Chapter 13: Statistics
Robert Johansson
Source code listings for Numerical Python - Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib (ISBN 979-8-8688-0412-0).
Imports¶
from scipy import statsfrom scipy import optimizeimport random
import numpy as np%matplotlib inline
import matplotlib.pyplot as pltimport matplotlib as mplmpl.rcParams["mathtext.fontset"] = "stix"
mpl.rcParams["font.family"] = "serif"
mpl.rcParams["font.sans-serif"] = "stix"import seaborn as snssns.set(style="whitegrid")Descriptive statistics¶
x = np.array([3.5, 1.1, 3.2, 2.8, 6.7, 4.4, 0.9, 2.2])np.mean(x)np.float64(3.1)np.median(x)np.float64(3.0)x.min(), x.max()(np.float64(0.9), np.float64(6.7))x.var()np.float64(3.0700000000000007)x.std()np.float64(1.7521415467935233)x.var(ddof=1)np.float64(3.5085714285714293)x.std(ddof=1)np.float64(1.8731181032095732)Random numbers¶
random.seed(123456789)random.random()0.6414006161858726random.randint(0, 10) # 0 and 10 inclusive8np.random.seed(123456789)np.random.rand()0.532833024789759np.random.randn()0.8768342101492541np.random.rand(5)array([0.71356403, 0.25699895, 0.75269361, 0.88387918, 0.15489908])np.random.randn(2, 4)array([[ 3.13325952, 1.15727052, 1.37591514, 0.94302846],
[ 0.8478706 , 0.52969142, -0.56940469, 0.83180456]])np.random.randint(10, size=10)array([0, 3, 8, 3, 9, 0, 6, 9, 2, 7])np.random.randint(low=10, high=20, size=(2, 10))array([[12, 18, 18, 17, 14, 12, 14, 10, 16, 19],
[15, 13, 15, 18, 11, 17, 17, 10, 13, 17]])fig, axes = plt.subplots(1, 3, figsize=(12, 3))
axes[0].hist(np.random.rand(10000))
axes[0].set_title("rand")
axes[1].hist(np.random.randn(10000))
axes[1].set_title("randn")
axes[2].hist(np.random.randint(low=1, high=10, size=10000), bins=9, align="left")
axes[2].set_title("randint(low=1, high=10)")
fig.tight_layout()
fig.savefig("ch13-random-hist.pdf")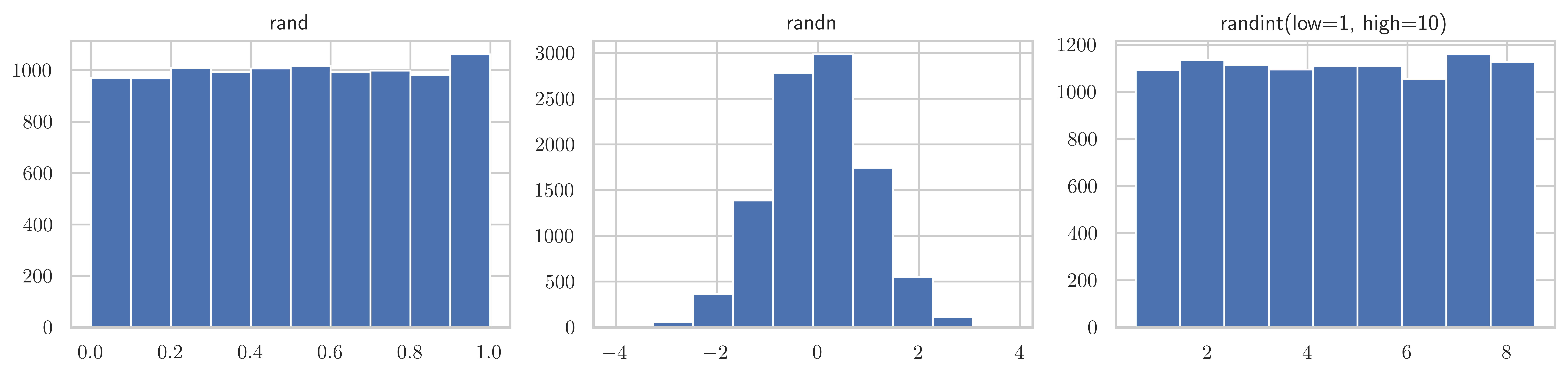
# random.sample(range(10), 5)np.random.choice(10, 5, replace=False)array([9, 0, 5, 8, 1])np.random.seed(123456789)np.random.rand()0.532833024789759np.random.seed(123456789)
np.random.rand()0.532833024789759np.random.seed(123456789)
np.random.rand()0.532833024789759prng = np.random.RandomState(123456789)prng.randn(2, 4)array([[ 2.212902 , 2.1283978 , 1.8417114 , 0.08238248],
[ 0.85896368, -0.82601643, 1.15727052, 1.37591514]])prng.chisquare(1, size=(2, 2))array([[1.26859720e+00, 2.02731988e+00],
[2.52605129e-05, 3.00376585e-04]])prng.standard_t(1, size=(2, 3))array([[ 0.59734384, -1.27669959, 0.09724793],
[ 0.22451466, 0.39697518, -0.19469463]])prng.f(5, 2, size=(2, 4))array([[ 0.77372119, 0.1213796 , 1.64779052, 1.21399831],
[ 0.45471421, 17.64891848, 1.48620557, 2.55433261]])prng.binomial(10, 0.5, size=10)array([8, 3, 4, 2, 4, 5, 4, 4, 7, 5])prng.poisson(5, size=10)array([7, 1, 3, 4, 6, 4, 9, 7, 3, 6])Probability distributions and random variables¶
np.random.seed(123456789)X = stats.norm(1, 0.5)X.mean()np.float64(1.0)X.median()np.float64(1.0)X.std()np.float64(0.5)X.var()np.float64(0.25)[X.moment(n) for n in range(5)][np.float64(1.0),
np.float64(1.0),
np.float64(1.25),
np.float64(1.75),
np.float64(2.6875)]X.stats()(np.float64(1.0), np.float64(0.25))X.pdf([0, 1, 2])array([0.10798193, 0.79788456, 0.10798193])X.cdf([0, 1, 2])array([0.02275013, 0.5 , 0.97724987])X.rvs(10)array([2.106451 , 2.0641989 , 1.9208557 , 1.04119124, 1.42948184,
0.58699179, 1.57863526, 1.68795757, 1.47151423, 1.4239353 ])stats.norm(1, 0.5).stats()(np.float64(1.0), np.float64(0.25))stats.norm.stats(loc=2, scale=0.5)(np.float64(2.0), np.float64(0.25))X.interval(0.95)(np.float64(0.020018007729972975), np.float64(1.979981992270027))X.interval(0.99)(np.float64(-0.2879146517744502), np.float64(2.28791465177445))def plot_rv_distribution(X, axes=None):
"""Plot the PDF, CDF, SF and PPF of a given random variable"""
if axes is None:
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
x_min_999, x_max_999 = X.interval(0.999)
x999 = np.linspace(x_min_999, x_max_999, 1000)
x_min_95, x_max_95 = X.interval(0.95)
x95 = np.linspace(x_min_95, x_max_95, 1000)
if hasattr(X.dist, "pdf"):
axes[0].plot(x999, X.pdf(x999), label="PDF")
axes[0].fill_between(x95, X.pdf(x95), alpha=0.25)
else:
x999_int = np.unique(x999.astype(int))
axes[0].bar(x999_int, X.pmf(x999_int), label="PMF")
axes[1].plot(x999, X.cdf(x999), label="CDF")
axes[1].plot(x999, X.sf(x999), label="SF")
axes[2].plot(x999, X.ppf(x999), label="PPF")
for ax in axes:
ax.legend()
return axesfig, axes = plt.subplots(3, 3, figsize=(12, 9))
X = stats.norm()
plot_rv_distribution(X, axes=axes[0, :])
axes[0, 0].set_ylabel("Normal dist.")
X = stats.f(2, 50)
plot_rv_distribution(X, axes=axes[1, :])
axes[1, 0].set_ylabel("F dist.")
X = stats.poisson(5)
plot_rv_distribution(X, axes=axes[2, :])
axes[2, 0].set_ylabel("Poisson dist.")
fig.tight_layout()
fig.savefig("ch13-distributions.pdf")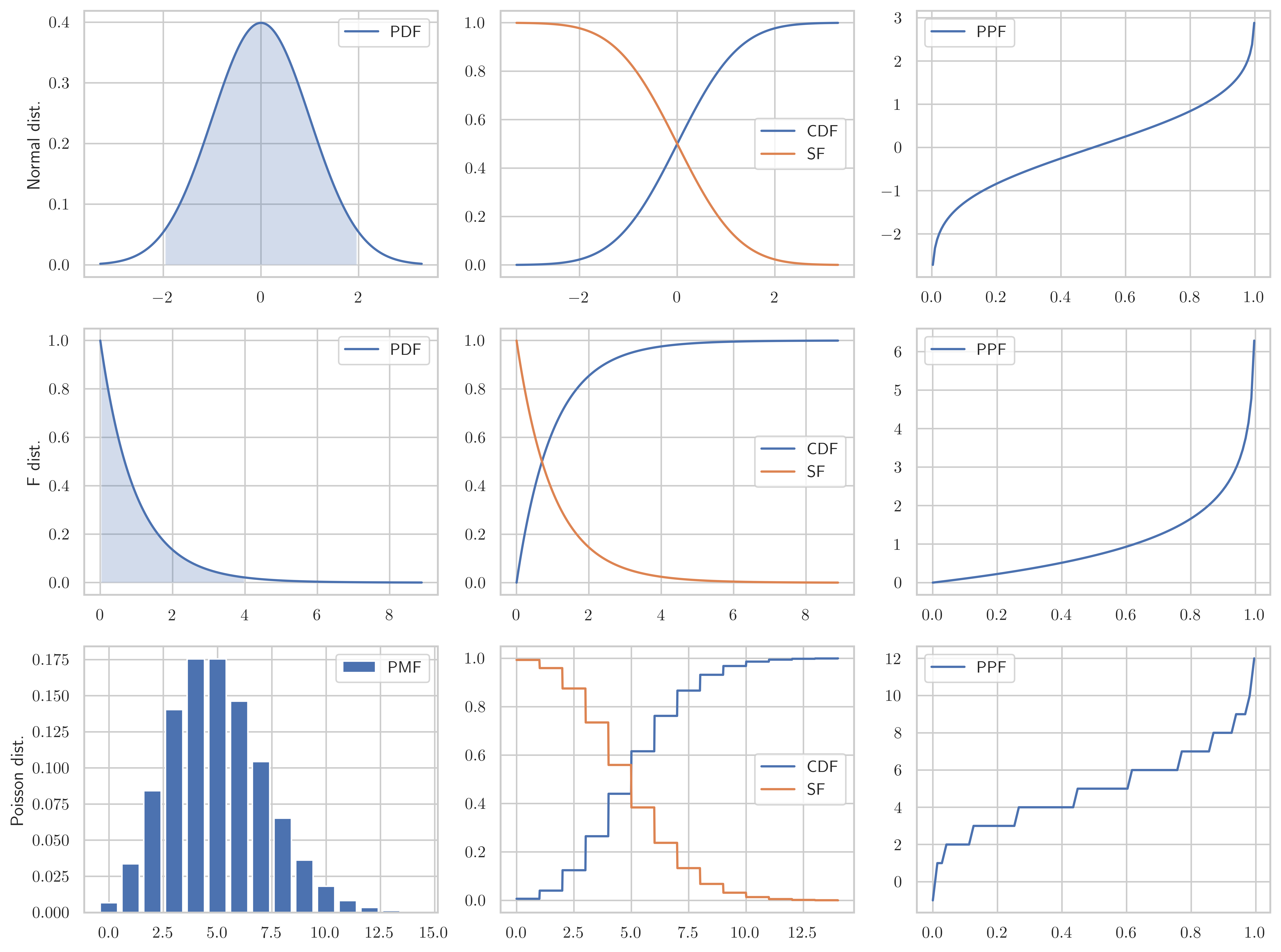
def plot_dist_samples(X, X_samples, title=None, ax=None):
"""Plot the PDF and histogram of samples of a continuous random variable"""
if ax is None:
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
x_lim = X.interval(0.99)
x = np.linspace(*x_lim, num=100)
ax.plot(x, X.pdf(x), label="PDF", lw=3)
ax.hist(X_samples, label="samples", density=True, bins=75)
ax.set_xlim(*x_lim)
ax.legend()
if title:
ax.set_title(title)
return axfig, axes = plt.subplots(1, 3, figsize=(12, 3))
X = stats.t(7.0)
plot_dist_samples(X, X.rvs(2000), "Student's t dist.", ax=axes[0])
X = stats.chi2(5.0)
plot_dist_samples(X, X.rvs(2000), r"$\chi^2$ dist.", ax=axes[1])
X = stats.expon(0.5)
plot_dist_samples(X, X.rvs(2000), "exponential dist.", ax=axes[2])
fig.tight_layout()
fig.savefig("ch13-dist-sample.pdf")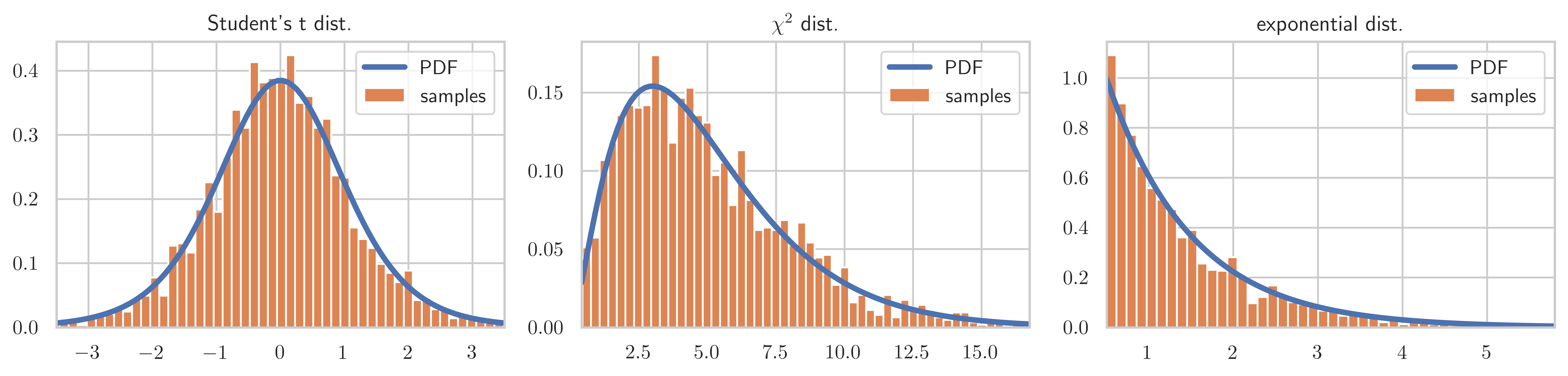
X = stats.chi2(df=5)X_samples = X.rvs(500)df, loc, scale = stats.chi2.fit(X_samples)df, loc, scale(np.float64(4.728645123391404),
np.float64(0.03257330219133387),
np.float64(1.0734482977974253))Y = stats.chi2(df=df, loc=loc, scale=scale)fig, ax = plt.subplots(1, 1, figsize=(8, 3))
x_lim = X.interval(0.99)
x = np.linspace(*x_lim, num=100)
ax.plot(x, X.pdf(x), label="original")
ax.plot(x, Y.pdf(x), label="recreated")
ax.legend()
fig.tight_layout()
fig.savefig("ch13-max-likelihood-fit.pdf")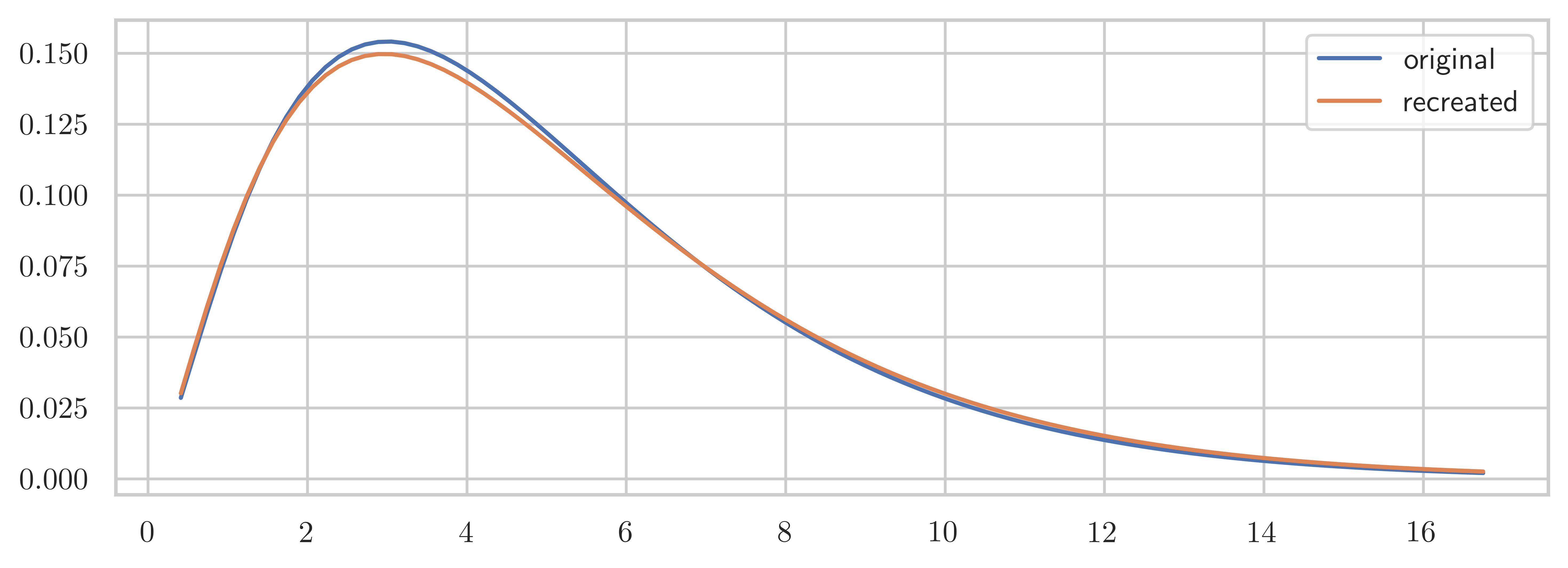
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
x_lim = X.interval(0.99)
x = np.linspace(*x_lim, num=100)
axes[0].plot(x, X.pdf(x), label="original")
axes[0].plot(x, Y.pdf(x), label="recreated")
axes[0].legend()
axes[1].plot(x, X.pdf(x) - Y.pdf(x), label="error")
axes[1].legend()
fig.tight_layout()
fig.savefig("ch13-max-likelihood-fit.pdf")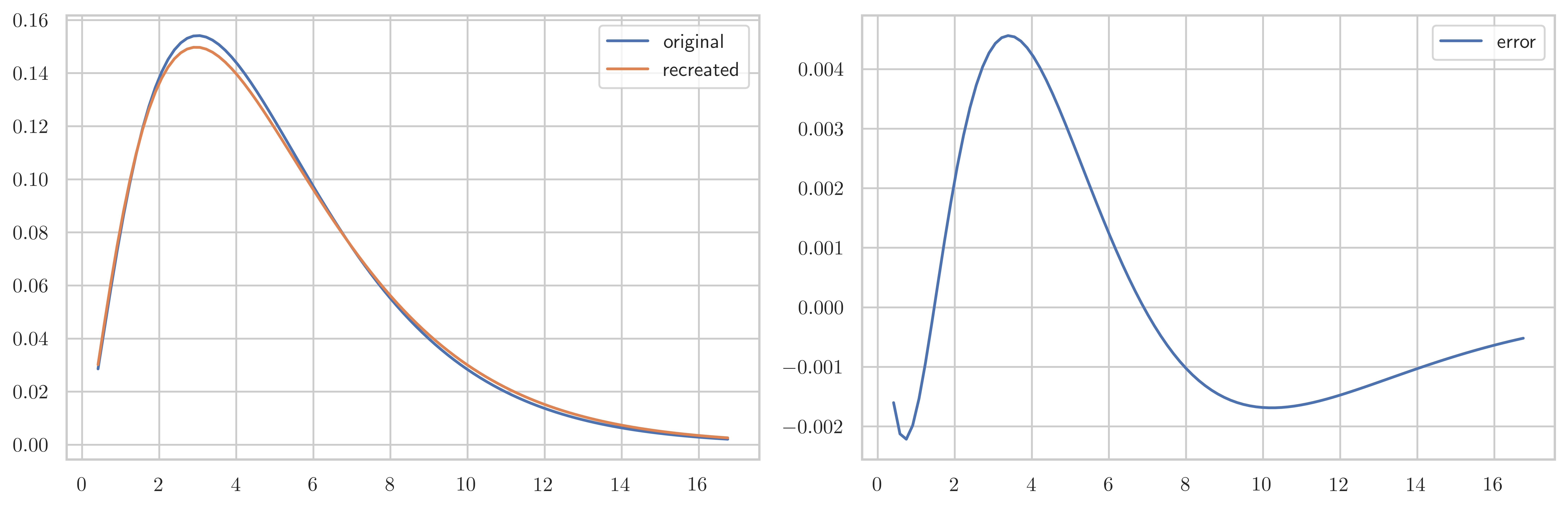
Hypothesis testing¶
np.random.seed(123456789)mu, sigma = 1.0, 0.5X = stats.norm(mu - 0.2, sigma)n = 100X_samples = X.rvs(n)z = (X_samples.mean() - mu) / (sigma / np.sqrt(n))znp.float64(-2.8338979550098298)t = (X_samples.mean() - mu) / (X_samples.std(ddof=1) / np.sqrt(n))tnp.float64(-2.9680338545657845)stats.norm().ppf(0.025)np.float64(-1.9599639845400545)2 * stats.norm().cdf(-abs(z))np.float64(0.004598401329075357)2 * stats.t(df=(n - 1)).cdf(-abs(t))np.float64(0.003758647967422724)t, p = stats.ttest_1samp(X_samples, mu)tnp.float64(-2.968033854565784)pnp.float64(0.003758647967422724)fig, ax = plt.subplots(figsize=(8, 3))
sns.histplot(X_samples, ax=ax, kde=True, stat="density")
x = np.linspace(*X.interval(0.999), num=100)
ax.plot(x, stats.norm(loc=mu, scale=sigma).pdf(x))
fig.tight_layout()
fig.savefig("ch13-hypothesis-test-dist-sample-mean.pdf")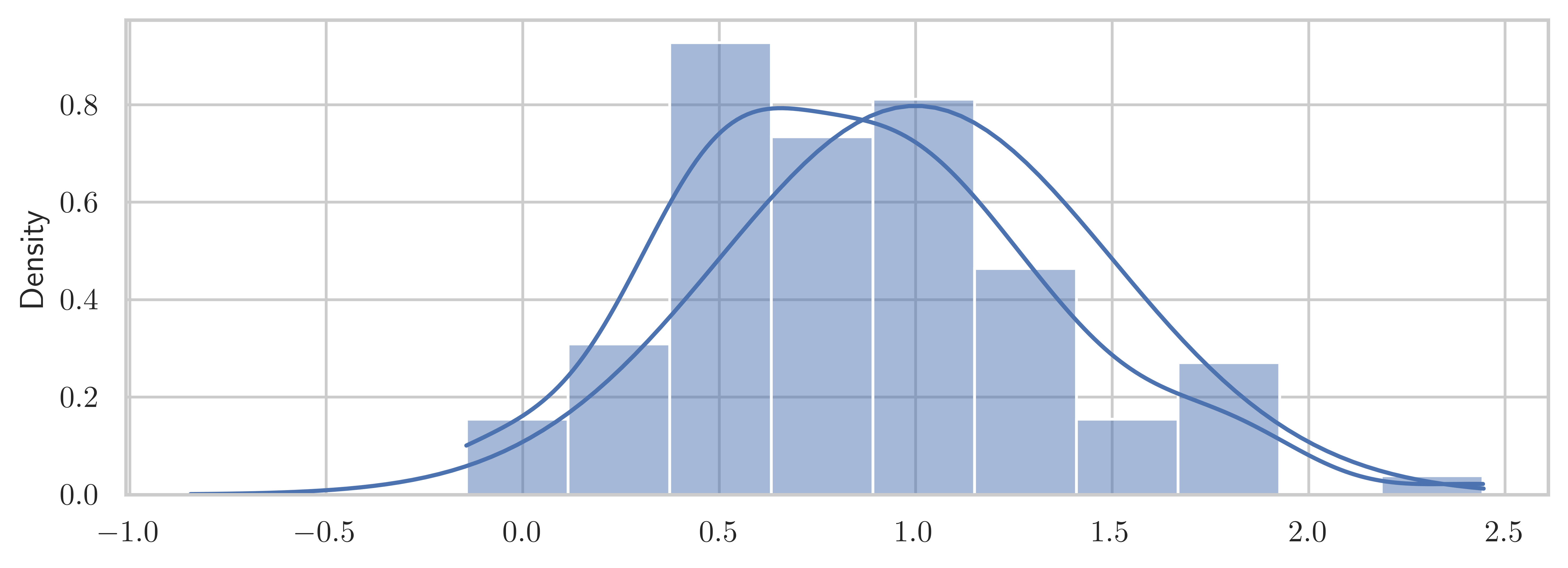
n = 50mu1, mu2 = np.random.rand(2)mu1, mu2(np.float64(0.24764580637159606), np.float64(0.42145435527527897))X1 = stats.norm(mu1, sigma)X1_sample = X1.rvs(n)X2 = stats.norm(mu2, sigma)X2_sample = X2.rvs(n)t, p = stats.ttest_ind(X1_sample, X2_sample)tnp.float64(-1.4283175246005888)pnp.float64(0.1563798105967323)mu1, mu2(np.float64(0.24764580637159606), np.float64(0.42145435527527897))sns.histplot(X1_sample)
sns.histplot(X2_sample)<Axes: ylabel='Count'>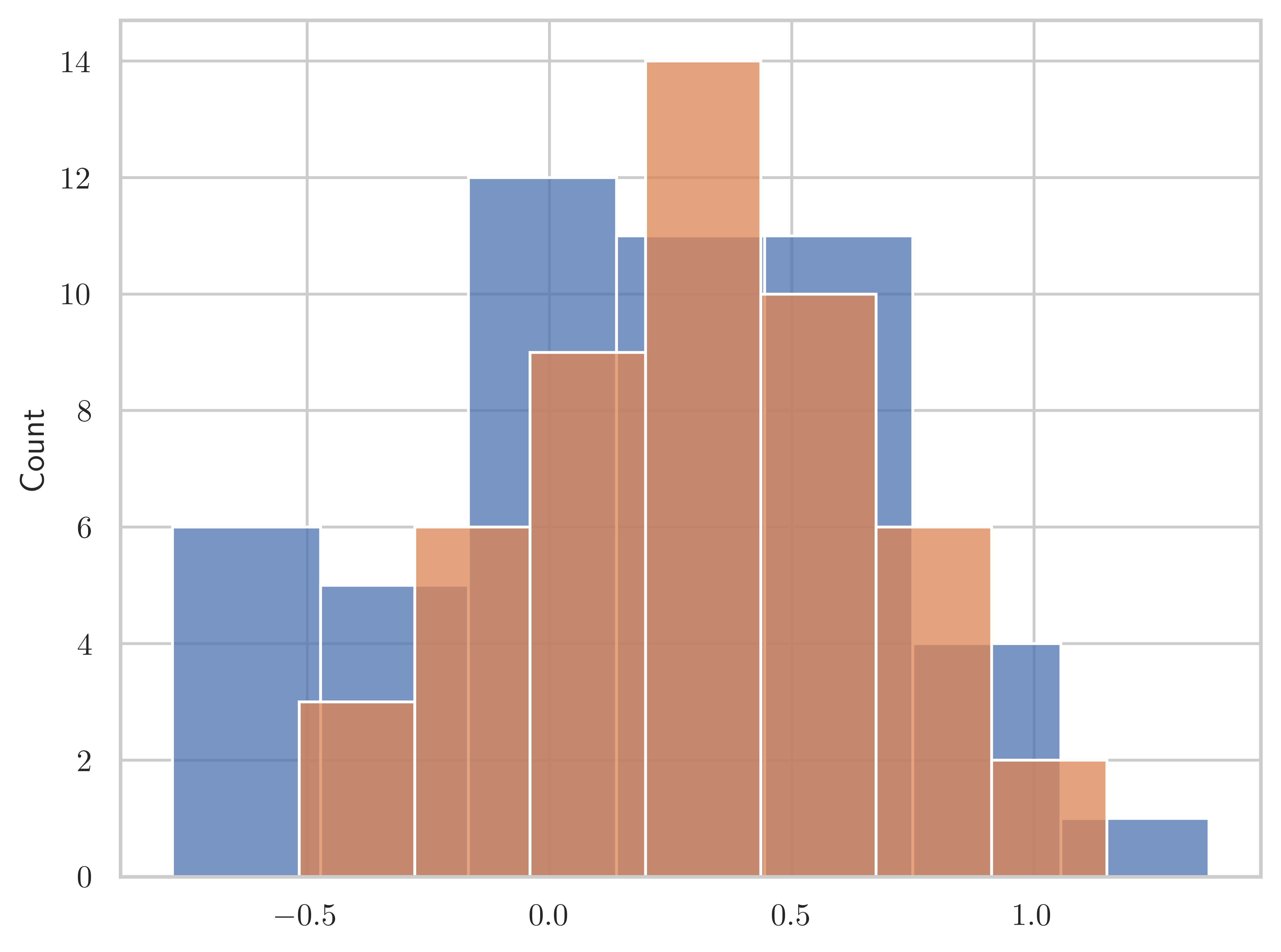
Nonparameteric methods¶
np.random.seed(0)X = stats.chi2(df=5)X_samples = X.rvs(100)kde = stats.gaussian_kde(X_samples)kde_low_bw = stats.gaussian_kde(X_samples, bw_method=0.25)x = np.linspace(0, 20, 100)fig, axes = plt.subplots(1, 3, figsize=(12, 3))
axes[0].hist(X_samples, density=True, alpha=0.5, bins=25)
axes[1].plot(x, kde(x), label="KDE")
axes[1].plot(x, kde_low_bw(x), label="KDE (low bw)")
axes[1].plot(x, X.pdf(x), label="True PDF")
axes[1].legend()
sns.histplot(X_samples, bins=25, ax=axes[2], kde=True, stat="density")
fig.tight_layout()
fig.savefig("ch13-hist-kde.pdf")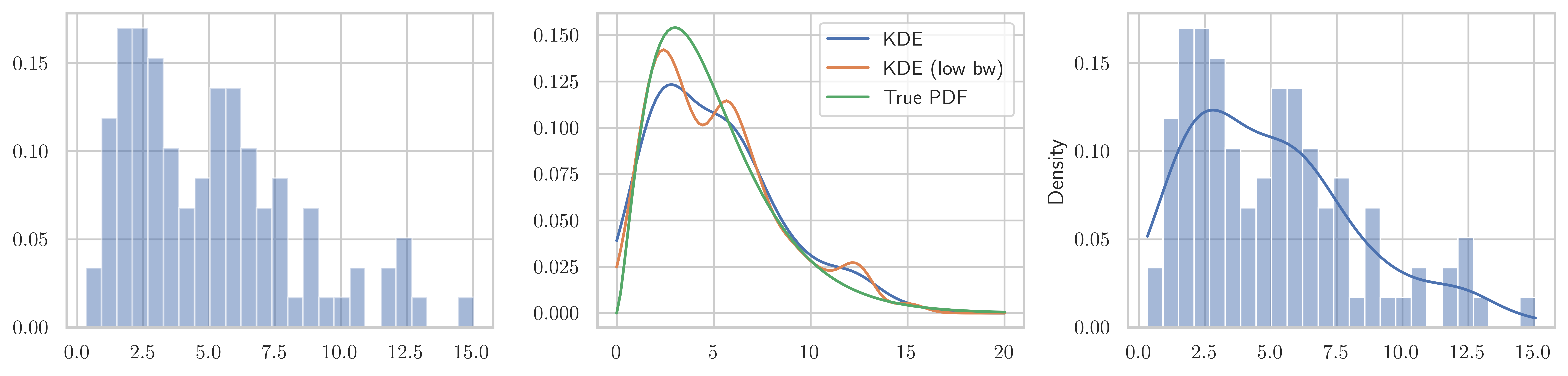
kde.resample(10)array([[ 1.10979087, 0.4379679 , 14.20879078, 5.94683846, 1.78490438,
5.58416739, 4.18349885, 2.78527976, 0.68112826, 7.7643985 ]])def _kde_cdf(x):
return kde.integrate_box_1d(-np.inf, x)kde_cdf = np.vectorize(_kde_cdf)fig, ax = plt.subplots(1, 1, figsize=(8, 3))
sns.histplot(X_samples, bins=25, ax=ax, kde=True, stat="density")
x = np.linspace(0, 20, 100)
ax.plot(x, kde_cdf(x))
fig.tight_layout()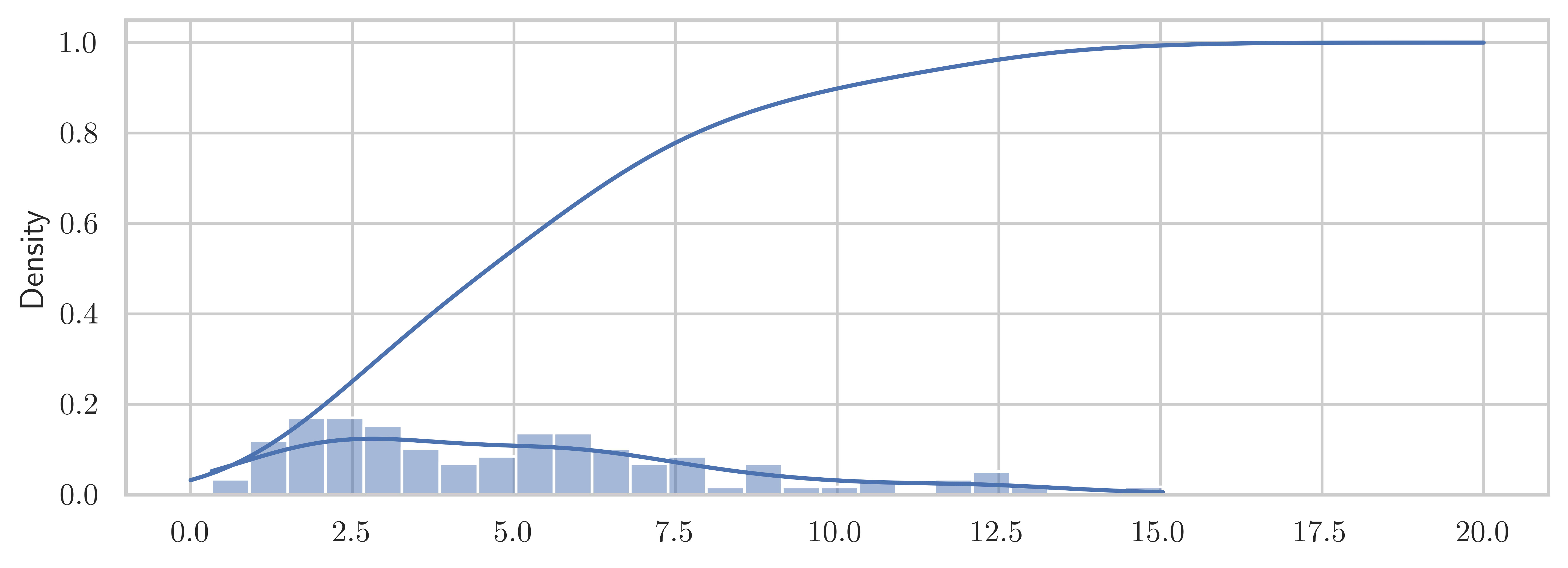
def _kde_ppf(q):
return optimize.fsolve(lambda x, q: kde_cdf(x) - q, kde.dataset.mean(), args=(q,))[
0
]kde_ppf = np.vectorize(_kde_ppf)kde_ppf([0.05, 0.95])array([ 0.39074674, 11.94993578])X.ppf([0.05, 0.95])array([ 1.14547623, 11.07049769])- Johansson, R. (2024). Numerical Python: Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib. Apress. 10.1007/979-8-8688-0413-7