Chapter 9: Ordinary differential equations
Robert Johansson
Source code listings for Numerical Python - Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib (ISBN 979-8-8688-0412-0).
import numpy as np%matplotlib inline
%config InlineBackend.figure_format='retina'
import matplotlib as mpl
import matplotlib.pyplot as plt
# mpl.rcParams['text.usetex'] = True
mpl.rcParams["mathtext.fontset"] = "stix"
mpl.rcParams["font.family"] = "serif"
mpl.rcParams["font.sans-serif"] = "stix"import sympy
sympy.init_printing()from scipy import integrateSymbolic ODE solving with SymPy¶
Newton’s law of cooling¶
t, k, T0, Ta = sympy.symbols("t, k, T_0, T_a")T = sympy.Function("T")ode = T(t).diff(t) + k * (T(t) - Ta)odeLoading...
ode_sol = sympy.dsolve(ode)ode_solLoading...
ode_sol.lhsLoading...
ode_sol.rhsLoading...
ics = {T(0): T0}icsLoading...
C_eq = ode_sol.subs(t, 0).subs(ics)C_eqLoading...
C_sol = sympy.solve(C_eq)C_solLoading...
ode_sol.subs(C_sol[0])Loading...
Function for applying initial conditions¶
def apply_ics(sol, ics, x, known_params):
"""
Apply the initial conditions (ics), given as a dictionary on
the form ics = {y(0): y0: y(x).diff(x).subs(x, 0): yp0, ...}
to the solution of the ODE with indepdendent variable x.
The undetermined integration constants C1, C2, ... are extracted
from the free symbols of the ODE solution, excluding symbols in
the known_params list.
"""
free_params = sol.free_symbols - set(known_params)
eqs = [(sol.lhs - sol.rhs).diff(x, n).subs(x, 0).subs(ics) for n in range(len(ics))]
sol_params = sympy.solve(eqs, free_params)
return sol.subs(sol_params)ode_solLoading...
apply_ics(ode_sol, ics, t, [k, Ta])Loading...
ode_sol = apply_ics(ode_sol, ics, t, [k, Ta]).simplify()ode_solLoading...
y_x = sympy.lambdify((t, k), ode_sol.rhs.subs({T0: 5, Ta: 1}), "numpy")fig, ax = plt.subplots(figsize=(8, 4))
x = np.linspace(0, 4, 100)
for k in [1, 2, 3]:
ax.plot(x, y_x(x, k), label=r"$k=%d$" % k)
ax.set_title(r"$%s$" % sympy.latex(ode_sol), fontsize=18)
ax.set_xlabel(r"$x$", fontsize=18)
ax.set_ylabel(r"$y$", fontsize=18)
ax.legend()
fig.tight_layout()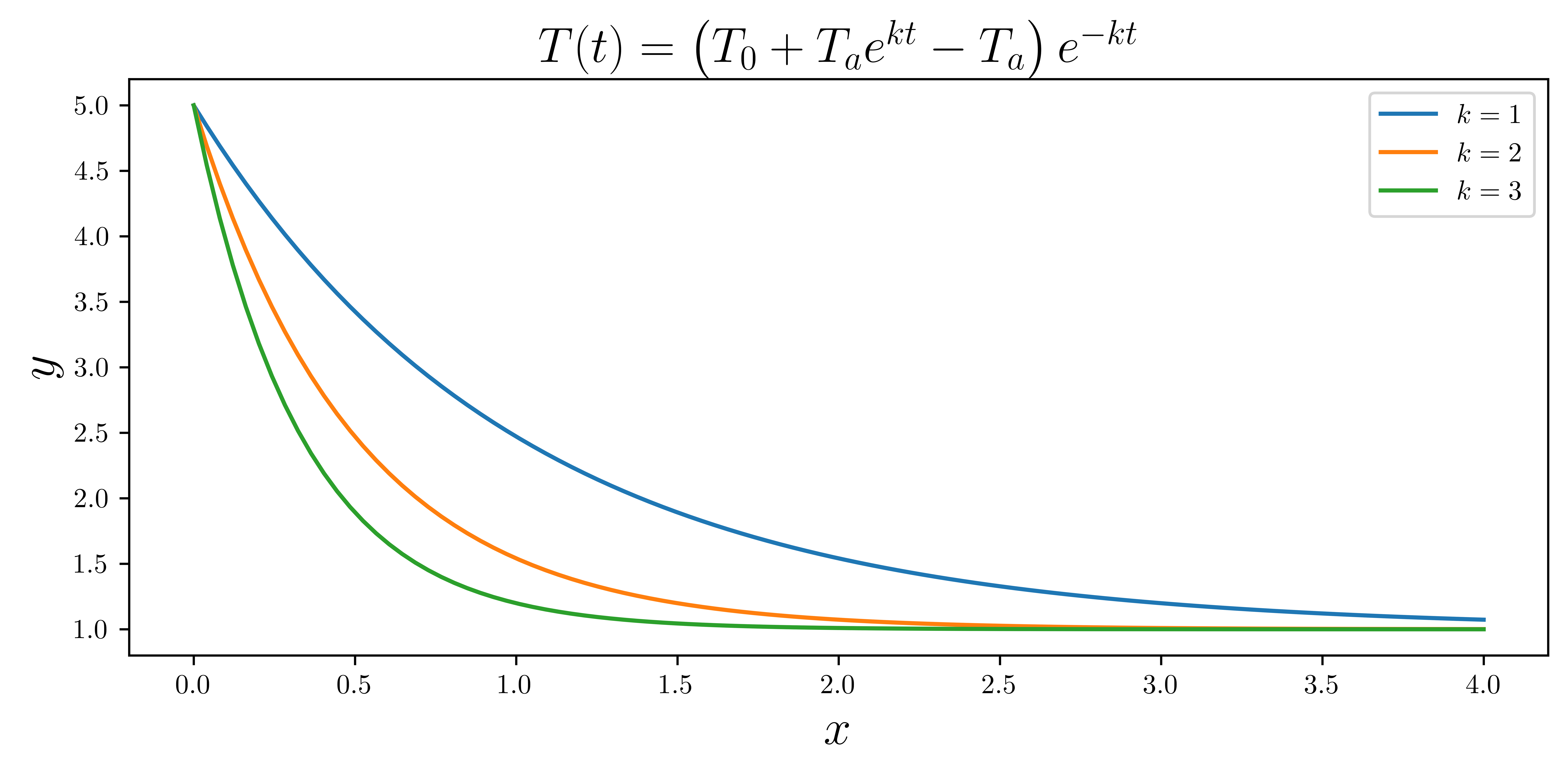
Damped harmonic oscillator¶
t, omega0 = sympy.symbols("t, omega_0", positive=True)
gamma = sympy.symbols("gamma", complex=True)x = sympy.Function("x")ode = x(t).diff(t, 2) + 2 * gamma * omega0 * x(t).diff(t) + omega0**2 * x(t)odeLoading...
ode_sol = sympy.dsolve(ode)ode_solLoading...
ics = {x(0): 1, x(t).diff(t).subs(t, 0): 0}icsLoading...
x_t_sol = apply_ics(ode_sol, ics, t, [omega0, gamma])x_t_solLoading...
x_t_critical = sympy.limit(x_t_sol.rhs, gamma, 1)x_t_criticalLoading...
fig, ax = plt.subplots(figsize=(8, 4))
tt = np.linspace(0, 3, 250)
for g in [0.1, 0.5, 1, 2.0, 5.0]:
if g == 1:
expr = x_t_critical.subs({omega0: 2.0 * sympy.pi})
else:
expr = x_t_sol.rhs.subs({omega0: 2.0 * sympy.pi, gamma: g})
x_t = sympy.lambdify(t, expr, "numpy")
ax.plot(tt, x_t(tt).real, label=r"$\gamma = %.1f$" % g)
ax.set_xlabel(r"$t$", fontsize=18)
ax.set_ylabel(r"$x(t)$", fontsize=18)
ax.set_xlim(0, 3)
ax.legend()
fig.tight_layout()
fig.savefig("ch9-harmonic-oscillator.pdf")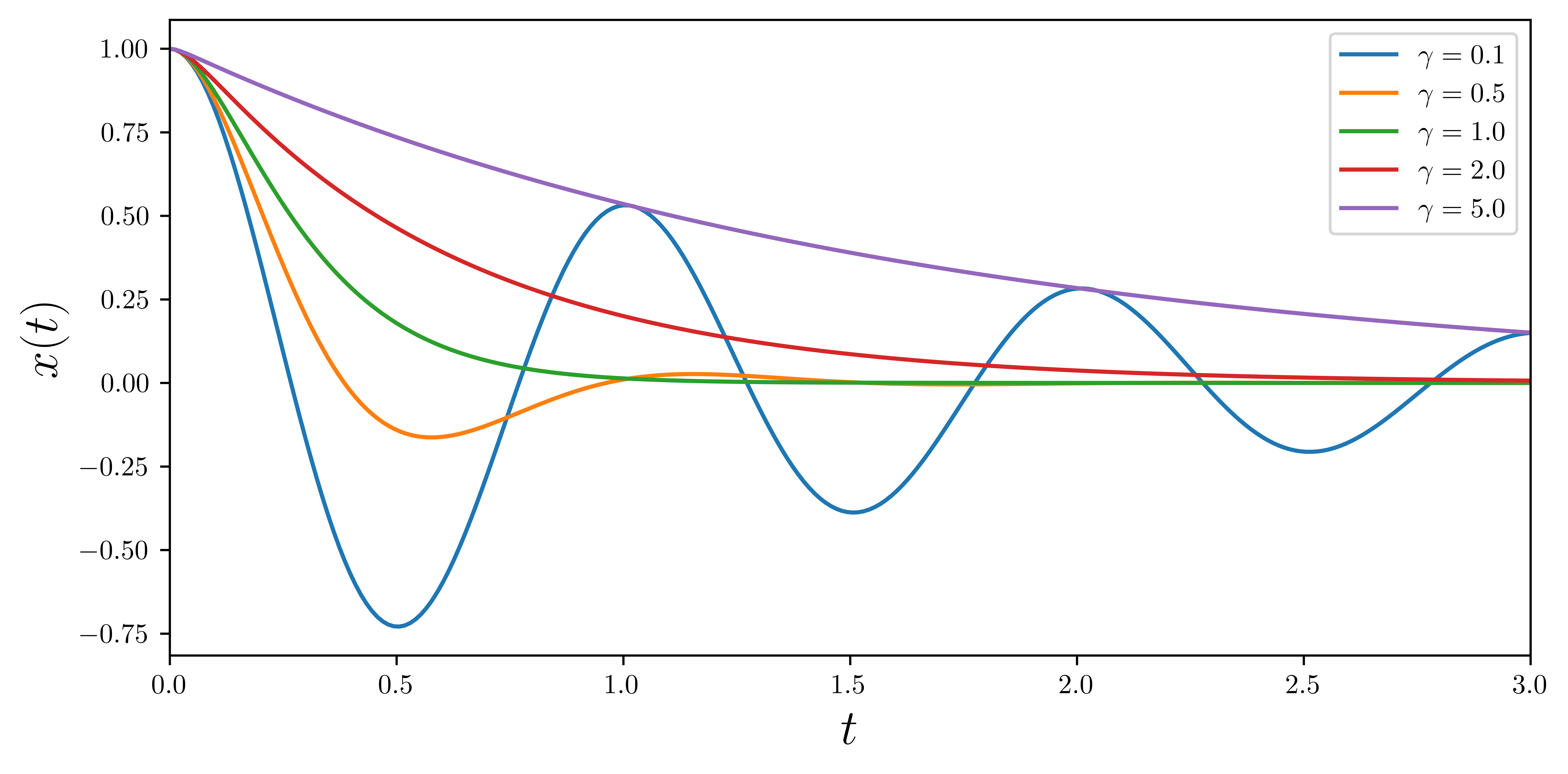
fig, ax = plt.subplots(figsize=(8, 4))
tt = np.linspace(0, 3, 250)
for g in [0.1, 0.5, 1, 2.0, 5.0]:
if g == 1:
x_t = sympy.lambdify(t, x_t_critical.subs({omega0: 2.0 * sympy.pi}), "numpy")
else:
x_t = sympy.lambdify(
t, x_t_sol.rhs.subs({omega0: 2.0 * sympy.pi, gamma: g}), "numpy"
)
ax.plot(tt, x_t(tt).real, label=r"$\gamma = %.1f$" % g)
ax.set_xlabel(r"$t$", fontsize=18)
ax.set_ylabel(r"$x(t)$", fontsize=18)
ax.set_xlim(0, 3)
ax.legend()
fig.tight_layout()
fig.savefig("ch9-harmonic-oscillator.pdf")
# example of equation that sympy cannot solvex = sympy.symbols("x")y = sympy.Function("y")f = y(x) ** 2 + xsympy.Eq(y(x).diff(x, x), f)Loading...
# sympy is unable to solve this differential equation exactly
# sympy.dsolve(y(x).diff(x, x) - f)Direction fields¶
def plot_direction_field(x, y_x, f_xy, x_lim=(-5, 5), y_lim=(-5, 5), ax=None):
f_np = sympy.lambdify((x, y_x), f_xy, "numpy")
x_vec = np.linspace(x_lim[0], x_lim[1], 20)
y_vec = np.linspace(y_lim[0], y_lim[1], 20)
if ax is None:
_, ax = plt.subplots(figsize=(4, 4))
dx = x_vec[1] - x_vec[0]
dy = y_vec[1] - y_vec[0]
for m, xx in enumerate(x_vec):
for n, yy in enumerate(y_vec):
Dy = f_np(xx, yy) * dx
Dx = 0.8 * dx**2 / np.sqrt(dx**2 + Dy**2)
Dy = 0.8 * Dy * dy / np.sqrt(dx**2 + Dy**2)
ax.plot([xx - Dx / 2, xx + Dx / 2], [yy - Dy / 2, yy + Dy / 2], "b", lw=0.5)
ax.axis("tight")
ax.set_title(r"$%s$" % (sympy.latex(sympy.Eq(y(x).diff(x), f_xy))), fontsize=18)
return axx = sympy.symbols("x")y = sympy.Function("y")fig, axes = plt.subplots(1, 3, figsize=(12, 4))
plot_direction_field(x, y(x), y(x) ** 2 + x, ax=axes[0])
plot_direction_field(x, y(x), -x / y(x), ax=axes[1])
plot_direction_field(x, y(x), y(x) ** 2 / x, ax=axes[2])
fig.tight_layout()
fig.savefig("ch9-direction-field.pdf")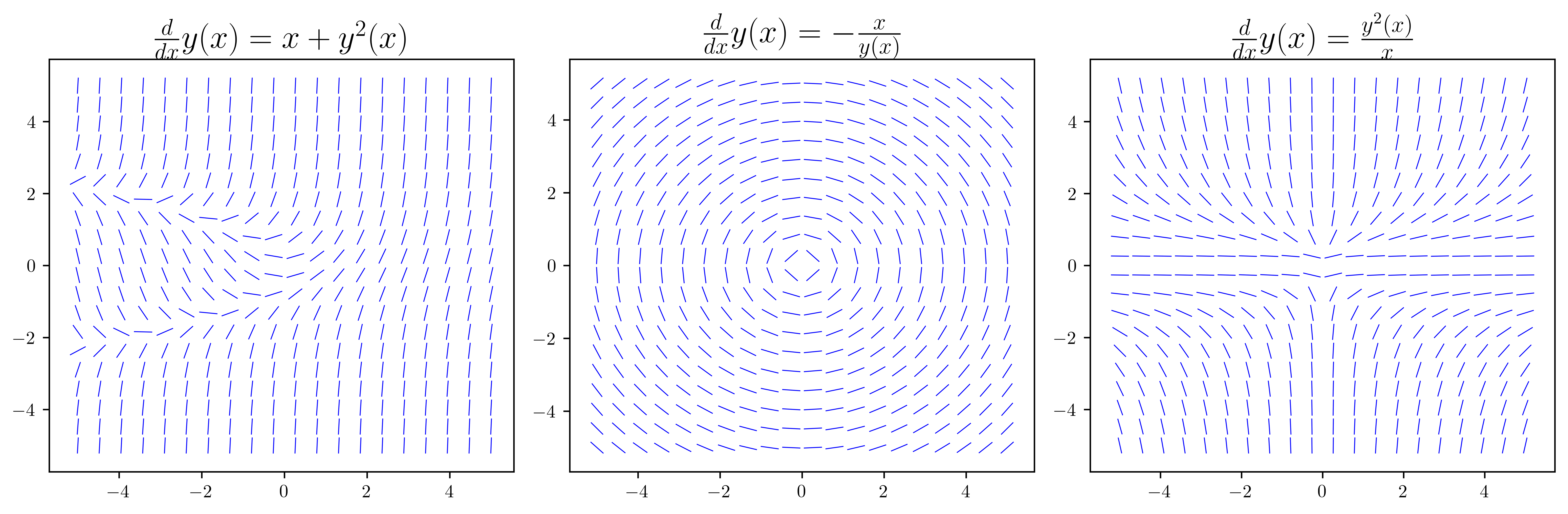
Inexact solutions to ODEs¶
x = sympy.symbols("x")y = sympy.Function("y")f = y(x) ** 2 + x
# f = sympy.cos(y(x)**2) + x
# f = sympy.sqrt(y(x)) * sympy.cos(x**2)
# f = y(x)**2 / xsympy.Eq(y(x).diff(x), f)Loading...
ics = {y(0): 0}sympy.dsolve(
y(x).diff(x) - f, hint="1st_power_series"
) # use hint to get power series approximationLoading...
ode_sol = sympy.dsolve(y(x).diff(x) - f, hint="1st_power_series")ode_solLoading...
ode_sol = apply_ics(ode_sol, {y(0): 0}, x, [])ode_solLoading...
sympy.classify_ode(y(x).diff(x) - f)('1st_rational_riccati', '1st_power_series', 'lie_group')ode_sol = sympy.dsolve(y(x).diff(x) - f, ics=ics, hint="1st_power_series")ode_solLoading...
fig, axes = plt.subplots(1, 1, figsize=(4, 4))
axes = [0, axes]
plot_direction_field(x, y(x), f, ax=axes[1])
x_vec = np.linspace(-1, 1, 100)
# axes[1].plot(x_vec, sympy.lambdify(x, ode_sol.rhs.removeO())(x_vec), 'b', lw=2)
ode_sol_m = ode_sol_p = ode_sol
dx = 0.125
for x0 in np.arange(0, 2.0, dx):
x_vec = np.linspace(x0, x0 + dx, 100)
ics = {y(x0): ode_sol_p.rhs.removeO().subs(x, x0)}
ode_sol_p = sympy.dsolve(y(x).diff(x) - f, ics=ics, n=6, hint="1st_power_series")
axes[1].plot(x_vec, sympy.lambdify(x, ode_sol_p.rhs.removeO())(x_vec), "r", lw=2)
for x0 in np.arange(0, -5, -dx):
x_vec = np.linspace(x0, x0 - dx, 100)
ics = {y(x0): ode_sol_m.rhs.removeO().subs(x, x0)}
ode_sol_m = sympy.dsolve(y(x).diff(x) - f, ics=ics, n=6, hint="1st_power_series")
axes[1].plot(x_vec, sympy.lambdify(x, ode_sol_m.rhs.removeO())(x_vec), "b", lw=2)
axes[1].set_ylim(-4.75, 4.75)
axes[1].set_ylim(-4.75, 4.75)
fig.tight_layout()
fig.savefig("ch9-direction-field-and-approx-sol.pdf")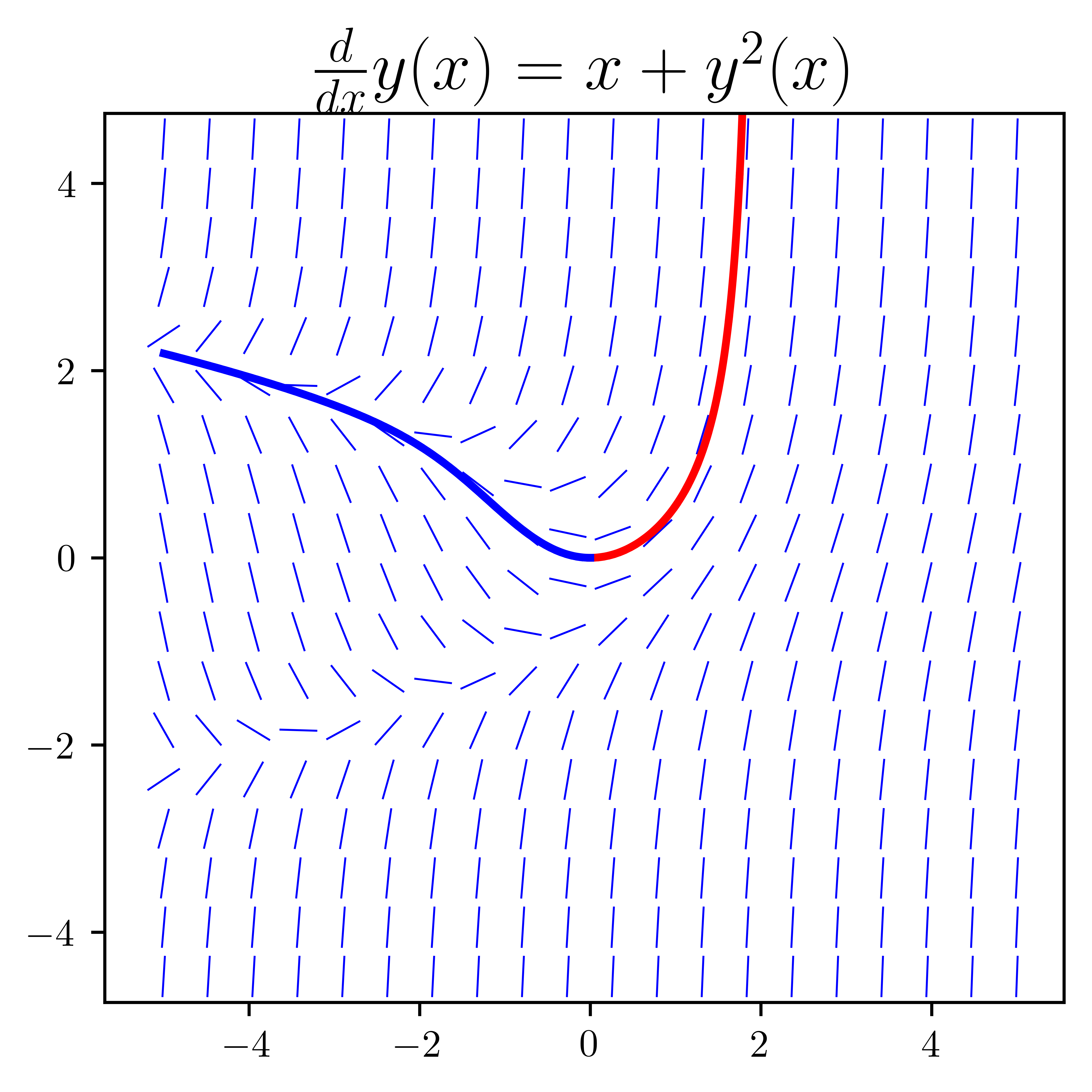
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
plot_direction_field(x, y(x), f, ax=axes[0])
x_vec = np.linspace(-3, 3, 100)
axes[0].plot(x_vec, sympy.lambdify(x, ode_sol.rhs.removeO())(x_vec), "b", lw=2)
axes[0].set_ylim(-5, 5)
plot_direction_field(x, y(x), f, ax=axes[1])
x_vec = np.linspace(-1, 1, 100)
axes[1].plot(x_vec, sympy.lambdify(x, ode_sol.rhs.removeO())(x_vec), "b", lw=2)
ode_sol_m = ode_sol_p = ode_sol
dx = 0.125
for x0 in np.arange(1, 2.0, dx):
x_vec = np.linspace(x0, x0 + dx, 100)
ics = {y(x0): ode_sol_p.rhs.removeO().subs(x, x0)}
ode_sol_p = sympy.dsolve(y(x).diff(x) - f, ics=ics, n=6, hint="1st_power_series")
axes[1].plot(x_vec, sympy.lambdify(x, ode_sol_p.rhs.removeO())(x_vec), "r", lw=2)
for x0 in np.arange(-1, -5, -dx):
x_vec = np.linspace(x0, x0 - dx, 100)
ics = {y(x0): ode_sol_m.rhs.removeO().subs(x, x0)}
ode_sol_m = sympy.dsolve(y(x).diff(x) - f, ics=ics, n=6, hint="1st_power_series")
axes[1].plot(x_vec, sympy.lambdify(x, ode_sol_m.rhs.removeO())(x_vec), "r", lw=2)
fig.tight_layout()
fig.savefig("ch9-direction-field-and-approx-sol.pdf")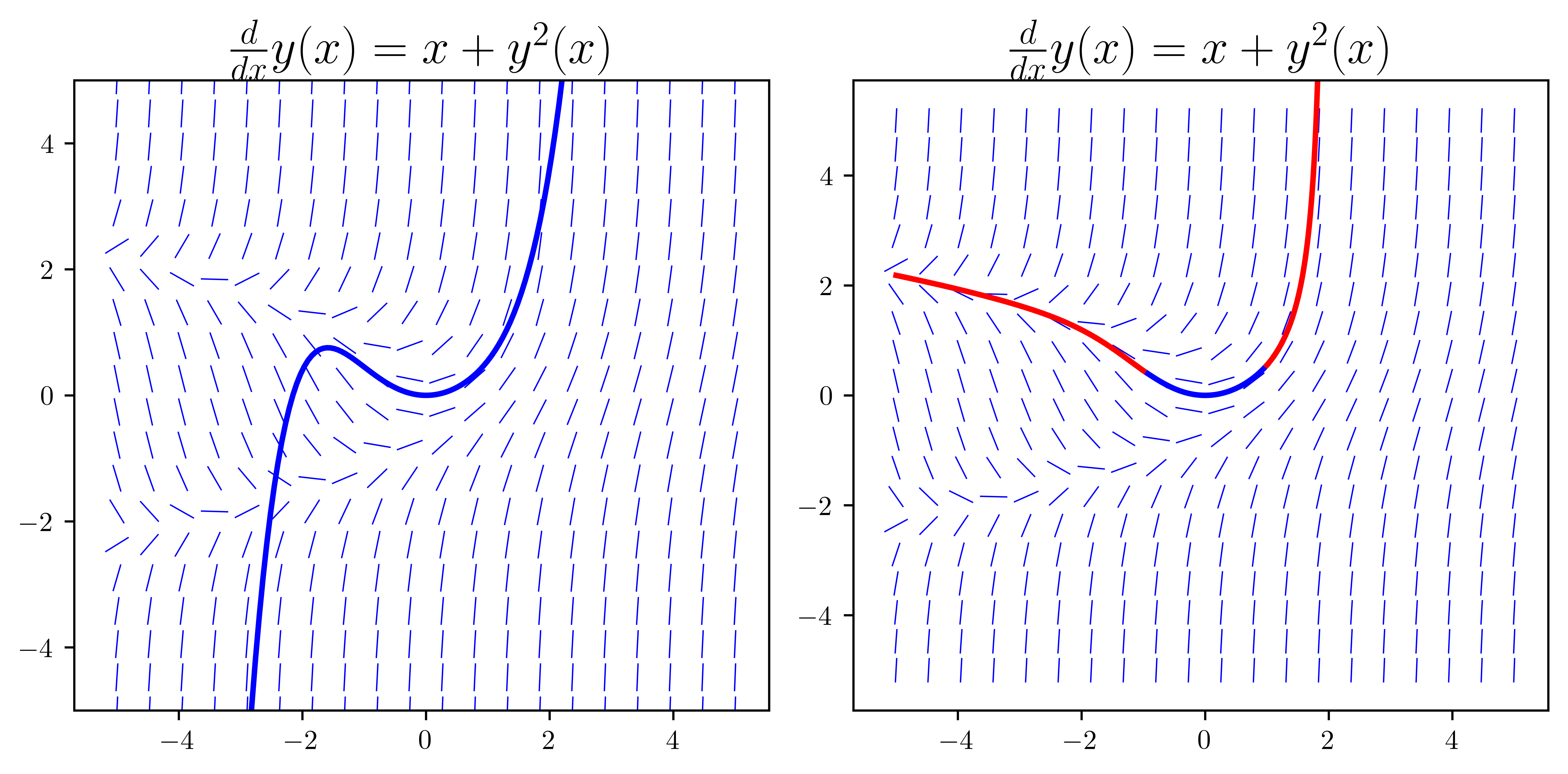
Laplace transformation method¶
t = sympy.symbols("t", positive=True)s, Y = sympy.symbols("s, Y", real=True)y = sympy.Function("y")ode = y(t).diff(t, 2) + 2 * y(t).diff(t) + 10 * y(t) - 2 * sympy.sin(3 * t)odeLoading...
L_y = sympy.laplace_transform(y(t), t, s)[0]L_yLoading...
# L_ode = sympy.laplace_transform(ode, t, s, noconds=True) # previously like this, not working now
L_ode = sympy.laplace_transform(ode, t, s)[0] # instead use thisL_odeLoading...
def laplace_transform_derivatives(e):
"""
Evaluate the laplace transforms of derivatives of functions
"""
if isinstance(e, sympy.LaplaceTransform):
if isinstance(e.args[0], sympy.Derivative):
d, t, s = e.args
n = d.args[1][1]
return (s**n) * sympy.LaplaceTransform(d.args[0], t, s) - sum(
[
s ** (n - i) * sympy.diff(d.args[0], t, i - 1).subs(t, 0)
for i in range(1, n + 1)
]
)
if isinstance(e, (sympy.Add, sympy.Mul)):
t = type(e)
return t(*[laplace_transform_derivatives(arg) for arg in e.args])
return eL_odeLoading...
L_ode_2 = laplace_transform_derivatives(L_ode)L_ode_2Loading...
L_ode_3 = L_ode_2.subs(L_y, Y)L_ode_3Loading...
ics = {y(0): 1, y(t).diff(t).subs(t, 0): 0}icsLoading...
L_ode_4 = L_ode_3.subs(ics)L_ode_4Loading...
Y_sol = sympy.solve(L_ode_4, Y)Y_solLoading...
sympy.apart(Y_sol[0])Loading...
y_sol = sympy.inverse_laplace_transform(Y_sol[0], s, t)sympy.simplify(y_sol)Loading...
sympy.simplify(y_sol).evalf()Loading...
y_t = sympy.lambdify(t, y_sol.evalf(), "numpy")fig, ax = plt.subplots(figsize=(8, 4))
tt = np.linspace(0, 10, 500)
ax.plot(tt, y_t(tt).real)
ax.set_xlabel(r"$t$", fontsize=18)
ax.set_ylabel(r"$y(t)$", fontsize=18)
fig.tight_layout()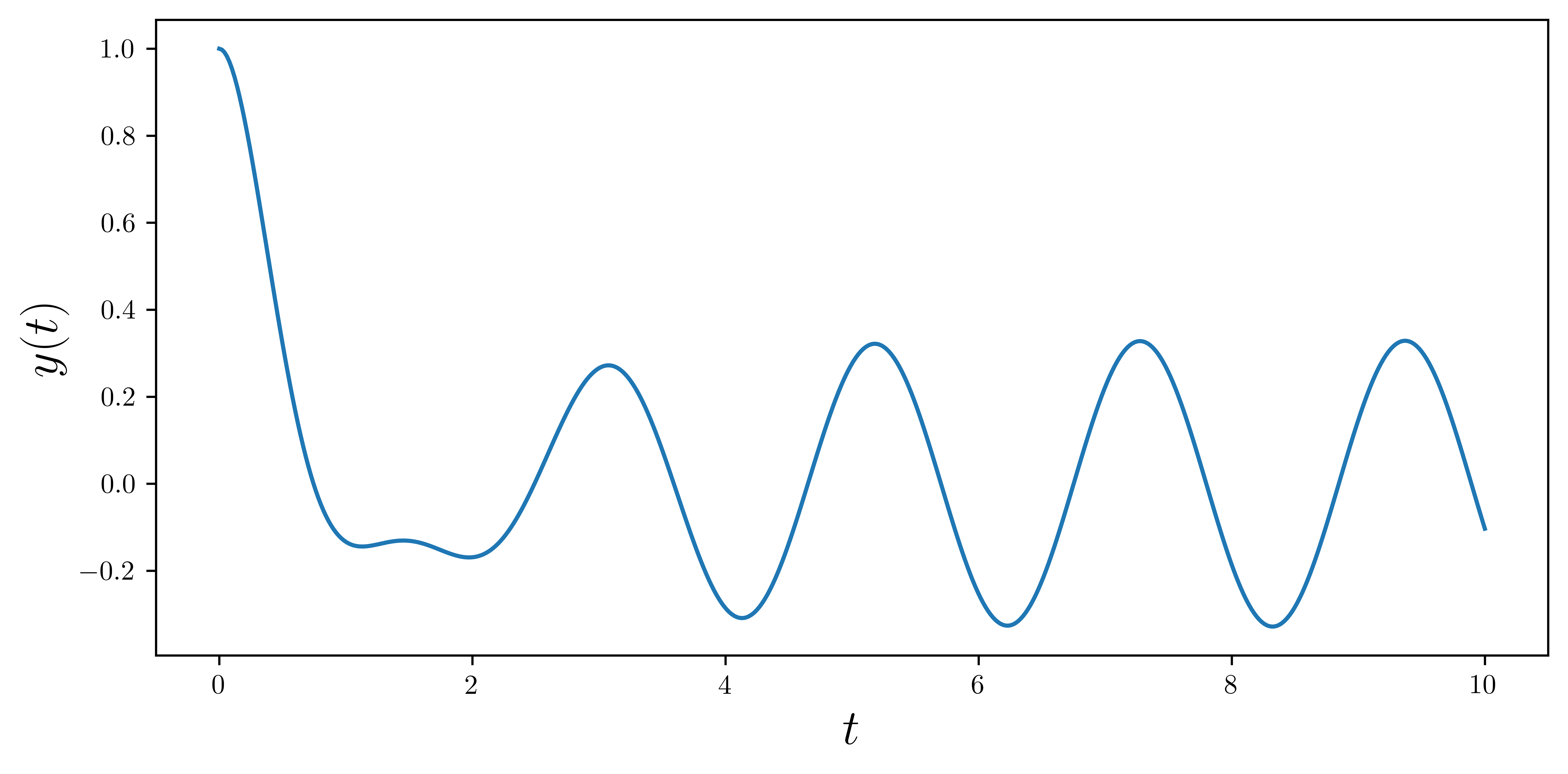
Numerical integration of ODEs using SciPy¶
x = sympy.symbols("x")y = sympy.Function("y")f = y(x) ** 2 + xf_np = sympy.lambdify((y(x), x), f, "math")y0 = 0xp = np.linspace(0, 1.9, 100)xp.shapeLoading...
yp = integrate.odeint(f_np, y0, xp)xm = np.linspace(0, -5, 100)ym = integrate.odeint(f_np, y0, xm)fig, ax = plt.subplots(1, 1, figsize=(4, 4))
plot_direction_field(x, y(x), f, ax=ax)
ax.plot(xm, ym, "b", lw=2)
ax.plot(xp, yp, "r", lw=2)
fig.savefig("ch9-odeint-single-eq-example.pdf")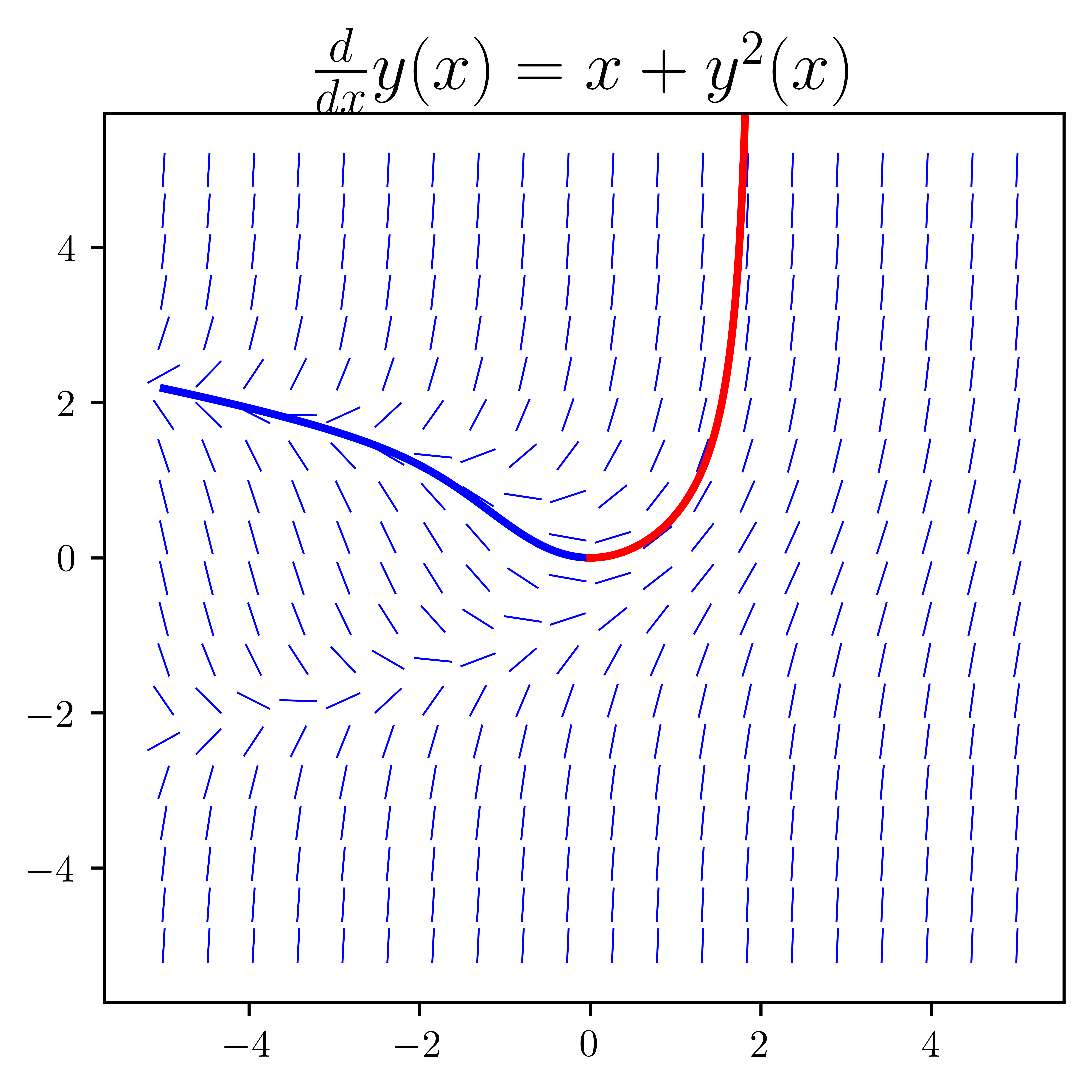
a, b, c, d = 0.4, 0.002, 0.001, 0.7def f(xy, t):
x, y = xy
return [a * x - b * x * y, c * x * y - d * y]xy0 = [600, 400]t = np.linspace(0, 50, 250)xy_t = integrate.odeint(f, xy0, t)xy_t.shapeLoading...
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
axes[0].plot(t, xy_t[:, 0], "r", label="Prey")
axes[0].plot(t, xy_t[:, 1], "b", label="Predator")
axes[0].set_xlabel("Time")
axes[0].set_ylabel("Number of animals")
axes[0].legend(loc=2, facecolor="white", framealpha=1)
axes[1].plot(xy_t[:, 0], xy_t[:, 1], "k")
axes[1].set_xlabel("Number of prey")
axes[1].set_ylabel("Number of predators")
fig.tight_layout()
fig.savefig("ch9-lokta-volterra.pdf")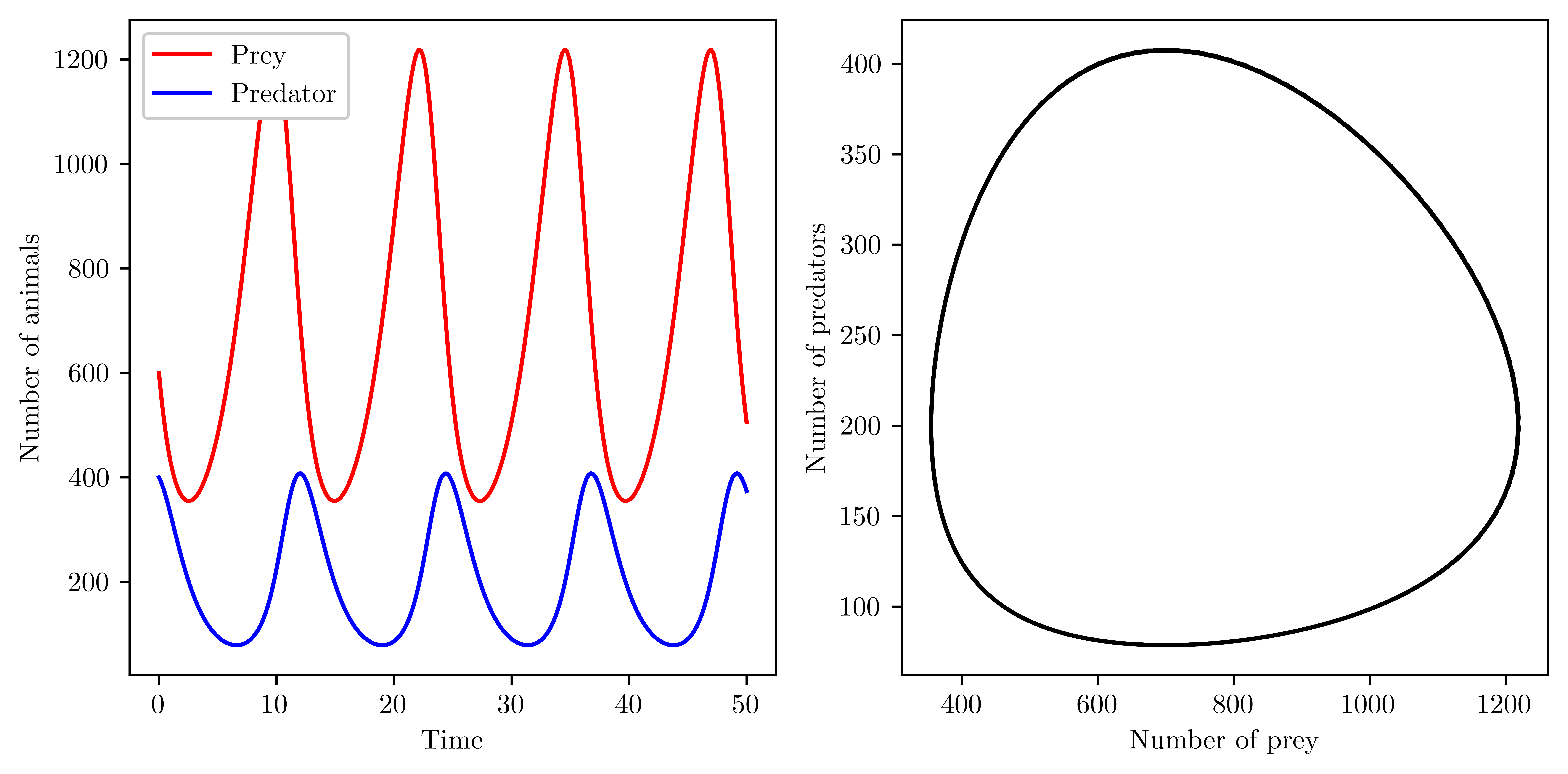
Lorenz equations¶
def f(xyz, t, rho, sigma, beta):
x, y, z = xyz
return [sigma * (y - x), x * (rho - z) - y, x * y - beta * z]rho = 28
sigma = 8
beta = 8 / 3.0t = np.linspace(0, 25, 10000)xyz0 = [1.0, 1.0, 1.0]xyz1 = integrate.odeint(f, xyz0, t, args=(rho, sigma, beta))xyz2 = integrate.odeint(f, xyz0, t, args=(rho, sigma, 0.6 * beta))xyz3 = integrate.odeint(f, xyz0, t, args=(rho, 2 * sigma, 0.6 * beta))xyz3.shapeLoading...
from mpl_toolkits.mplot3d.axes3d import Axes3Dfig, (ax1, ax2, ax3) = plt.subplots(
1, 3, figsize=(12, 3.5), subplot_kw={"projection": "3d"}
)
for ax, xyz, c, p in [
(ax1, xyz1, "r", (rho, sigma, beta)),
(ax2, xyz2, "b", (rho, sigma, 0.6 * beta)),
(ax3, xyz3, "g", (rho, 2 * sigma, 0.6 * beta)),
]:
ax.plot(xyz[:, 0], xyz[:, 1], xyz[:, 2], c, alpha=0.5)
ax.set_xlabel("$x$", fontsize=16)
ax.set_ylabel("$y$", fontsize=16)
ax.set_zlabel("$z$", fontsize=16)
ax.set_xticks([-15, 0, 15])
ax.set_yticks([-20, 0, 20])
ax.set_zticks([0, 20, 40])
ax.set_title(r"$\rho=%.1f, \sigma=%.1f, \beta=%.1f$" % (p[0], p[1], p[2]))
fig.tight_layout()
fig.savefig("ch9-lorenz-equations.pdf")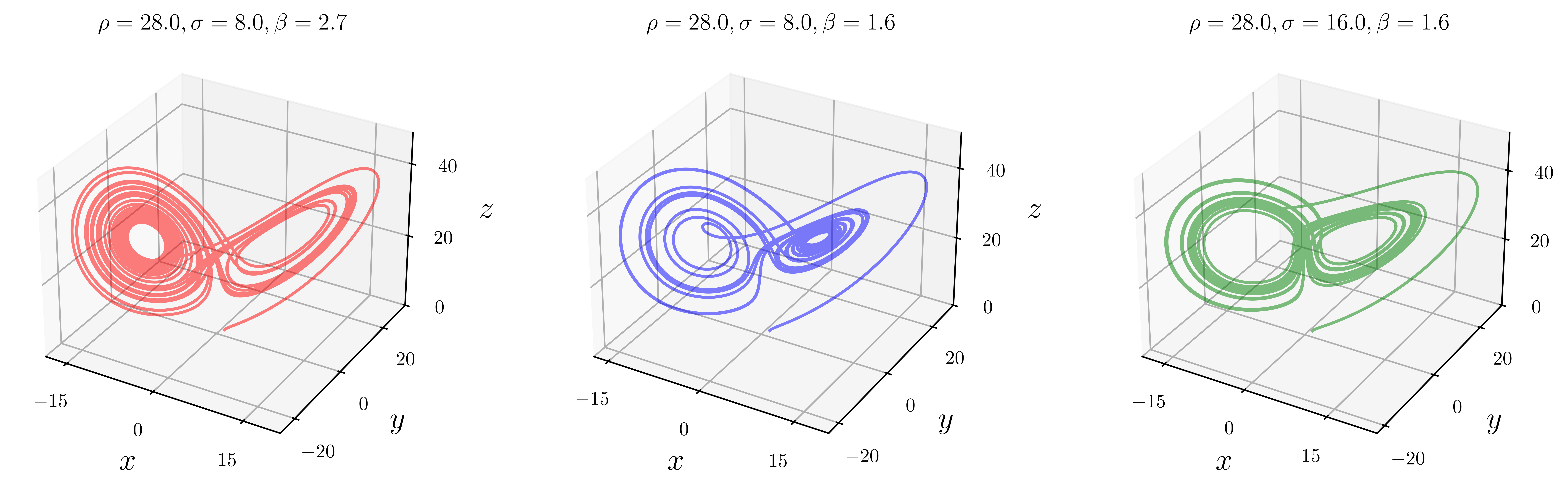
Coupled damped springs¶
def f(t, y, args):
m1, k1, g1, m2, k2, g2 = args
return [
y[1],
-k1 / m1 * y[0] + k2 / m1 * (y[2] - y[0]) - g1 / m1 * y[1],
y[3],
-k2 / m2 * (y[2] - y[0]) - g2 / m2 * y[3],
]m1, k1, g1 = 1.0, 10.0, 0.5m2, k2, g2 = 2.0, 40.0, 0.25args = (m1, k1, g1, m2, k2, g2)y0 = [1.0, 0, 0.5, 0]t = np.linspace(0, 20, 1000)r = integrate.ode(f)r.set_integrator("lsoda")<scipy.integrate._ode.ode at 0x73d31c3becf0>r.set_initial_value(y0, t[0])<scipy.integrate._ode.ode at 0x73d31c3becf0>r.set_f_params(args)<scipy.integrate._ode.ode at 0x73d31c3becf0>dt = t[1] - t[0]
y = np.zeros((len(t), len(y0)))
idx = 0
while r.successful() and r.t < t[-1]:
y[idx, :] = r.y
r.integrate(r.t + dt)
idx += 1fig = plt.figure(figsize=(10, 4))
ax1 = plt.subplot2grid((2, 5), (0, 0), colspan=3)
ax2 = plt.subplot2grid((2, 5), (1, 0), colspan=3)
ax3 = plt.subplot2grid((2, 5), (0, 3), colspan=2, rowspan=2)
ax1.plot(t, y[:, 0], "r")
ax1.set_ylabel("$x_1$", fontsize=18)
ax1.set_yticks([-1, -0.5, 0, 0.5, 1])
ax2.plot(t, y[:, 2], "b")
ax2.set_xlabel("$t$", fontsize=18)
ax2.set_ylabel("$x_2$", fontsize=18)
ax2.set_yticks([-1, -0.5, 0, 0.5, 1])
ax3.plot(y[:, 0], y[:, 2], "k")
ax3.set_xlabel("$x_1$", fontsize=18)
ax3.set_ylabel("$x_2$", fontsize=18)
ax3.set_xticks([-1, -0.5, 0, 0.5, 1])
ax3.set_yticks([-1, -0.5, 0, 0.5, 1])
fig.tight_layout()
fig.savefig("ch9-coupled-damped-springs.pdf")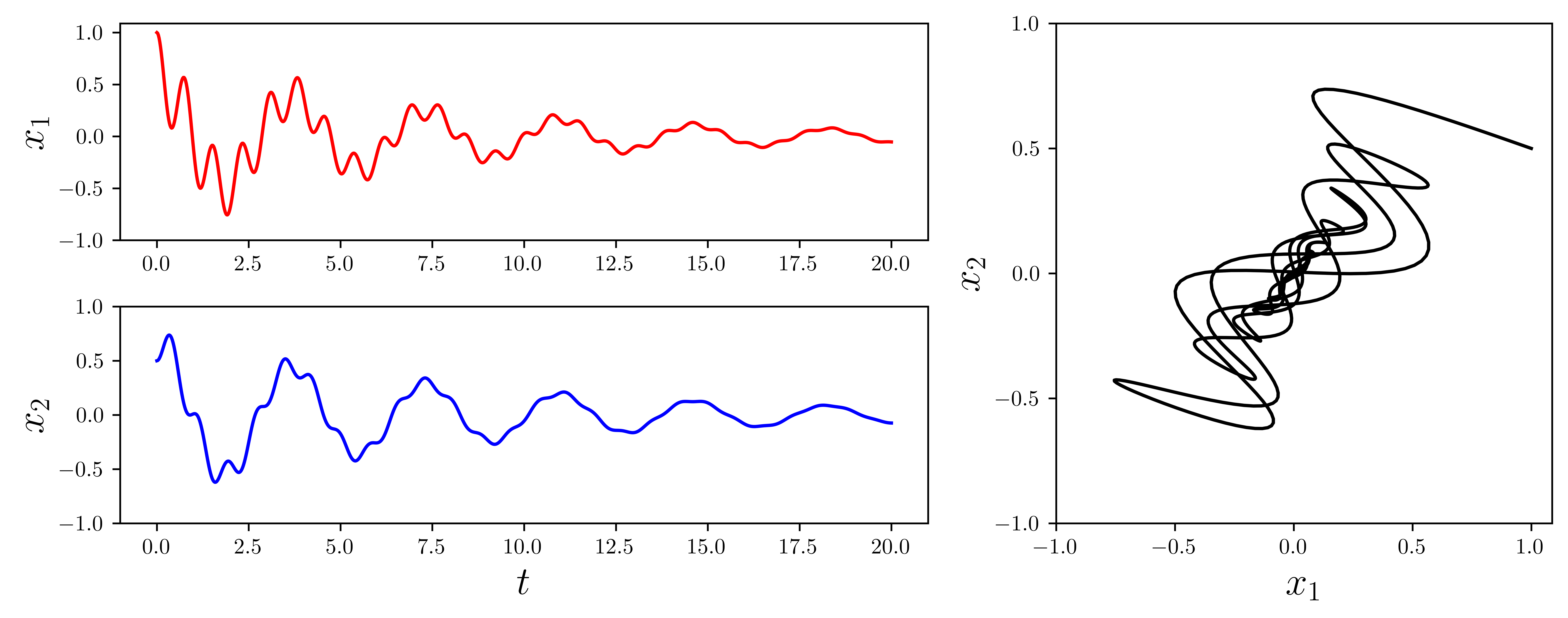
Same calculation as above, but with specifying the Jacobian as well:¶
def jac(t, y, args):
m1, k1, g1, m2, k2, g2 = args
return [
[0, 1, 0, 0],
[-k1 / m1 - k2 / m1, -g1 / m1, k2 / m1, 0],
[0, 0, 0, 1],
[k2 / m2, 0, -k2 / m2, -g2 / m2],
]r = (
integrate.ode(f, jac)
.set_f_params(args)
.set_jac_params(args)
.set_initial_value(y0, t[0])
)dt = t[1] - t[0]
y = np.zeros((len(t), len(y0)))
idx = 0
while r.successful() and r.t < t[-1]:
y[idx, :] = r.y
r.integrate(r.t + dt)
idx += 1fig = plt.figure(figsize=(10, 4))
ax1 = plt.subplot2grid((2, 5), (0, 0), colspan=3)
ax2 = plt.subplot2grid((2, 5), (1, 0), colspan=3)
ax3 = plt.subplot2grid((2, 5), (0, 3), colspan=2, rowspan=2)
ax1.plot(t, y[:, 0], "r")
ax1.set_ylabel("$x_1$", fontsize=18)
ax1.set_yticks([-1, -0.5, 0, 0.5, 1])
ax2.plot(t, y[:, 2], "b")
ax2.set_xlabel("$t$", fontsize=18)
ax2.set_ylabel("$x_2$", fontsize=18)
ax2.set_yticks([-1, -0.5, 0, 0.5, 1])
ax3.plot(y[:, 0], y[:, 2], "k")
ax3.set_xlabel("$x_1$", fontsize=18)
ax3.set_ylabel("$x_2$", fontsize=18)
ax3.set_xticks([-1, -0.5, 0, 0.5, 1])
ax3.set_yticks([-1, -0.5, 0, 0.5, 1])
fig.tight_layout()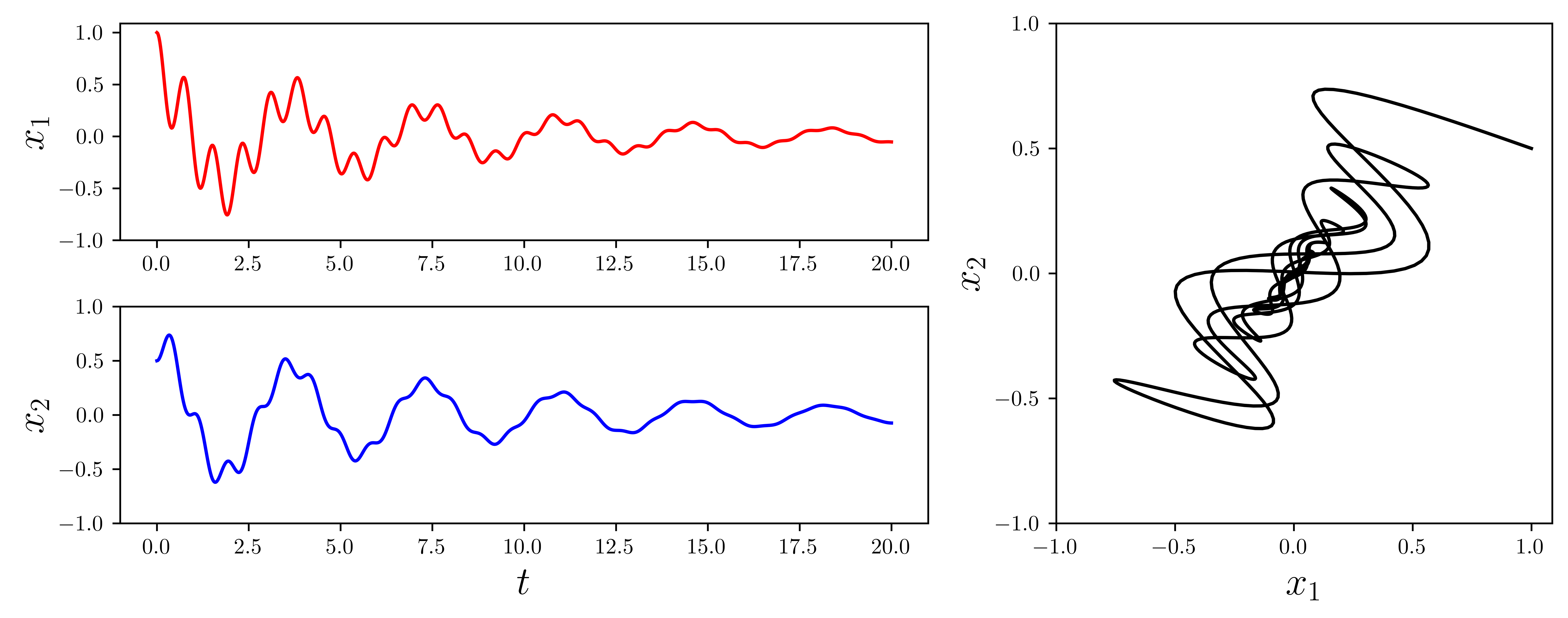
Same calculation, but using SymPy to setup the problem for SciPy¶
L1 = L2 = 0
t = sympy.symbols("t")
m1, k1, b1 = sympy.symbols("m_1, k_1, b_1")
m2, k2, b2 = sympy.symbols("m_2, k_2, b_2")x1 = sympy.Function("x_1")
x2 = sympy.Function("x_2")ode1 = sympy.Eq(
m1
* x1(t).diff(
t,
t,
)
+ b1 * x1(t).diff(t)
+ k1 * (x1(t) - L1)
- k2 * (x2(t) - x1(t) - L2),
0,
)ode2 = sympy.Eq(
m2
* x2(t).diff(
t,
t,
)
+ b2 * x2(t).diff(t)
+ k2 * (x2(t) - x1(t) - L2),
0,
)params = {m1: 1.0, k1: 10.0, b1: 0.5, m2: 2.0, k2: 40.0, b2: 0.25}argsLoading...
y1 = sympy.Function("y_1")
y2 = sympy.Function("y_2")
y3 = sympy.Function("y_3")
y4 = sympy.Function("y_4")varchange = {
x1(t).diff(t, t): y2(t).diff(t),
x1(t): y1(t),
x2(t).diff(t, t): y4(t).diff(t),
x2(t): y3(t),
}(ode1.subs(varchange).lhs, ode2.subs(varchange).lhs)Loading...
ode3 = y1(t).diff(t) - y2(t)ode4 = y3(t).diff(t) - y4(t)vcsol = sympy.solve(
(ode1.subs(varchange), ode2.subs(varchange), ode3, ode4),
(y1(t).diff(t), y2(t).diff(t), y3(t).diff(t), y4(t).diff(t)),
)vcsolLoading...
ode_rhs = sympy.Matrix(
[y1(t).diff(t), y2(t).diff(t), y3(t).diff(t), y4(t).diff(t)]
).subs(vcsol)y = sympy.Matrix([y1(t), y2(t), y3(t), y4(t)])sympy.Eq(y.diff(t), ode_rhs)Loading...
ode_rhs.subs(params)Loading...
_f_np = sympy.lambdify((t, y), ode_rhs.subs(params), "numpy")
f_np = lambda _x, _y, *args: _f_np(_x, _y).flatten()y0 = [1.0, 0, 0.5, 0]
tt = np.linspace(0, 20, 1000)
r = integrate.ode(f_np)
r.set_integrator("lsoda")
r.set_initial_value(y0, tt[0])
dt = tt[1] - tt[0]
yy = np.zeros((len(tt), len(y0)))
idx = 0
while r.successful() and r.t < tt[-1]:
yy[idx, :] = r.y
r.integrate(r.t + dt)
idx += 1fig = plt.figure(figsize=(10, 4))
ax1 = plt.subplot2grid((2, 5), (0, 0), colspan=3)
ax2 = plt.subplot2grid((2, 5), (1, 0), colspan=3)
ax3 = plt.subplot2grid((2, 5), (0, 3), colspan=2, rowspan=2)
ax1.plot(tt, yy[:, 0], "r")
ax1.set_ylabel("$x_1$", fontsize=18)
ax1.set_yticks([-1, -0.5, 0, 0.5, 1])
ax2.plot(tt, yy[:, 2], "b")
ax2.set_xlabel("$t$", fontsize=18)
ax2.set_ylabel("$x_2$", fontsize=18)
ax2.set_yticks([-1, -0.5, 0, 0.5, 1])
ax3.plot(yy[:, 0], yy[:, 2], "k")
ax3.set_xlabel("$x_1$", fontsize=18)
ax3.set_ylabel("$x_2$", fontsize=18)
ax3.set_xticks([-1, -0.5, 0, 0.5, 1])
ax3.set_yticks([-1, -0.5, 0, 0.5, 1])
fig.tight_layout()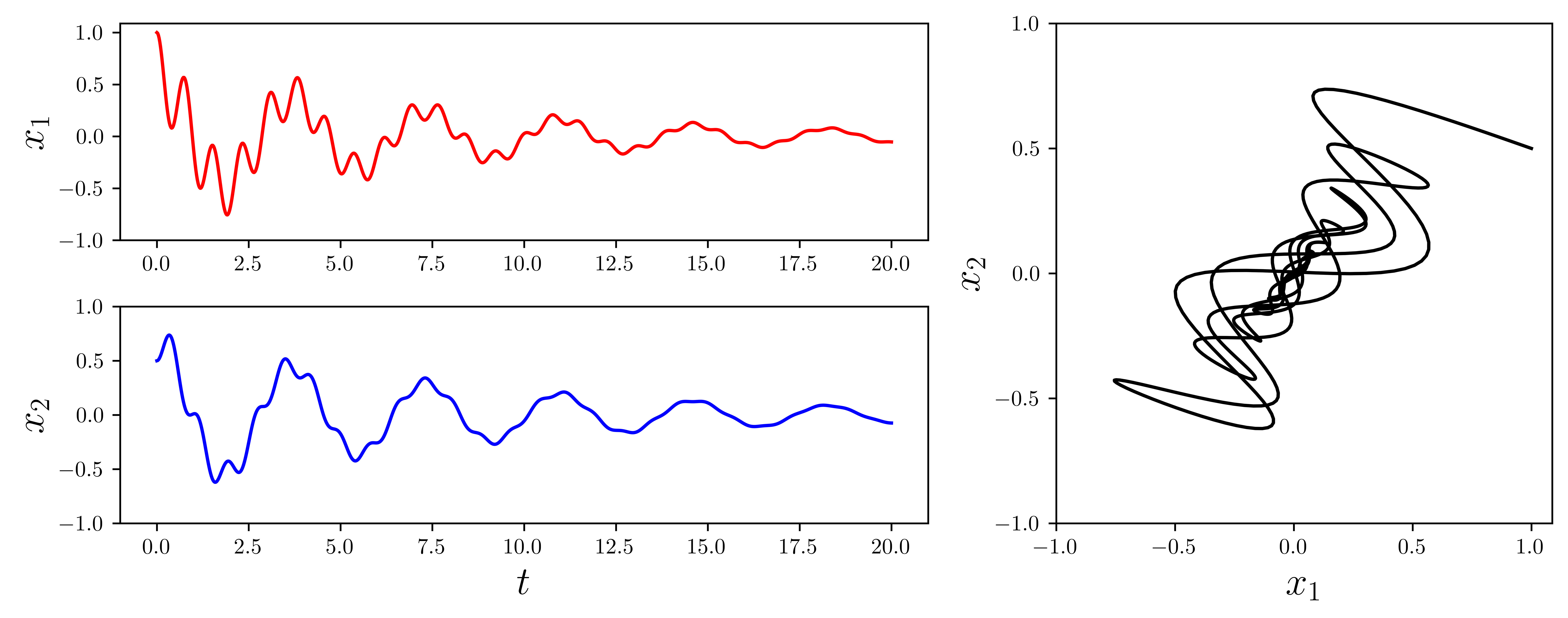
Doube pendulum¶
t, g, m1, l1, m2, l2 = sympy.symbols("t, g, m_1, l_1, m_2, l_2")theta1, theta2 = sympy.symbols("theta_1, theta_2", cls=sympy.Function)ode1 = sympy.Eq(
(m1 + m2) * l1 * theta1(t).diff(t, t)
+ m2 * l2 * theta2(t).diff(t, t) * sympy.cos(theta1(t) - theta2(t))
+ m2 * l2 * theta2(t).diff(t) ** 2 * sympy.sin(theta1(t) - theta2(t))
+ g * (m1 + m2) * sympy.sin(theta1(t)),
0,
)
ode1Loading...
ode2 = sympy.Eq(
m2 * l2 * theta2(t).diff(t, t)
+ m2 * l1 * theta1(t).diff(t, t) * sympy.cos(theta1(t) - theta2(t))
- m2 * l1 * theta1(t).diff(t) ** 2 * sympy.sin(theta1(t) - theta2(t))
+ m2 * g * sympy.sin(theta2(t)),
0,
)
ode2Loading...
# this is fruitless, sympy cannot solve these ODEs
try:
sympy.dsolve(ode1, ode2)
except Exception as e:
print(e)cannot determine truth value of Relational: Eq(g*m_2*sin(theta_2(t)) - l_1*m_2*sin(theta_1(t) - theta_2(t))*Derivative(theta_1(t), t)**2 + l_1*m_2*cos(theta_1(t) - theta_2(t))*Derivative(theta_1(t), (t, 2)) + l_2*m_2*Derivative(theta_2(t), (t, 2)), 0)
y1, y2, y3, y4 = sympy.symbols("y_1, y_2, y_3, y_4", cls=sympy.Function)varchange = {
theta1(t).diff(t, t): y2(t).diff(t),
theta1(t): y1(t),
theta2(t).diff(t, t): y4(t).diff(t),
theta2(t): y3(t),
}ode1_vc = ode1.subs(varchange)ode2_vc = ode2.subs(varchange)ode3 = y1(t).diff(t) - y2(t)ode4 = y3(t).diff(t) - y4(t)ode3 = sympy.Eq(y1(t).diff(t), y2(t))
ode4 = sympy.Eq(y3(t).diff(t), y4(t))y = sympy.Matrix([y1(t), y2(t), y3(t), y4(t)])vcsol = sympy.solve((ode1_vc, ode2_vc, ode3, ode4), y.diff(t), dict=True)f = y.diff(t).subs(vcsol[0])sympy.Eq(y.diff(t), f)Loading...
params = {m1: 5.0, l1: 2.0, m2: 1.0, l2: 1.0, g: 10.0}_f_np = sympy.lambdify((t, y), f.subs(params), "numpy")
f_np = lambda _t, _y, *args: _f_np(_t, _y).flatten()jac = sympy.Matrix([[fj.diff(yi) for yi in y] for fj in f])_jac_np = sympy.lambdify((t, y), jac.subs(params), "numpy")
jac_np = lambda _t, _y, *args: _jac_np(_t, _y)y0 = [2.0, 0, 0.0, 0]tt = np.linspace(0, 20, 1000)%%time
r = integrate.ode(f_np, jac_np).set_initial_value(y0, tt[0])
dt = tt[1] - tt[0]
yy = np.zeros((len(tt), len(y0)))
idx = 0
while r.successful() and r.t < tt[-1]:
yy[idx, :] = r.y
r.integrate(r.t + dt)
idx += 1CPU times: user 34 ms, sys: 6.67 ms, total: 40.6 ms
Wall time: 30.7 ms
fig = plt.figure(figsize=(10, 4))
ax1 = plt.subplot2grid((2, 5), (0, 0), colspan=3)
ax2 = plt.subplot2grid((2, 5), (1, 0), colspan=3)
ax3 = plt.subplot2grid((2, 5), (0, 3), colspan=2, rowspan=2)
ax1.plot(tt, yy[:, 0], "r")
ax1.set_ylabel(r"$\theta_1$", fontsize=18)
ax2.plot(tt, yy[:, 2], "b")
ax2.set_xlabel("$t$", fontsize=18)
ax2.set_ylabel(r"$\theta_2$", fontsize=18)
ax3.plot(yy[:, 0], yy[:, 2], "k")
ax3.set_xlabel(r"$\theta_1$", fontsize=18)
ax3.set_ylabel(r"$\theta_2$", fontsize=18)
fig.tight_layout()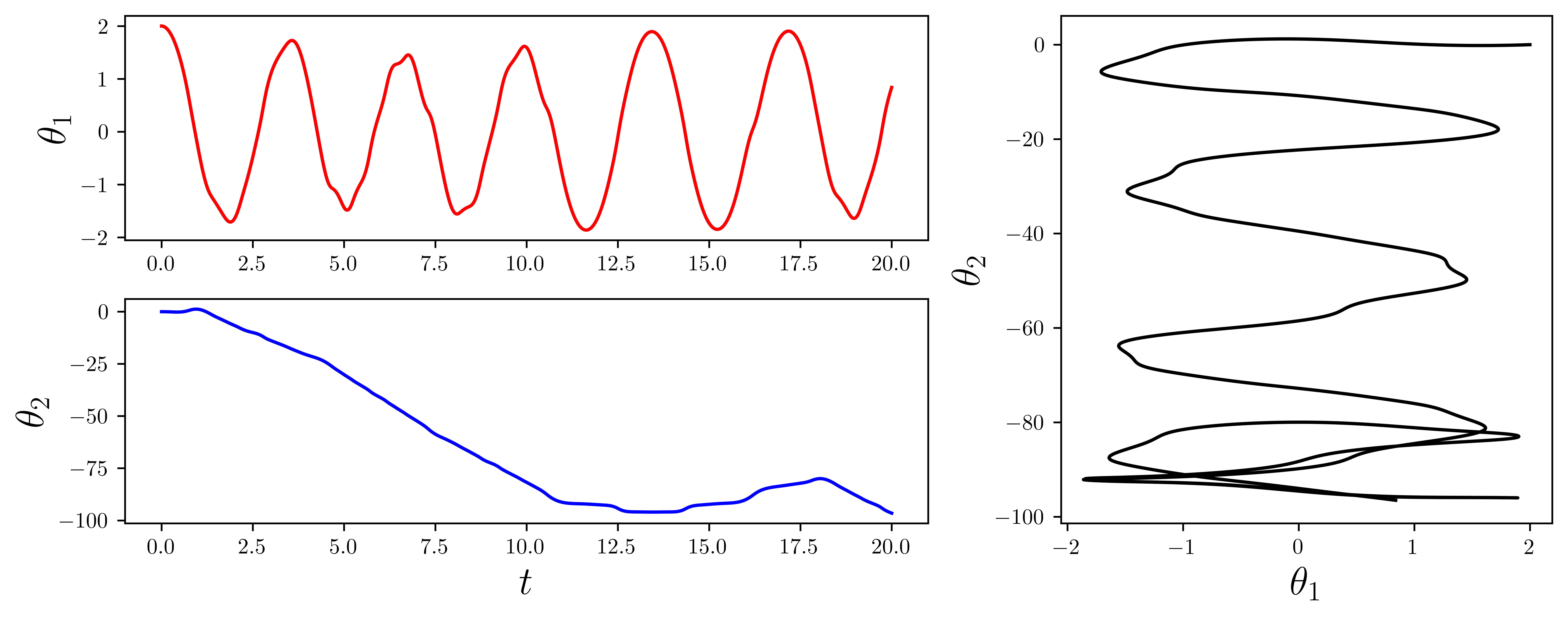
theta1_np, theta2_np = yy[:, 0], yy[:, 2]x1 = params[l1] * np.sin(theta1_np)
y1 = -params[l1] * np.cos(theta1_np)
x2 = x1 + params[l2] * np.sin(theta2_np)
y2 = y1 - params[l2] * np.cos(theta2_np)fig = plt.figure(figsize=(10, 4))
ax1 = plt.subplot2grid((2, 5), (0, 0), colspan=3)
ax2 = plt.subplot2grid((2, 5), (1, 0), colspan=3)
ax3 = plt.subplot2grid((2, 5), (0, 3), colspan=2, rowspan=2)
ax1.plot(tt, x1, "r")
ax1.plot(tt, y1, "b")
ax1.set_ylabel("$x_1, y_1$", fontsize=18)
ax1.set_yticks([-3, 0, 3])
ax2.plot(tt, x2, "r")
ax2.plot(tt, y2, "b")
ax2.set_xlabel("$t$", fontsize=18)
ax2.set_ylabel("$x_2, y_2$", fontsize=18)
ax2.set_yticks([-3, 0, 3])
ax3.plot(x1, y1, "r", lw=2.0, label="trajectory of pendulum 1")
ax3.plot(x2, y2, "b", lw=0.5, label="trajectory of pendulum 2")
ax3.set_xlabel("$x$", fontsize=18)
ax3.set_ylabel("$y$", fontsize=18)
ax3.set_xticks([-3, 0, 3])
ax3.set_yticks([-3, 0, 3])
ax3.legend()
fig.tight_layout()
fig.savefig("ch9-double-pendulum.pdf")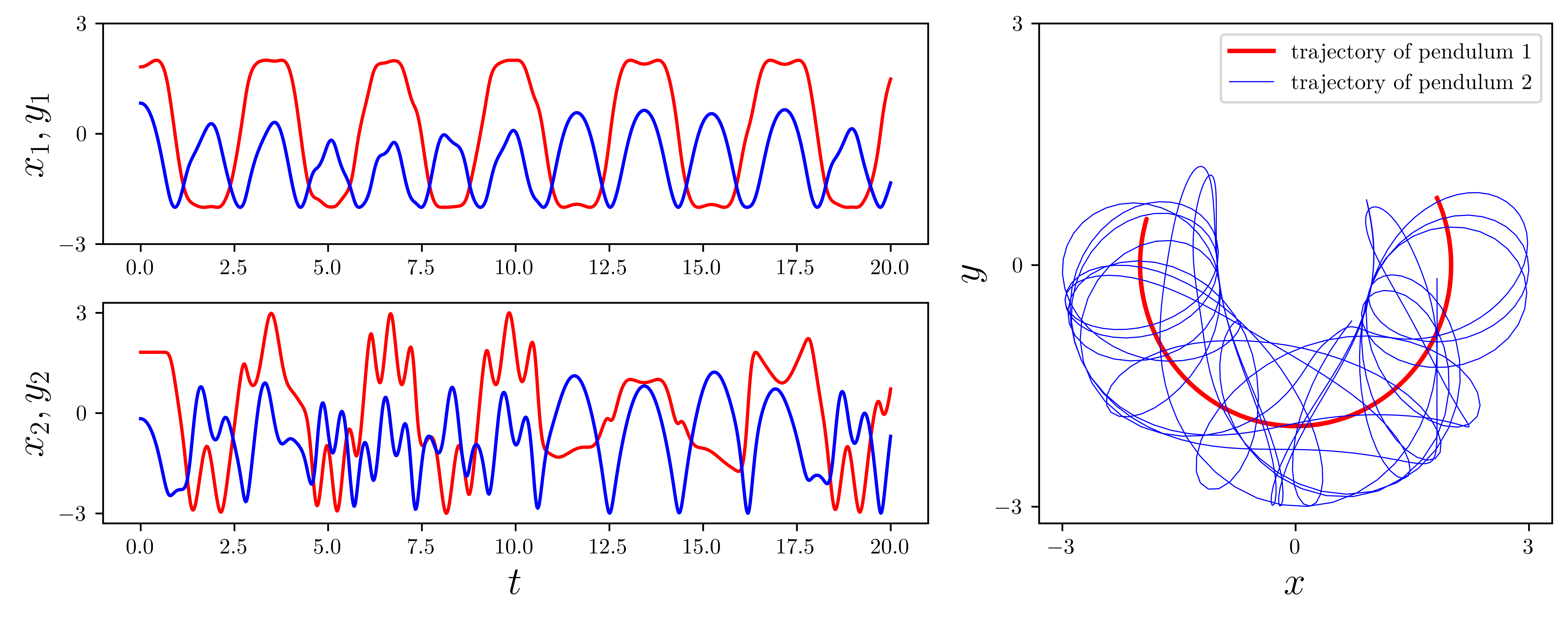
- Johansson, R. (2024). Numerical Python: Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib. Apress. 10.1007/979-8-8688-0413-7