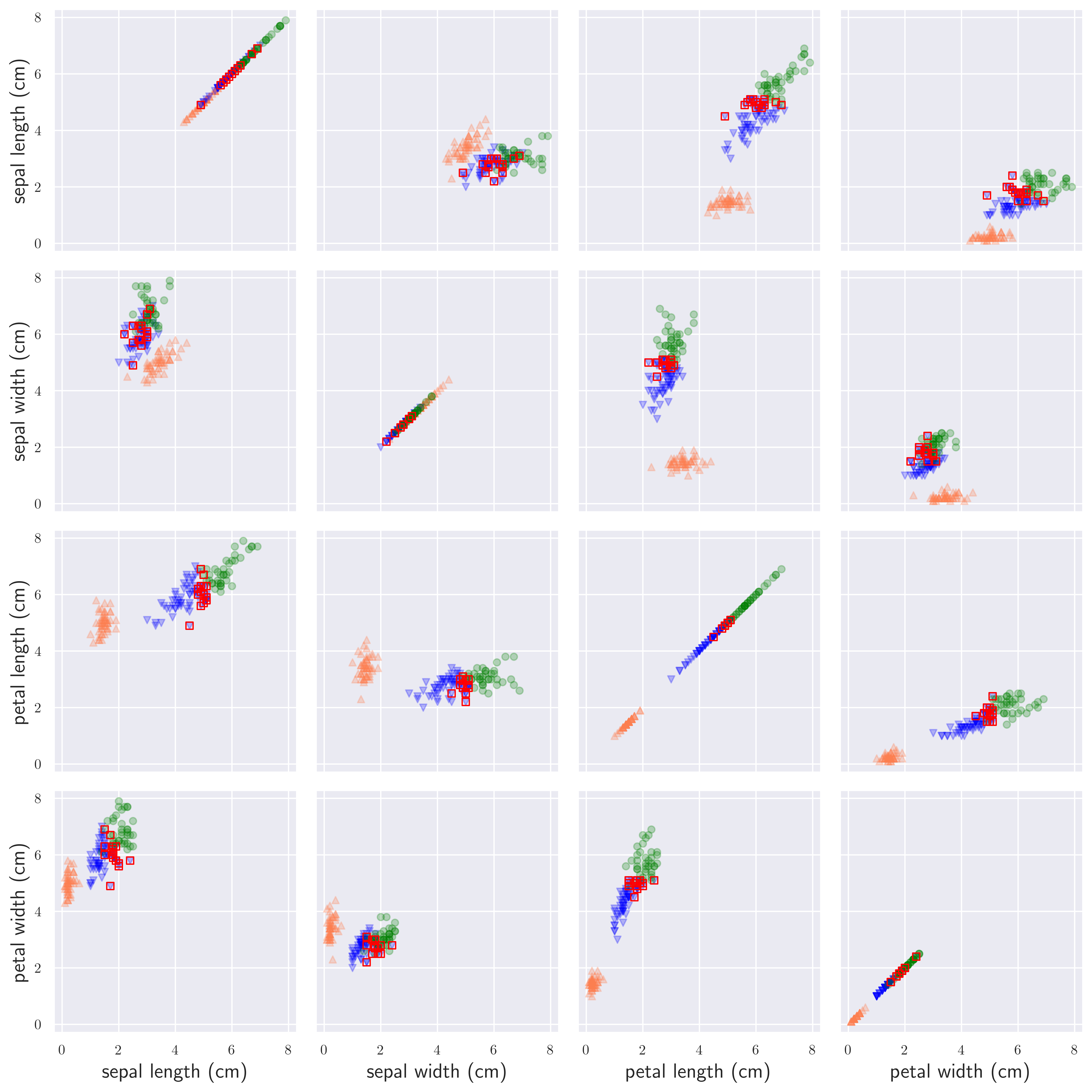
- Johansson, R. (2024). Numerical Python: Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib. Apress. 10.1007/979-8-8688-0413-7