Chapter 11: Partial differential equations
Robert Johansson
Source code listings for Numerical Python - Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib (ISBN 979-8-8688-0412-0).
import numpy as np%matplotlib inline
%config InlineBackend.figure_format='retina'
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams["mathtext.fontset"] = "stix"
mpl.rcParams["font.family"] = "stix"
mpl.rcParams["font.sans-serif"] = "stix"import mpl_toolkits.mplot3dimport scipy.sparse as spimport scipy.sparse.linalgimport scipy.linalg as laFinite-difference method¶
1d example¶
N = 5u0 = 1
u1 = 2dx = 1.0 / (N + 1)A = (np.eye(N, k=-1) - 2 * np.eye(N) + np.eye(N, k=1)) / dx**2Aarray([[-72., 36., 0., 0., 0.],
[ 36., -72., 36., 0., 0.],
[ 0., 36., -72., 36., 0.],
[ 0., 0., 36., -72., 36.],
[ 0., 0., 0., 36., -72.]])d = -5 * np.ones(N)
d[0] -= u0 / dx**2
d[N - 1] -= u1 / dx**2u = np.linalg.solve(A, d)x = np.linspace(0, 1, N + 2)U = np.hstack([[u0], u, [u1]])fig, ax = plt.subplots(figsize=(8, 4))
ax.plot(x, U)
ax.plot(x[1:-1], u, "ks")
ax.set_xlim(0, 1)
ax.set_xlabel(r"$x$", fontsize=18)
ax.set_ylabel(r"$u(x)$", fontsize=18)
fig.savefig("ch11-fdm-1d.pdf")
fig.tight_layout();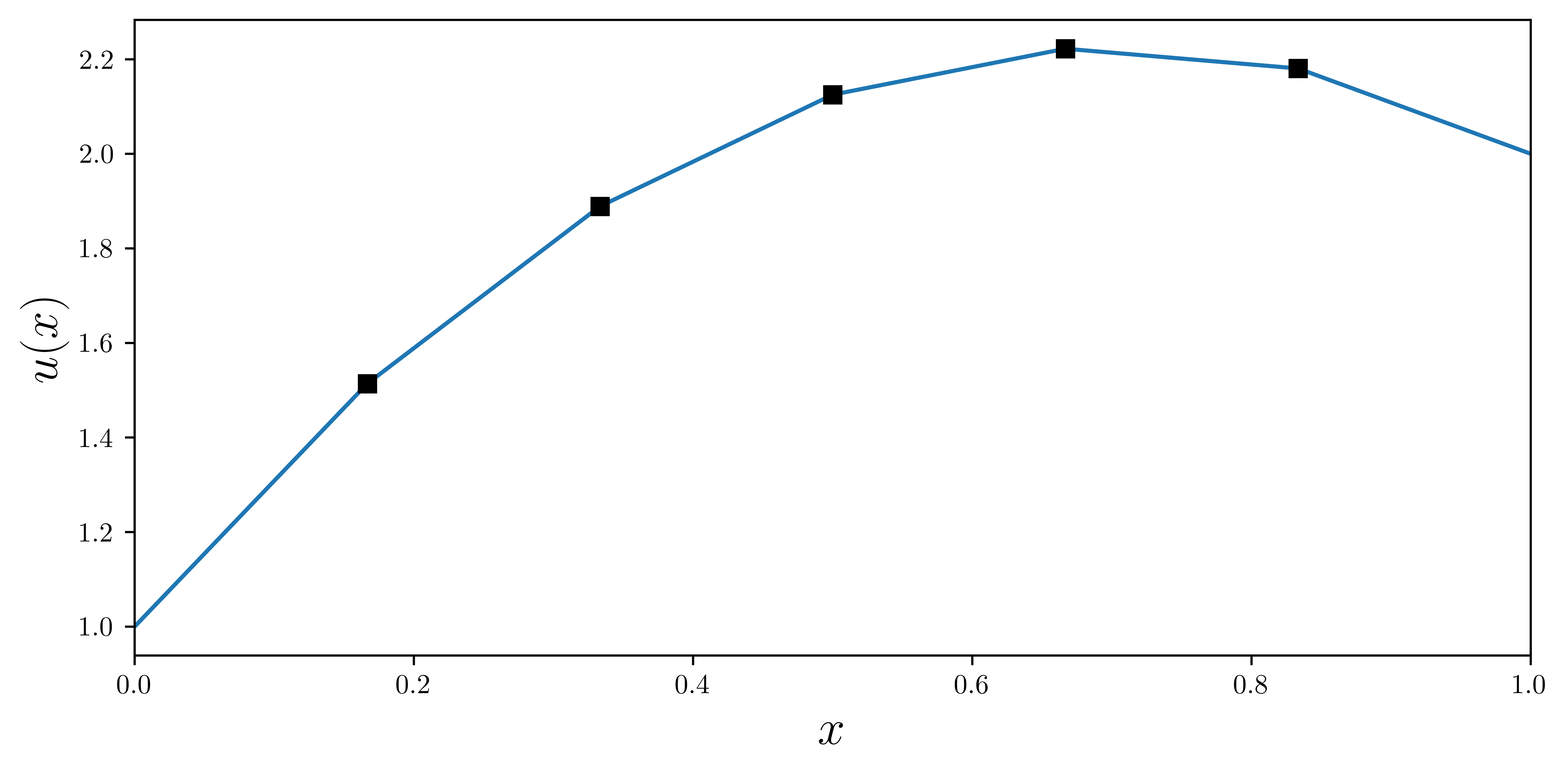
2d example¶
N = 100u0_t, u0_b = 5, -5u0_l, u0_r = 3, -1dx = 1.0 / (N + 1)A_1d = (sp.eye(N, k=-1) + sp.eye(N, k=1) - 4 * sp.eye(N)) / dx**2A = sp.kron(sp.eye(N), A_1d) + (sp.eye(N**2, k=-N) + sp.eye(N**2, k=N)) / dx**2A<Compressed Sparse Row sparse matrix of dtype 'float64'
with 49600 stored elements and shape (10000, 10000)>A.nnz * 1.0 / np.prod(A.shape) * 2000np.float64(0.992)d = np.zeros((N, N))
d[0, :] += -u0_b
d[-1, :] += -u0_t
d[:, 0] += -u0_l
d[:, -1] += -u0_r
d = d.reshape(N**2) / dx**2u = sp.linalg.spsolve(A, d).reshape(N, N)U = np.vstack(
[
np.ones((1, N + 2)) * u0_b,
np.hstack([np.ones((N, 1)) * u0_l, u, np.ones((N, 1)) * u0_r]),
np.ones((1, N + 2)) * u0_t,
]
)fig, ax = plt.subplots(1, 1, figsize=(8, 6))
x = np.linspace(0, 1, N + 2)
X, Y = np.meshgrid(x, x)
c = ax.pcolor(X, Y, U, vmin=-5, vmax=5, cmap=mpl.cm.get_cmap("RdBu_r"))
cb = plt.colorbar(c, ax=ax)
ax.set_xlabel(r"$x_1$", fontsize=18)
ax.set_ylabel(r"$x_2$", fontsize=18)
cb.set_label(r"$u(x_1, x_2)$", fontsize=18)
fig.savefig("ch11-fdm-2d.pdf")
fig.tight_layout()/tmp/ipykernel_46899/2799813999.py:6: MatplotlibDeprecationWarning: The get_cmap function was deprecated in Matplotlib 3.7 and will be removed in 3.11. Use ``matplotlib.colormaps[name]`` or ``matplotlib.colormaps.get_cmap()`` or ``pyplot.get_cmap()`` instead.
c = ax.pcolor(X, Y, U, vmin=-5, vmax=5, cmap=mpl.cm.get_cmap("RdBu_r"))
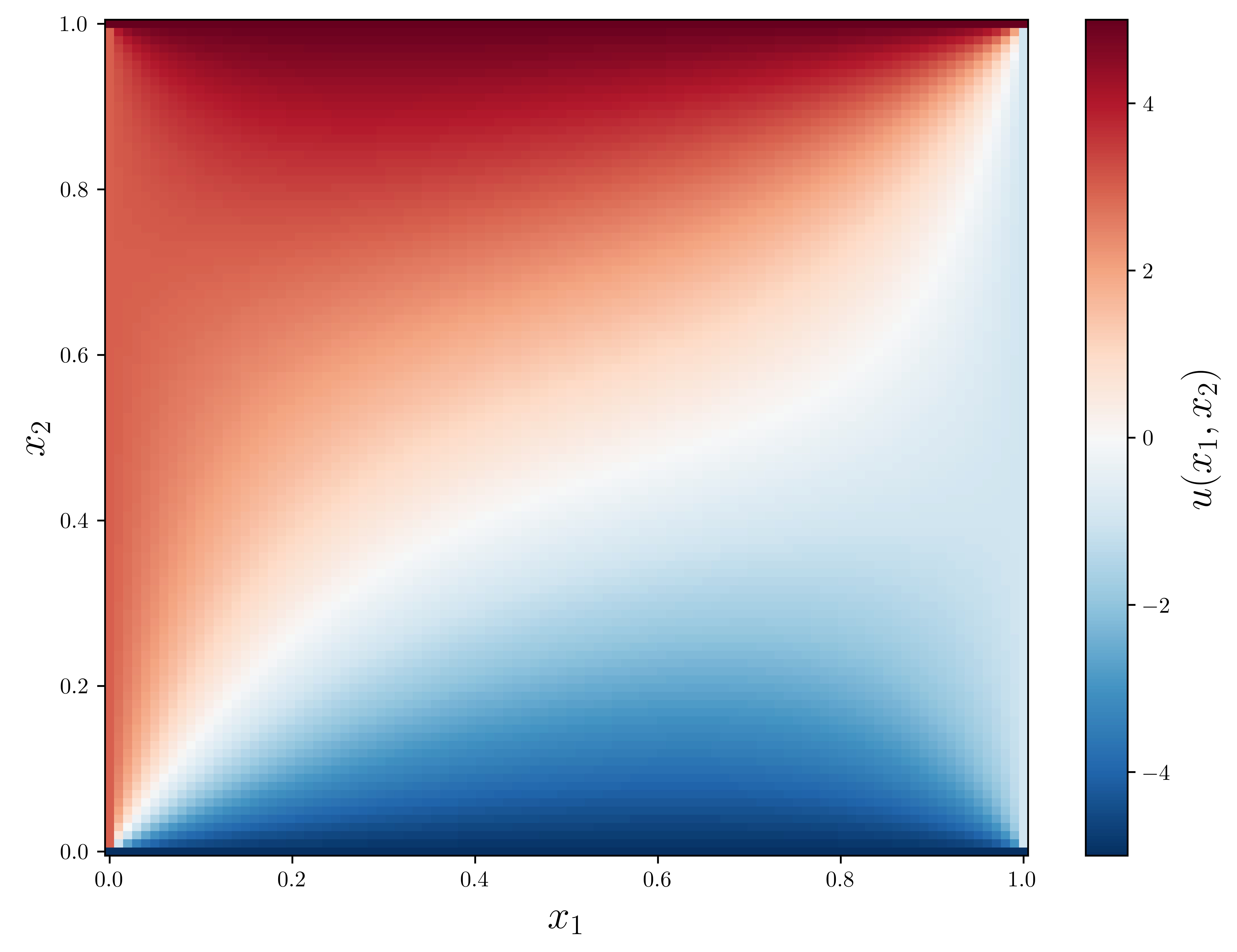
x = np.linspace(0, 1, N + 2)
X, Y = np.meshgrid(x, x)fig = plt.figure(figsize=(12, 5.5))
cmap = mpl.cm.get_cmap("RdBu_r")
ax = fig.add_subplot(1, 2, 1)
p = ax.pcolor(X, Y, U, vmin=-5, vmax=5, cmap=cmap)
ax.set_xlabel(r"$x_1$", fontsize=18)
ax.set_ylabel(r"$x_2$", fontsize=18)
ax = fig.add_subplot(1, 2, 2, projection="3d")
p = ax.plot_surface(
X, Y, U, vmin=-5, vmax=5, rstride=3, cstride=3, linewidth=0, cmap=cmap
)
ax.set_xlabel(r"$x_1$", fontsize=16)
ax.set_ylabel(r"$x_2$", fontsize=16)
cb = plt.colorbar(p, ax=ax, shrink=0.75)
cb.set_label(r"$u(x_1, x_2)$", fontsize=18)
fig.savefig("ch11-fdm-2d.pdf")
fig.savefig("ch11-fdm-2d.png")
fig.tight_layout()/tmp/ipykernel_46899/386270017.py:2: MatplotlibDeprecationWarning: The get_cmap function was deprecated in Matplotlib 3.7 and will be removed in 3.11. Use ``matplotlib.colormaps[name]`` or ``matplotlib.colormaps.get_cmap()`` or ``pyplot.get_cmap()`` instead.
cmap = mpl.cm.get_cmap("RdBu_r")
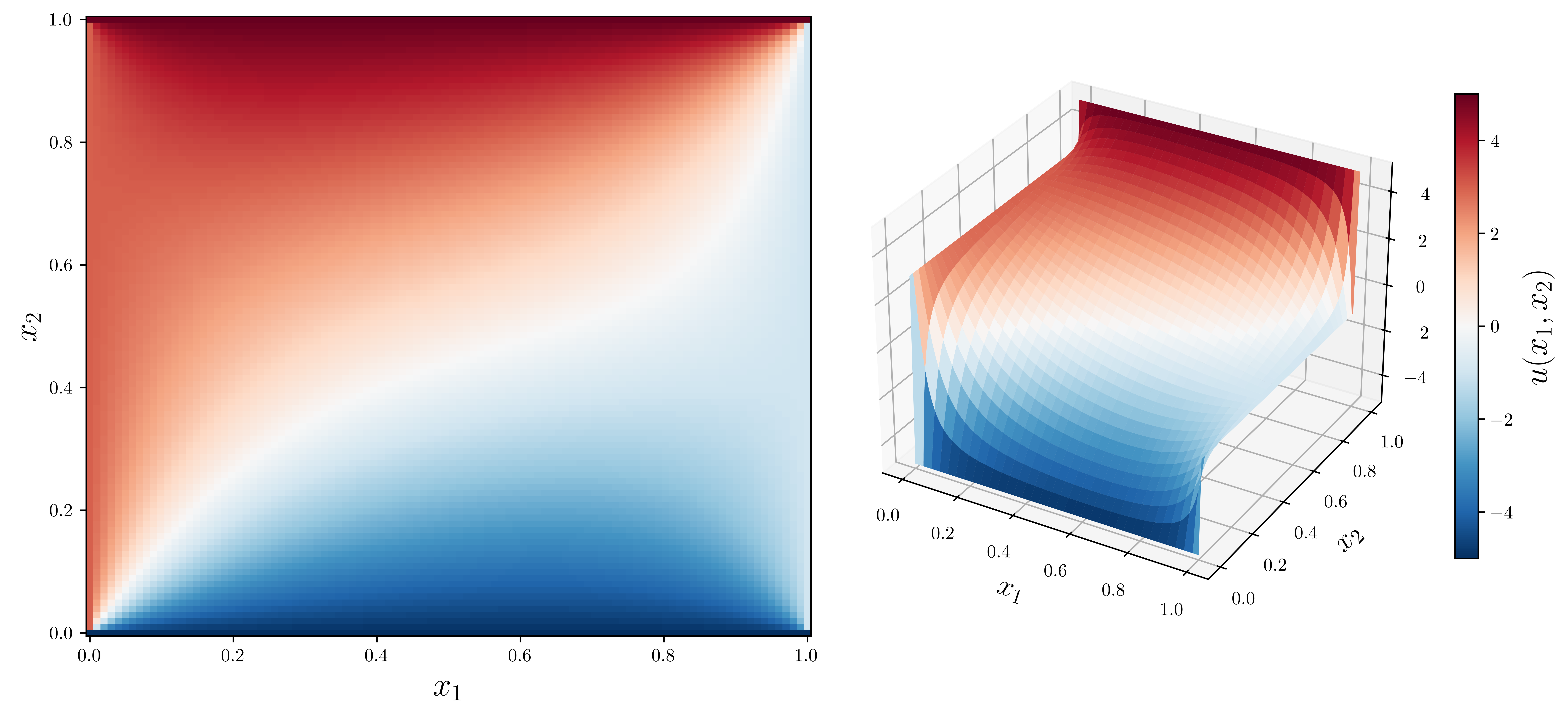
Compare performance when using dense/sparse matrices¶
A_dense = A.todense()%timeit np.linalg.solve(A_dense, d)15.1 s ± 520 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
%timeit la.solve(A_dense, d)11.9 s ± 2.66 s per loop (mean ± std. dev. of 7 runs, 1 loop each)
%timeit sp.linalg.spsolve(A, d)23.5 ms ± 1.84 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
10.8 / 31.9e-3338.55799373040762d example with source term¶
d = -np.ones((N, N))
d = d.reshape(N**2)u = sp.linalg.spsolve(A, d).reshape(N, N)U = np.vstack(
[
np.zeros((1, N + 2)),
np.hstack([np.zeros((N, 1)), u, np.zeros((N, 1))]),
np.zeros((1, N + 2)),
]
)x = np.linspace(0, 1, N + 2)
X, Y = np.meshgrid(x, x)fig, ax = plt.subplots(1, 1, figsize=(8, 6), subplot_kw={"projection": "3d"})
p = ax.plot_surface(
X, Y, U, rstride=4, cstride=4, linewidth=0, cmap=mpl.cm.get_cmap("Reds")
)
cb = fig.colorbar(p, shrink=0.5)
ax.set_xlabel(r"$x_1$", fontsize=18)
ax.set_ylabel(r"$x_2$", fontsize=18)
cb.set_label(r"$u(x_1, x_2)$", fontsize=18)/tmp/ipykernel_46899/3899247969.py:4: MatplotlibDeprecationWarning: The get_cmap function was deprecated in Matplotlib 3.7 and will be removed in 3.11. Use ``matplotlib.colormaps[name]`` or ``matplotlib.colormaps.get_cmap()`` or ``pyplot.get_cmap()`` instead.
X, Y, U, rstride=4, cstride=4, linewidth=0, cmap=mpl.cm.get_cmap("Reds")
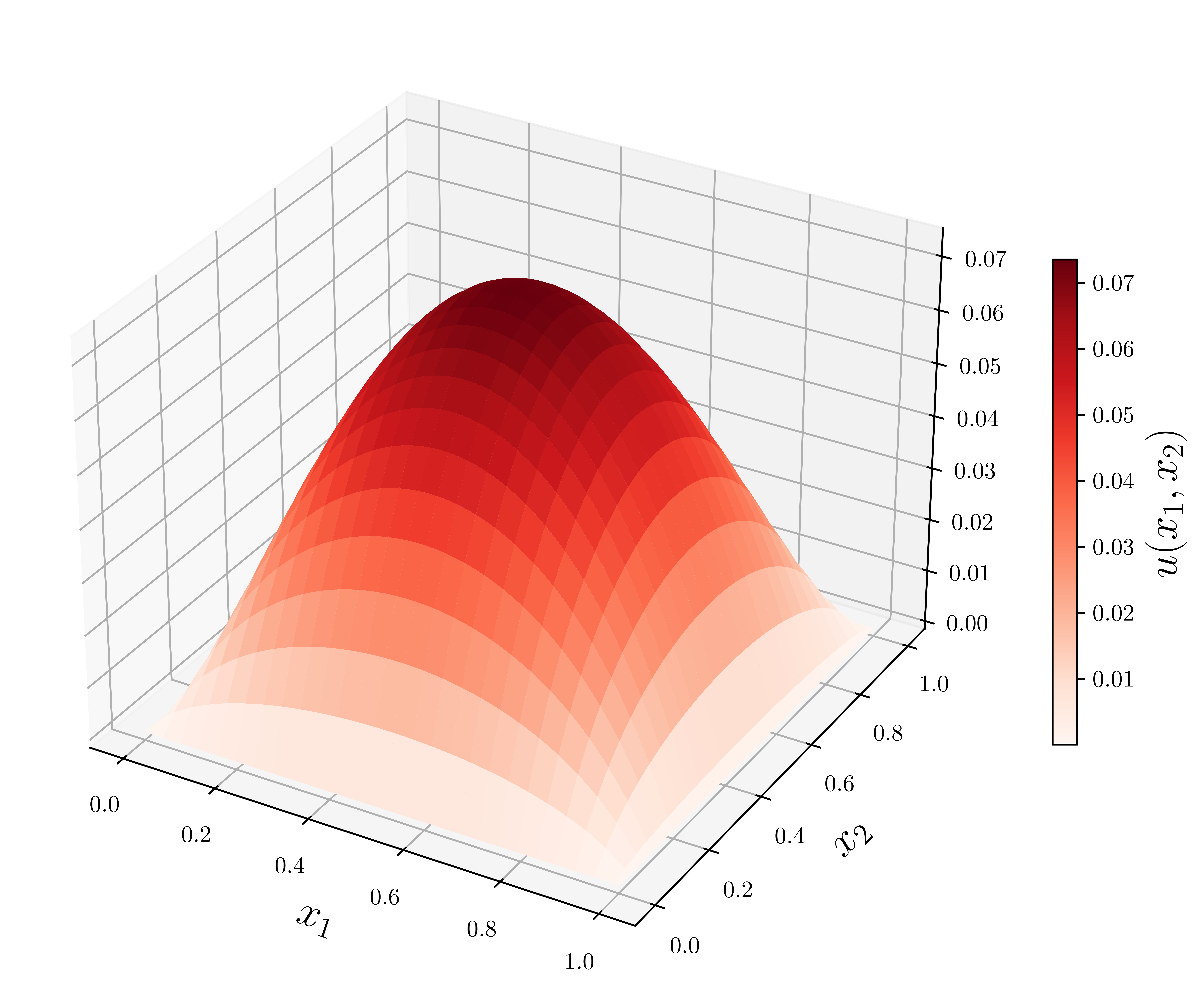
- Johansson, R. (2024). Numerical Python: Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib. Apress. 10.1007/979-8-8688-0413-7