Chapter 7: Interpolation
Robert Johansson
Source code listings for Numerical Python - Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib (ISBN 979-8-8688-0412-0).
%matplotlib inline
import matplotlib as mpl
mpl.rcParams["font.family"] = "serif"
mpl.rcParams["font.size"] = "12"
mpl.rcParams["mathtext.fontset"] = "stix"
mpl.rcParams["font.family"] = "serif"
mpl.rcParams["font.sans-serif"] = "stix"import numpy as npfrom numpy import polynomial as Pfrom scipy import interpolateimport matplotlib.pyplot as pltfrom scipy import linalgPolynomials¶
p1 = P.Polynomial([1, 2, 3])p1Loading...
p1.__repr__()"Polynomial([1., 2., 3.], domain=[-1., 1.], window=[-1., 1.], symbol='x')"p2 = P.Polynomial.fromroots([-1, 1])p2Loading...
p1.roots()array([-0.33333333-0.47140452j, -0.33333333+0.47140452j])p2.roots()array([-1., 1.])p1.coefarray([1., 2., 3.])p1.domainarray([-1., 1.])p1.windowarray([-1., 1.])p1(np.array([1.5, 2.5, 3.5]))array([10.75, 24.75, 44.75])(p1 + p2).__repr__<bound method ABCPolyBase.__repr__ of Polynomial([0., 2., 4.], domain=[-1., 1.], window=[-1., 1.], symbol='x')>p2 / 5Loading...
p1 = P.Polynomial.fromroots([1, 2, 3])p1Loading...
p2 = P.Polynomial.fromroots([2])p2Loading...
p3 = p1 // p2p3Loading...
p3.roots()array([1., 3.])p2Loading...
c1 = P.Chebyshev([1, 2, 3])c1Loading...
c1.roots()array([-0.76759188, 0.43425855])c = P.Chebyshev.fromroots([-1, 1])cLoading...
l = P.Legendre.fromroots([-1, 1])lLoading...
c(np.array([0.5, 1.5, 2.5]))array([-0.75, 1.25, 5.25])l(np.array([0.5, 1.5, 2.5]))array([-0.75, 1.25, 5.25])Polynomial interpolation¶
x = np.array([1, 2, 3, 4])
y = np.array([1, 3, 5, 4])deg = len(x) - 1A = P.polynomial.polyvander(x, deg)c = linalg.solve(A, y)carray([ 2. , -3.5, 3. , -0.5])f1 = P.Polynomial(c)f1(2.5)np.float64(4.1875)A = P.chebyshev.chebvander(x, deg)c = linalg.solve(A, y)carray([ 3.5 , -3.875, 1.5 , -0.125])f2 = P.Chebyshev(c)f2(2.5)np.float64(4.187500000000001)xx = np.linspace(x.min(), x.max(), 100)
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
ax.plot(xx, f1(xx), "b", lw=2, label="Power basis interp.")
ax.plot(xx, f2(xx), "r--", lw=2, label="Chebyshev basis interp.")
ax.scatter(x, y, label="data points")
ax.legend(loc=4)
ax.set_xticks(x)
ax.set_ylabel(r"$y$", fontsize=18)
ax.set_xlabel(r"$x$", fontsize=18)
fig.tight_layout()
fig.savefig("ch7-polynomial-interpolation.pdf");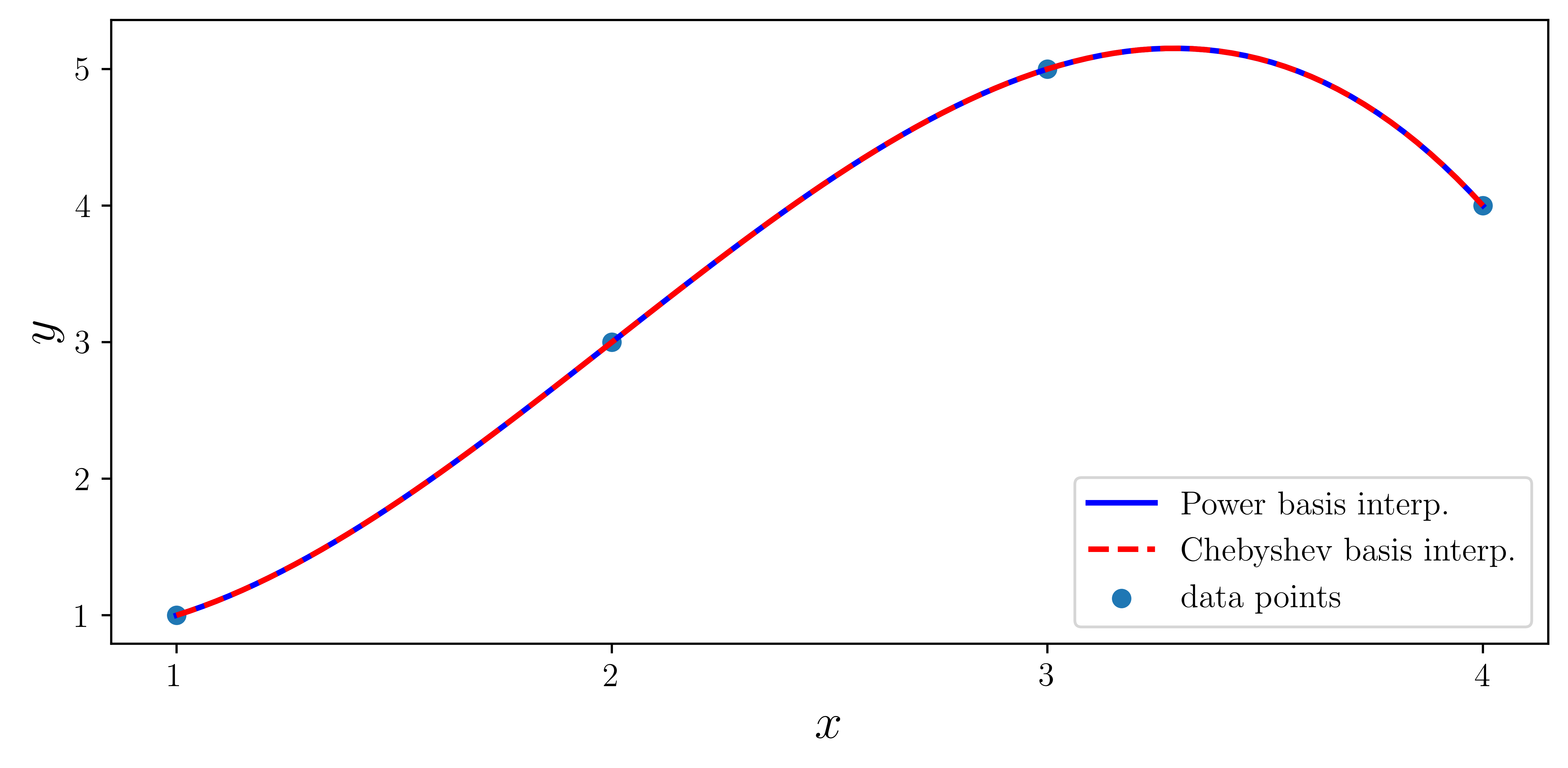
f1b = P.Polynomial.fit(x, y, deg)f1bLoading...
f2b = P.Chebyshev.fit(x, y, deg)f2bLoading...
np.linalg.cond(P.chebyshev.chebvander(x, deg))np.float64(4659.738424140432)np.linalg.cond(P.chebyshev.chebvander((2 * x - 5) / 3.0, deg))np.float64(1.8542033440472903)(2 * x - 5) / 3.0array([-1. , -0.33333333, 0.33333333, 1. ])f1 = P.Polynomial.fit(x, y, 1)
f2 = P.Polynomial.fit(x, y, 2)
f3 = P.Polynomial.fit(x, y, 3)xx = np.linspace(x.min(), x.max(), 100)
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
ax.plot(xx, f1(xx), "r", lw=2, label="1st order")
ax.plot(xx, f2(xx), "g", lw=2, label="2nd order")
ax.plot(xx, f3(xx), "b", lw=2, label="3rd order")
ax.scatter(x, y, label="data points")
ax.legend(loc=4)
ax.set_xticks(x)
ax.set_ylabel(r"$y$", fontsize=18)
ax.set_xlabel(r"$x$", fontsize=18);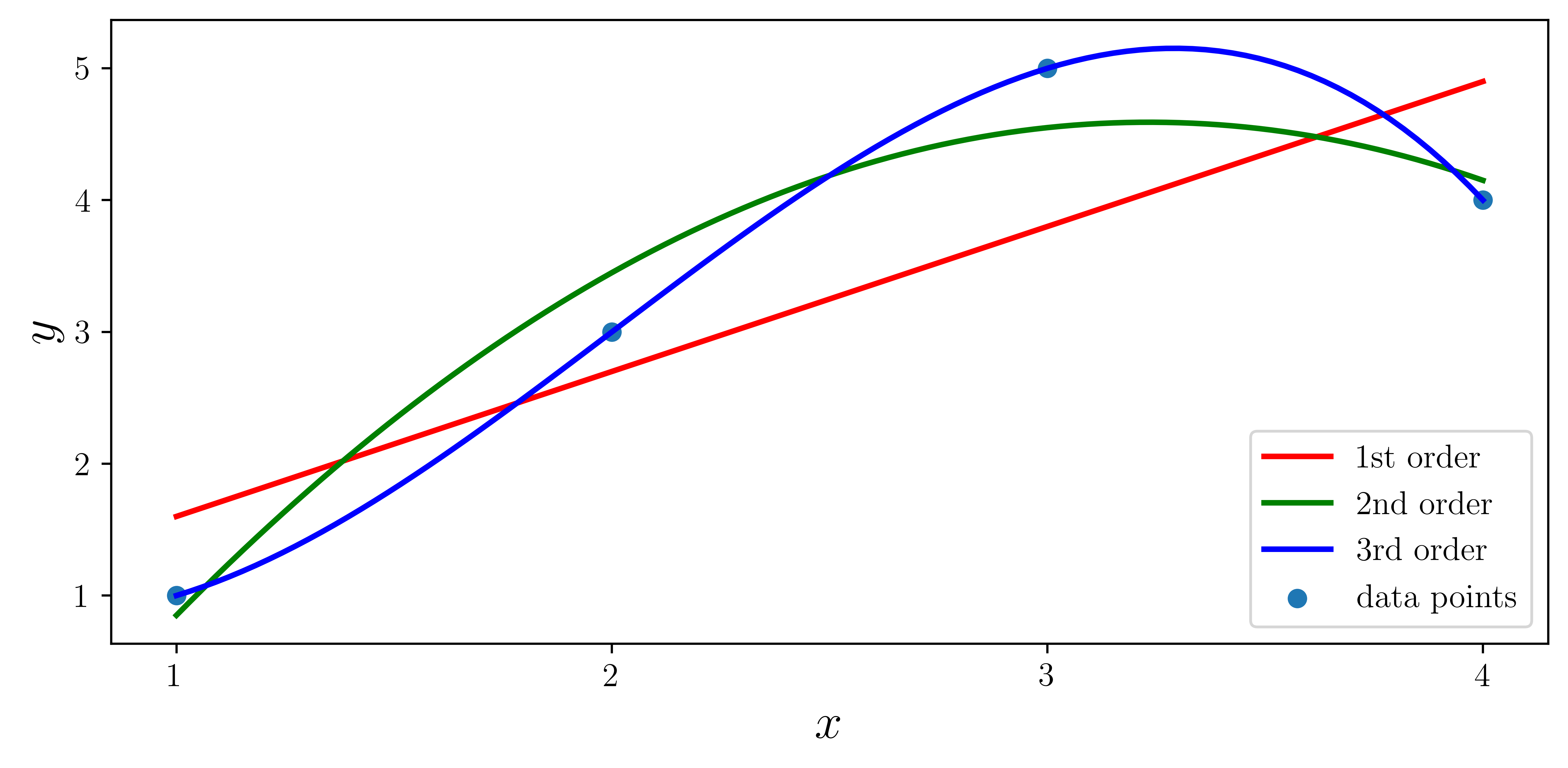
Runge problem¶
def runge(x):
return 1 / (1 + 25 * x**2)def runge_interpolate(n):
x = np.linspace(-1, 1, n + 1)
p = P.Polynomial.fit(x, runge(x), deg=n)
return x, pxx = np.linspace(-1, 1, 250)fig, ax = plt.subplots(1, 1, figsize=(8, 4))
ax.plot(xx, runge(xx), "k", lw=2, label="Runge's function")
n = 13
x, p = runge_interpolate(n)
ax.plot(x, runge(x), "ro")
ax.plot(xx, p(xx), "r", label="interp. order %d" % n)
n = 14
x, p = runge_interpolate(n)
ax.plot(x, runge(x), "go")
ax.plot(xx, p(xx), "g", label="interp. order %d" % n)
ax.legend(loc=8)
ax.set_xlim(-1.1, 1.1)
ax.set_ylim(-1, 2)
ax.set_xticks([-1, -0.5, 0, 0.5, 1])
ax.set_ylabel(r"$y$", fontsize=18)
ax.set_xlabel(r"$x$", fontsize=18)
fig.tight_layout()
fig.savefig("ch7-polynomial-interpolation-runge.pdf");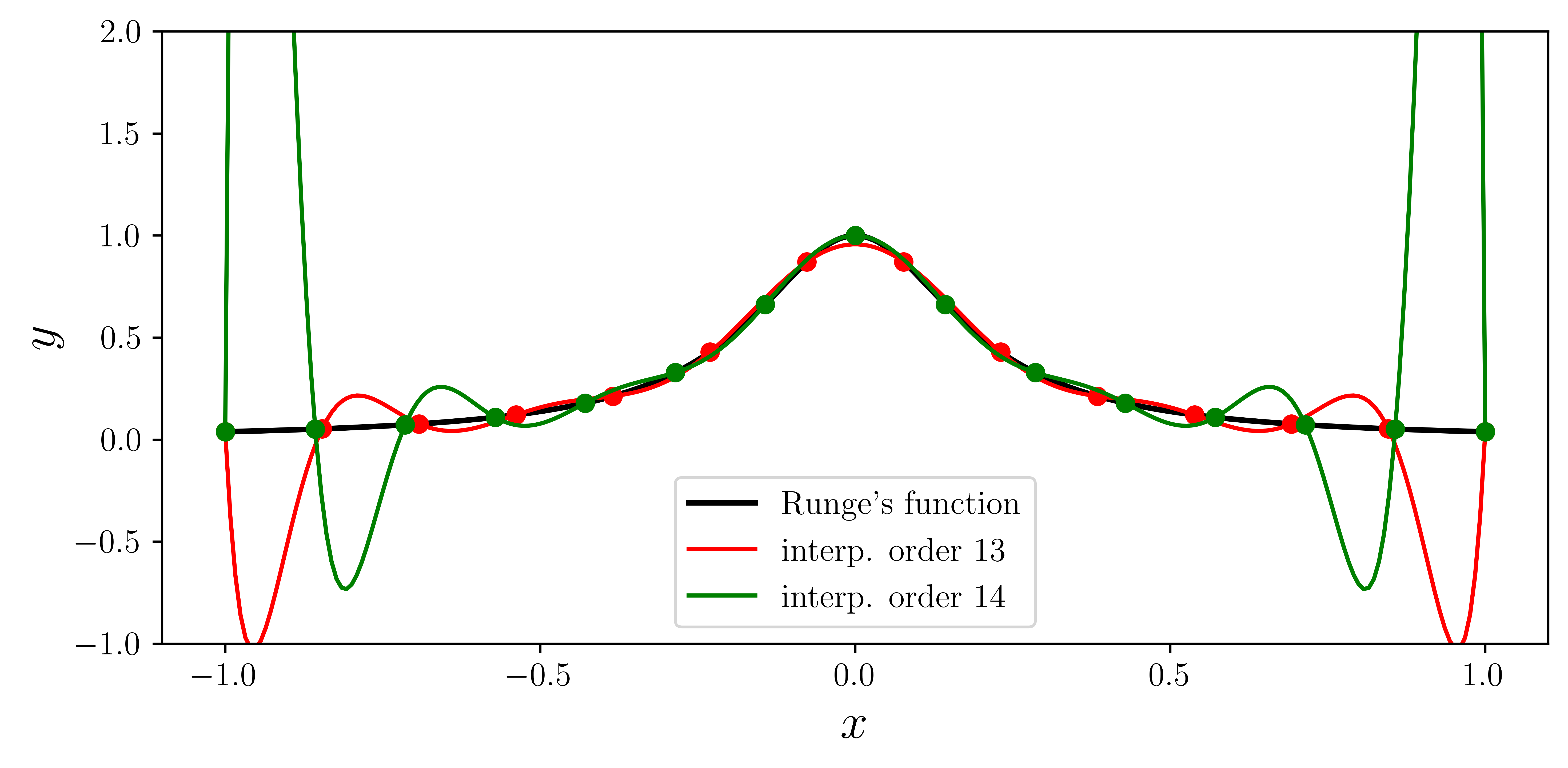
Spline interpolation¶
x = np.linspace(-1, 1, 11)y = runge(x)f = interpolate.interp1d(x, y, kind=3)xx = np.linspace(-1, 1, 100)fig, ax = plt.subplots(figsize=(8, 4))
ax.plot(xx, runge(xx), "k", lw=1, label="Runge's function")
ax.plot(x, y, "ro", label="sample points")
ax.plot(xx, f(xx), "r--", lw=2, label="spline order 3")
ax.legend()
ax.set_ylim(0, 1.1)
ax.set_xticks([-1, -0.5, 0, 0.5, 1])
ax.set_ylabel(r"$y$", fontsize=18)
ax.set_xlabel(r"$x$", fontsize=18)
fig.tight_layout()
fig.savefig("ch7-spline-interpolation-runge.pdf");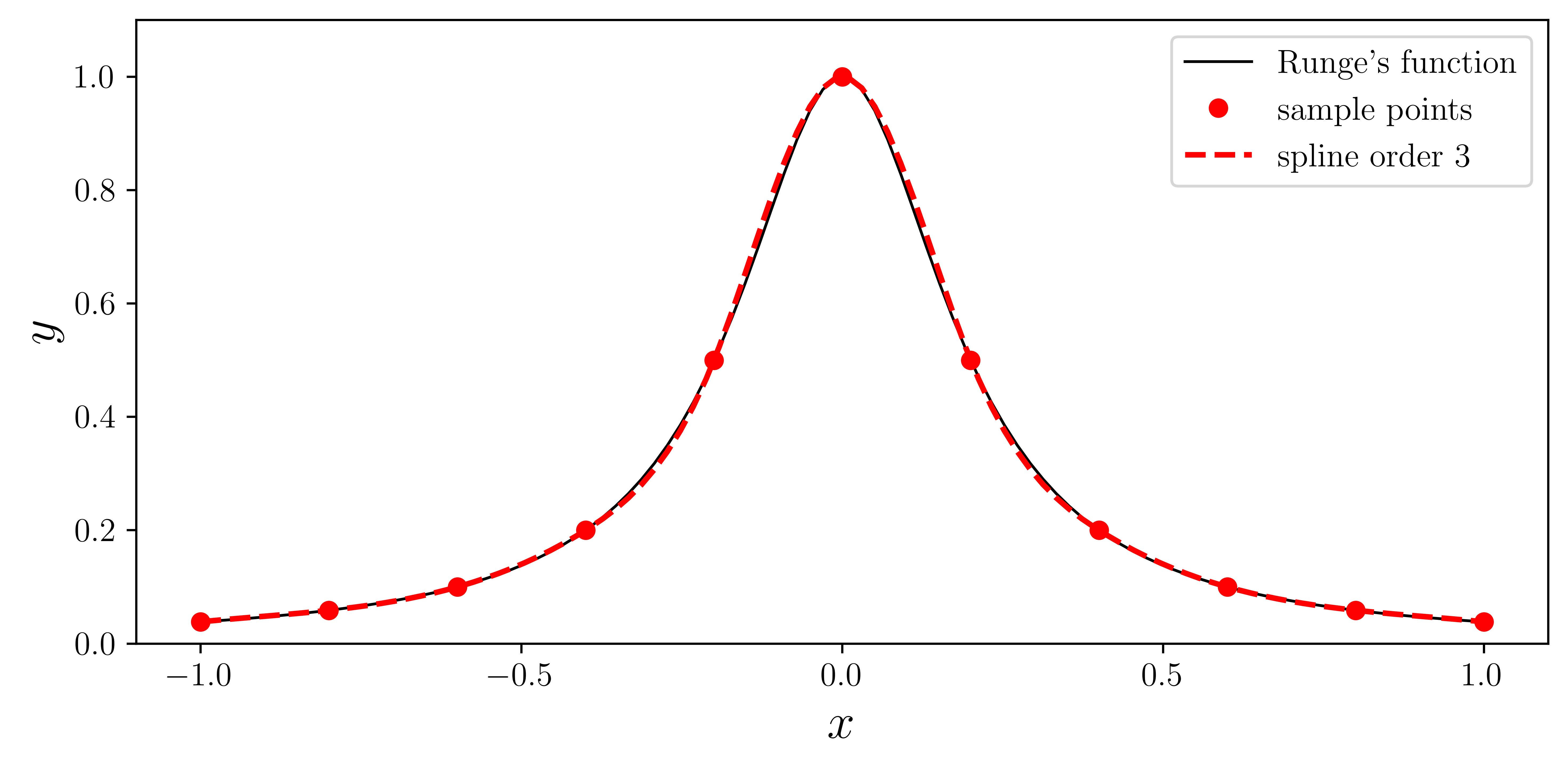
x = np.array([0, 1, 2, 3, 4, 5, 6, 7])y = np.array([3, 4, 3.5, 2, 1, 1.5, 1.25, 0.9])xx = np.linspace(x.min(), x.max(), 100)fig, ax = plt.subplots(figsize=(8, 4))
ax.scatter(x, y)
for n in [1, 2, 3, 5]:
f = interpolate.interp1d(x, y, kind=n)
ax.plot(xx, f(xx), label="order %d" % n)
ax.legend()
ax.set_ylabel(r"$y$", fontsize=18)
ax.set_xlabel(r"$x$", fontsize=18)
fig.tight_layout()
fig.savefig("ch7-spline-interpolation-orders.pdf");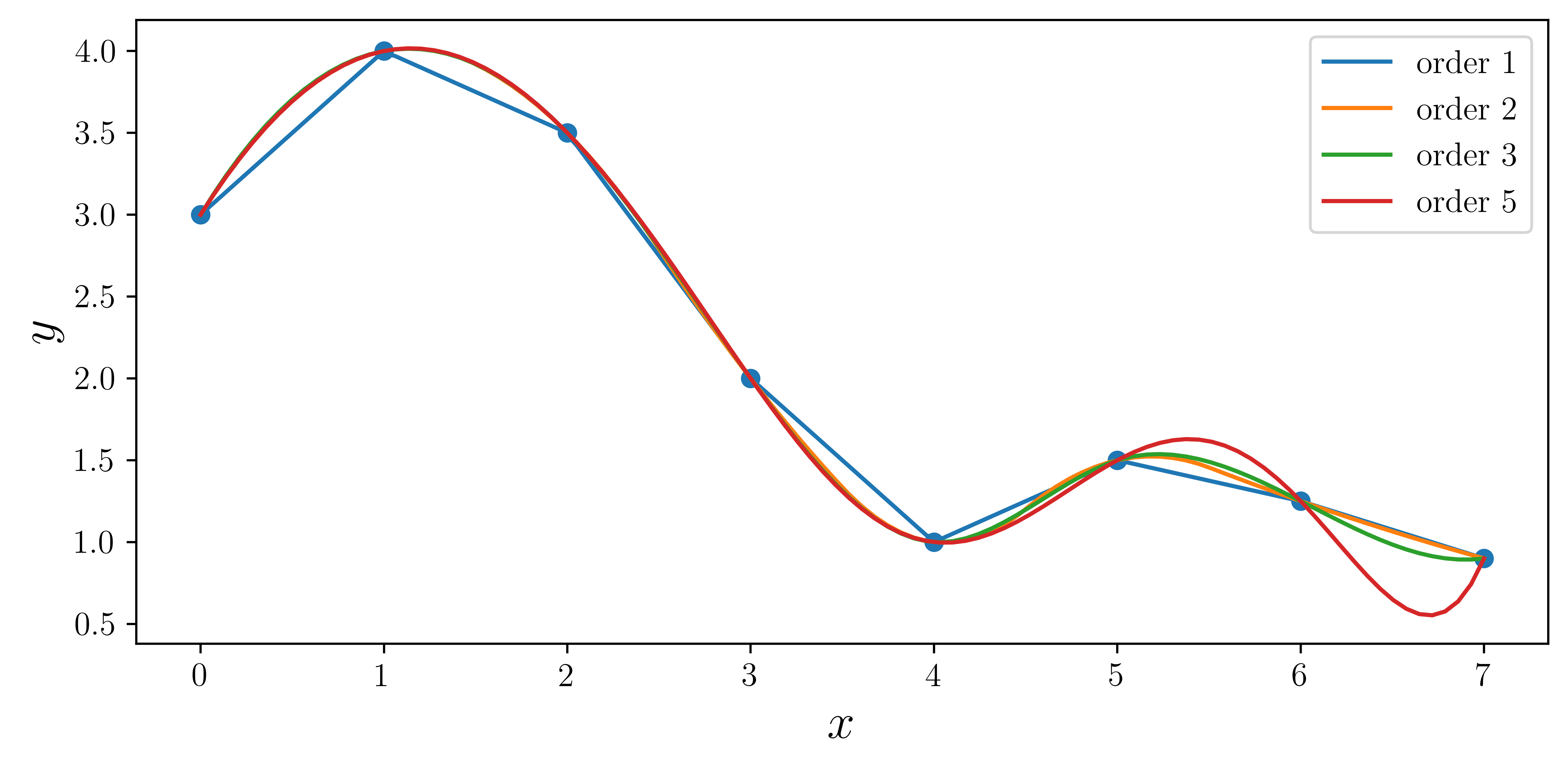
Multivariate interpolation¶
Regular grid¶
x = y = np.linspace(-2, 2, 10)def f(x, y):
return np.exp(-((x + 0.5) ** 2) - 2 * (y + 0.5) ** 2) - np.exp(
-((x - 0.5) ** 2) - 2 * (y - 0.5) ** 2
)X, Y = np.meshgrid(x, y)# simulate noisy data at fixed grid points X, Y
Z = f(X, Y) + 0.05 * np.random.randn(*X.shape)f_interp = interpolate.RegularGridInterpolator((x, y), Z, method="cubic")xx = yy = np.linspace(x.min(), x.max(), 100)XX, YY = np.meshgrid(xx, yy)
ZZi = f_interp(np.vstack((XX.ravel(), YY.ravel())).T).reshape(XX.shape)XX, YY = np.meshgrid(xx, yy)fig, axes = plt.subplots(1, 2, figsize=(12, 5))
c = axes[0].contourf(XX, YY, f(XX, YY), 15, cmap=plt.cm.RdBu)
axes[0].set_xlabel(r"$x$", fontsize=20)
axes[0].set_ylabel(r"$y$", fontsize=20)
axes[0].set_title("exact / high sampling")
cb = fig.colorbar(c, ax=axes[0])
cb.set_label(r"$z$", fontsize=20)
c = axes[1].contourf(XX, YY, ZZi, 15, cmap=plt.cm.RdBu)
axes[1].set_ylim(-2.1, 2.1)
axes[1].set_xlim(-2.1, 2.1)
axes[1].set_xlabel(r"$x$", fontsize=20)
axes[1].set_ylabel(r"$y$", fontsize=20)
axes[1].scatter(X, Y, marker="x", color="k")
axes[1].set_title("interpolation of noisy data / low sampling")
cb = fig.colorbar(c, ax=axes[1])
cb.set_label(r"$z$", fontsize=20)
fig.tight_layout()
fig.savefig("ch7-multivariate-interpolation-regular-grid.pdf")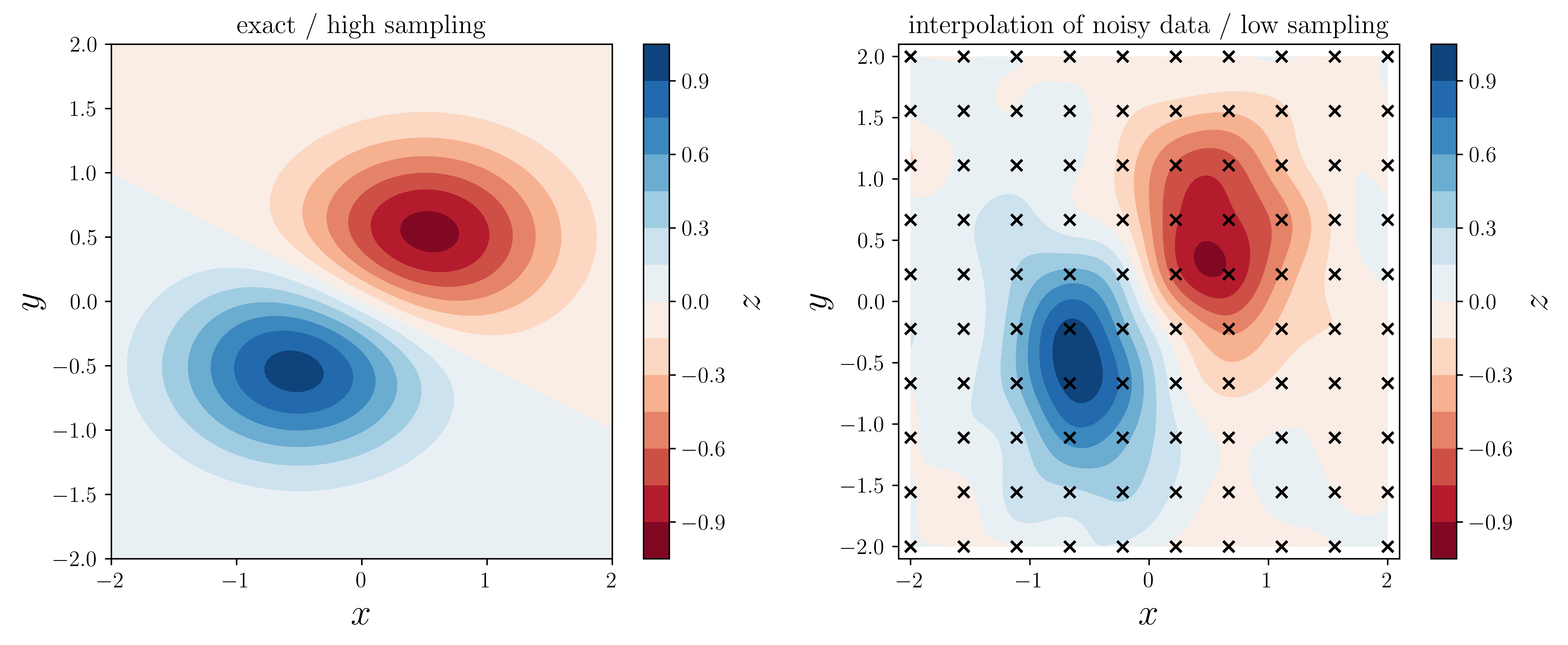
fig, ax = plt.subplots(1, 1, figsize=(6, 5))
c = ax.contourf(XX, YY, ZZi, 15, cmap=plt.cm.RdBu)
ax.set_ylim(-2.1, 2.1)
ax.set_xlim(-2.1, 2.1)
ax.set_xlabel(r"$x$", fontsize=20)
ax.set_ylabel(r"$y$", fontsize=20)
ax.scatter(X, Y, marker="x", color="k")
cb = fig.colorbar(c, ax=ax)
cb.set_label(r"$z$", fontsize=20)
fig.tight_layout()
# fig.savefig('ch7-multivariate-interpolation-regular-grid.pdf')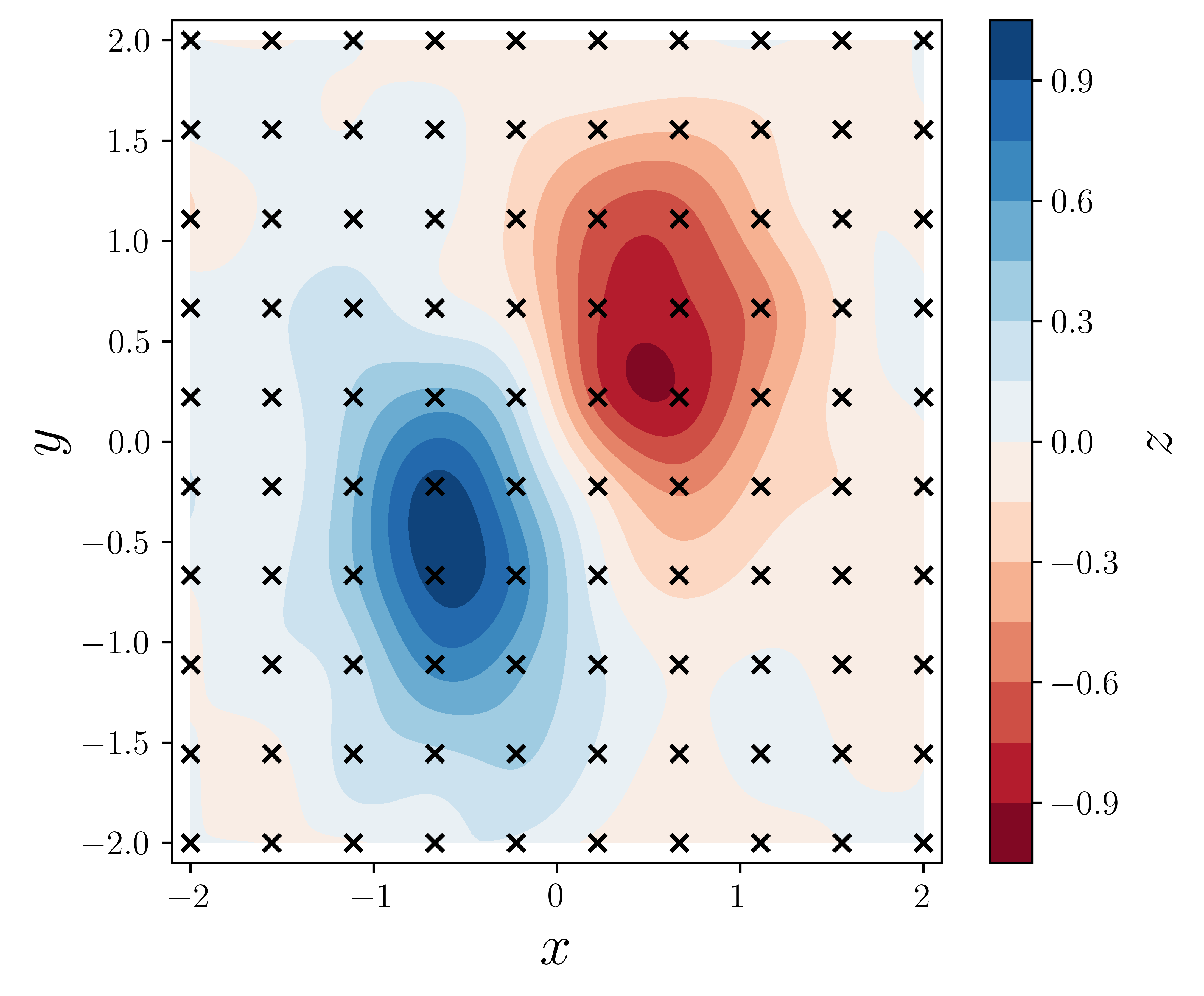
Irregular grid¶
np.random.seed(115925231)x = y = np.linspace(-1, 1, 100)X, Y = np.meshgrid(x, y)def f(x, y):
return np.exp(-(x**2) - y**2) * np.cos(4 * x) * np.sin(6 * y)Z = f(X, Y)N = 500xdata = np.random.uniform(-1, 1, N)ydata = np.random.uniform(-1, 1, N)zdata = f(xdata, ydata)fig, ax = plt.subplots(figsize=(8, 6))
c = ax.contourf(X, Y, Z, 15, cmap=plt.cm.RdBu)
ax.scatter(xdata, ydata, marker=".", color="black")
ax.set_ylim(-1, 1)
ax.set_xlim(-1, 1)
ax.set_xlabel(r"$x$", fontsize=20)
ax.set_ylabel(r"$y$", fontsize=20)
cb = fig.colorbar(c, ax=ax)
cb.set_label(r"$z$", fontsize=20)
fig.tight_layout()
fig.savefig("ch7-multivariate-interpolation-exact.pdf");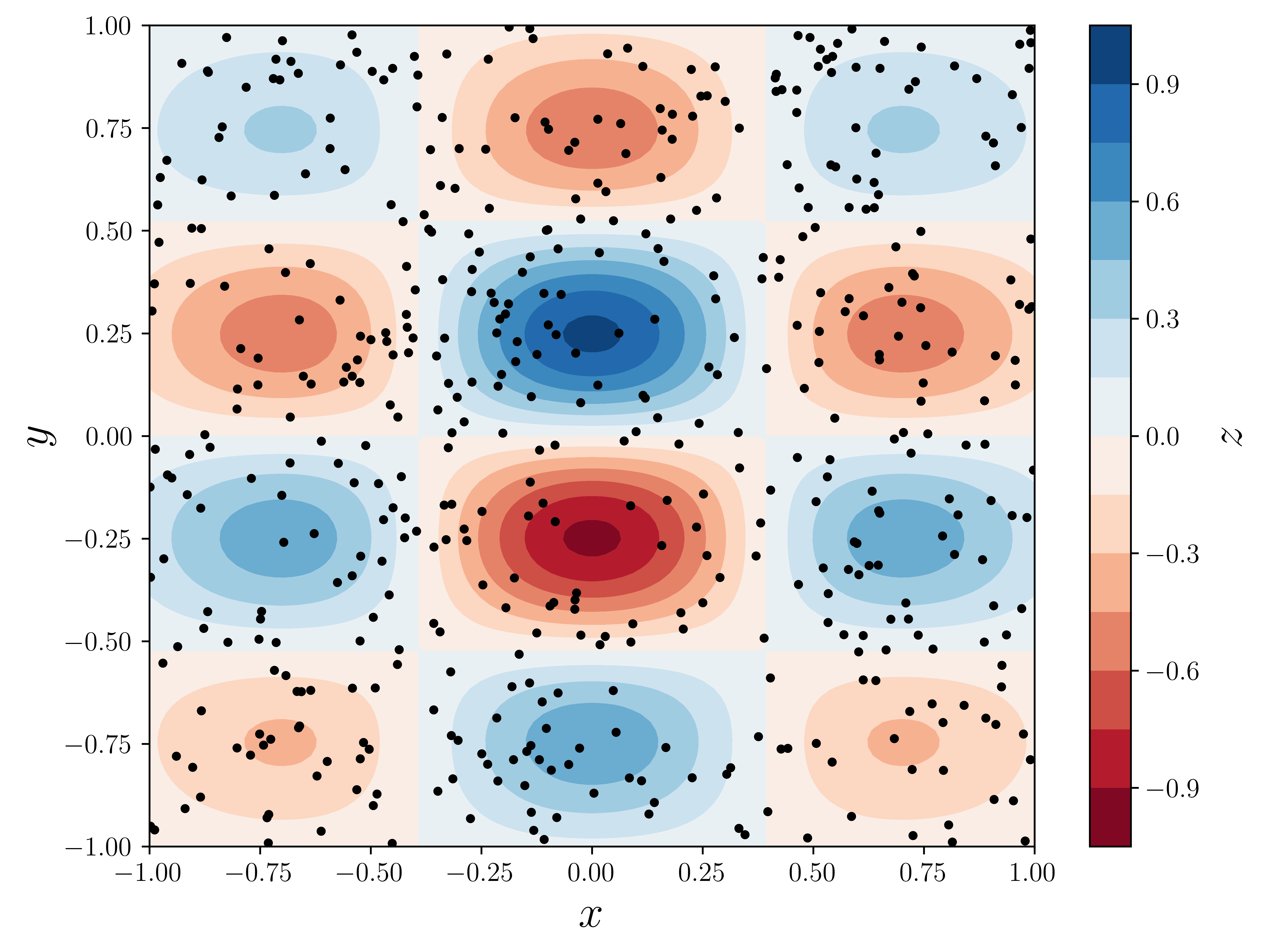
def z_interpolate(xdata, ydata, zdata):
Zi_0 = interpolate.griddata((xdata, ydata), zdata, (X, Y), method="nearest")
Zi_1 = interpolate.griddata((xdata, ydata), zdata, (X, Y), method="linear")
Zi_3 = interpolate.griddata((xdata, ydata), zdata, (X, Y), method="cubic")
return Zi_0, Zi_1, Zi_3fig, axes = plt.subplots(3, 3, figsize=(12, 12), sharex=True, sharey=True)
n_vec = [50, 150, 500]
for idx, n in enumerate(n_vec):
Zi_0, Zi_1, Zi_3 = z_interpolate(xdata[:n], ydata[:n], zdata[:n])
axes[idx, 0].contourf(X, Y, Zi_0, 15, cmap=plt.cm.RdBu)
axes[idx, 0].set_ylabel("%d data points\ny" % n, fontsize=16)
axes[idx, 0].set_title("nearest", fontsize=16)
axes[idx, 1].contourf(X, Y, Zi_1, 15, cmap=plt.cm.RdBu)
axes[idx, 1].set_title("linear", fontsize=16)
axes[idx, 2].contourf(X, Y, Zi_3, 15, cmap=plt.cm.RdBu)
axes[idx, 2].set_title("cubic", fontsize=16)
for m in range(len(n_vec)):
axes[2, m].set_xlabel("x", fontsize=16)
fig.tight_layout()
fig.savefig("ch7-multivariate-interpolation-interp.pdf");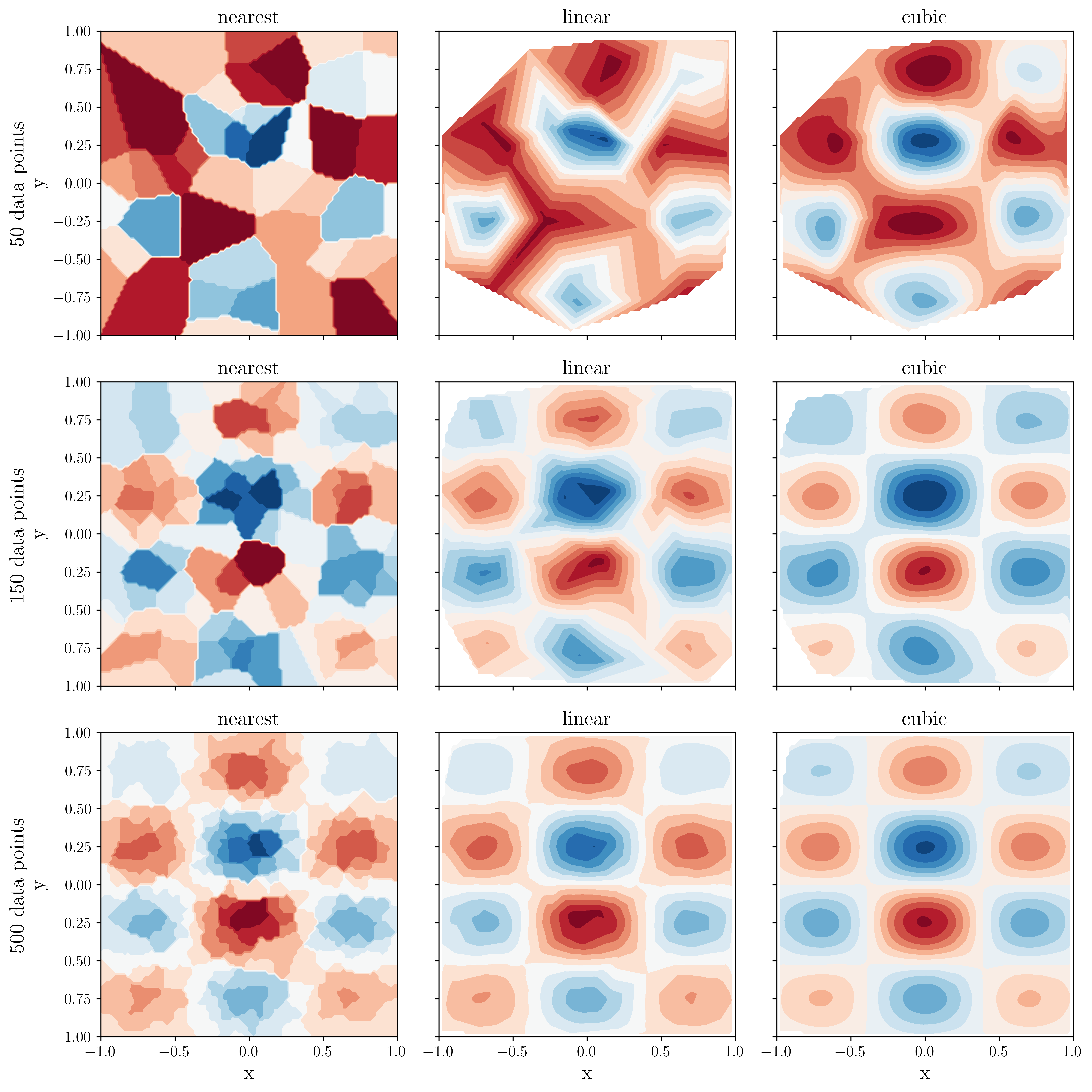
- Johansson, R. (2024). Numerical Python: Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib. Apress. 10.1007/979-8-8688-0413-7