Chapter 10: Sparse matrices and graphs
Robert Johansson
Source code listings for Numerical Python - Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib (ISBN 979-8-8688-0412-0).
%matplotlib inline
%config InlineBackend.figure_format='retina'import matplotlib as mpl
import matplotlib.pyplot as plt
# mpl.rcParams['text.usetex'] = True
# mpl.rcParams['mathtext.fontset'] = 'stix'
# mpl.rcParams['font.family'] = 'serif'
# mpl.rcParams['font.sans-serif'] = 'stix'import scipy.sparse as spimport scipy.sparse.linalgimport numpy as npimport scipy.linalg as laimport networkx as nxCoordinate list format¶
values = [1, 2, 3, 4]rows = [0, 1, 2, 3]cols = [1, 3, 2, 0]A = sp.coo_matrix((values, (rows, cols)), shape=[4, 4])A.todense()matrix([[0, 1, 0, 0],
[0, 0, 0, 2],
[0, 0, 3, 0],
[4, 0, 0, 0]])A<COOrdinate sparse matrix of dtype 'int64'
with 4 stored elements and shape (4, 4)>A.shape, A.size, A.dtype, A.ndim((4, 4), 4, dtype('int64'), 2)A.nnz, A.data(4, array([1, 2, 3, 4]))A.rowarray([0, 1, 2, 3], dtype=int32)A.colarray([1, 3, 2, 0], dtype=int32)A.tocsr()<Compressed Sparse Row sparse matrix of dtype 'int64'
with 4 stored elements and shape (4, 4)>A.toarray()array([[0, 1, 0, 0],
[0, 0, 0, 2],
[0, 0, 3, 0],
[4, 0, 0, 0]])A.todense()matrix([[0, 1, 0, 0],
[0, 0, 0, 2],
[0, 0, 3, 0],
[4, 0, 0, 0]])Not all sparse matrix formats supports indexing:
# A[1, 2]# A.tobsr()[1, 2]But some do:
A.tocsr()[1, 2]np.int64(0)A.tolil()[1:3, 3]<List of Lists sparse matrix of dtype 'int64'
with 1 stored elements and shape (2, 1)>CSR¶
A = np.array([[1, 2, 0, 0], [0, 3, 4, 0], [0, 0, 5, 6], [7, 0, 8, 9]])
Aarray([[1, 2, 0, 0],
[0, 3, 4, 0],
[0, 0, 5, 6],
[7, 0, 8, 9]])A = sp.csr_matrix(A)A.dataarray([1, 2, 3, 4, 5, 6, 7, 8, 9])A.indicesarray([0, 1, 1, 2, 2, 3, 0, 2, 3], dtype=int32)A.indptrarray([0, 2, 4, 6, 9], dtype=int32)i = 2A.indptr[i], A.indptr[i + 1] - 1(np.int32(4), np.int32(5))A.indices[A.indptr[i] : A.indptr[i + 1]]array([2, 3], dtype=int32)A.data[A.indptr[i] : A.indptr[i + 1]]array([5, 6])Functions for constructing sparse matrices¶
N = 10A = -2 * sp.eye(N) + sp.eye(N, k=1) + sp.eye(N, k=-1)A<DIAgonal sparse matrix of dtype 'float64'
with 28 stored elements (3 diagonals) and shape (10, 10)>A.todense()matrix([[-2., 1., 0., 0., 0., 0., 0., 0., 0., 0.],
[ 1., -2., 1., 0., 0., 0., 0., 0., 0., 0.],
[ 0., 1., -2., 1., 0., 0., 0., 0., 0., 0.],
[ 0., 0., 1., -2., 1., 0., 0., 0., 0., 0.],
[ 0., 0., 0., 1., -2., 1., 0., 0., 0., 0.],
[ 0., 0., 0., 0., 1., -2., 1., 0., 0., 0.],
[ 0., 0., 0., 0., 0., 1., -2., 1., 0., 0.],
[ 0., 0., 0., 0., 0., 0., 1., -2., 1., 0.],
[ 0., 0., 0., 0., 0., 0., 0., 1., -2., 1.],
[ 0., 0., 0., 0., 0., 0., 0., 0., 1., -2.]])fig, ax = plt.subplots()
ax.spy(A)
fig.tight_layout()
fig.savefig("ch10-sparse-matrix-1.pdf");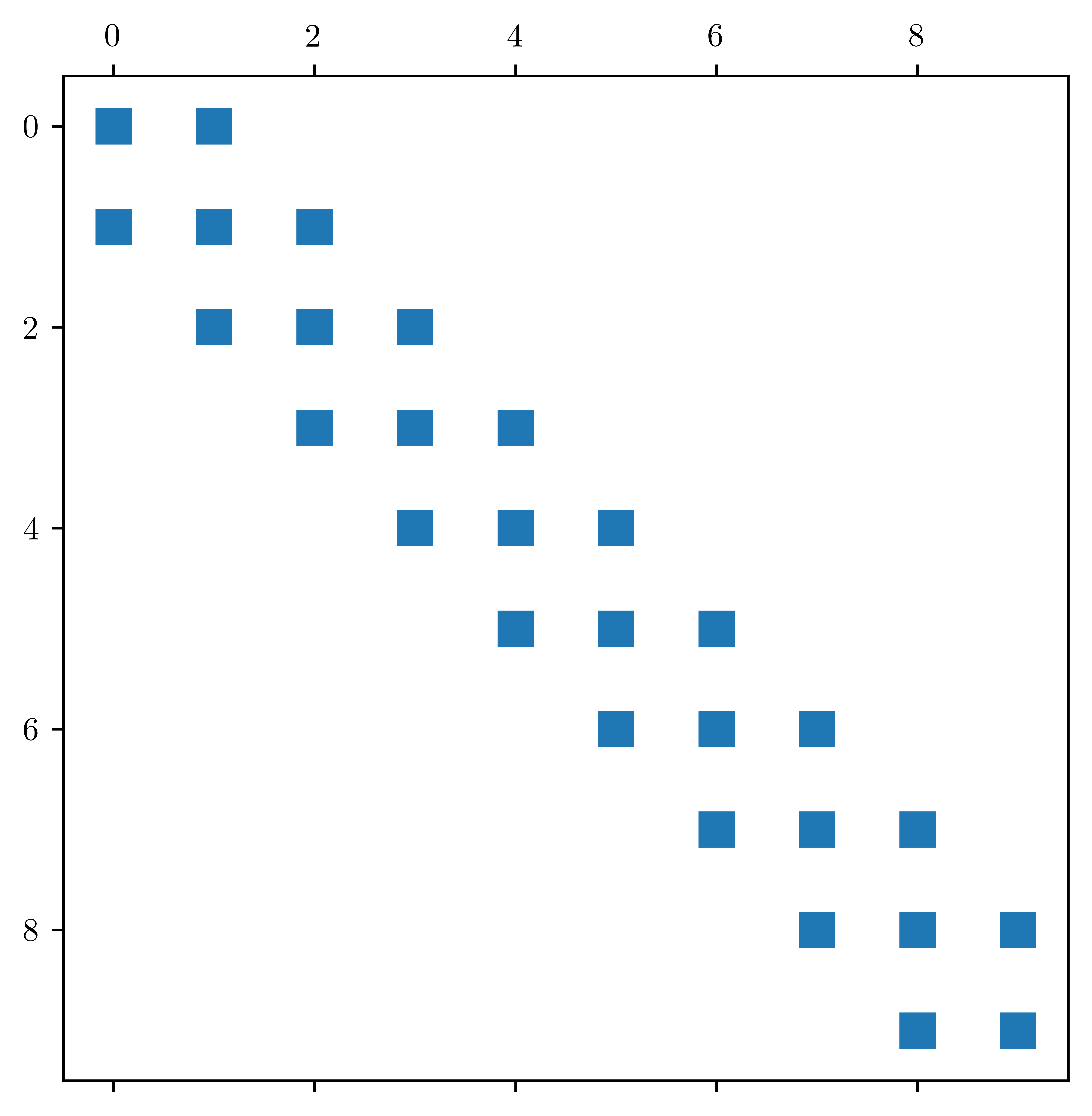
A = sp.diags([1, -2, 1], [1, 0, -1], shape=[N, N], format="csc")/tmp/ipykernel_46607/547814692.py:1: FutureWarning: Input has data type int64, but the output has been cast to float64. In the future, the output data type will match the input. To avoid this warning, set the `dtype` parameter to `None` to have the output dtype match the input, or set it to the desired output data type.
A = sp.diags([1, -2, 1], [1, 0, -1], shape=[N, N], format="csc")
A<Compressed Sparse Column sparse matrix of dtype 'float64'
with 28 stored elements and shape (10, 10)>fig, ax = plt.subplots()
ax.spy(A);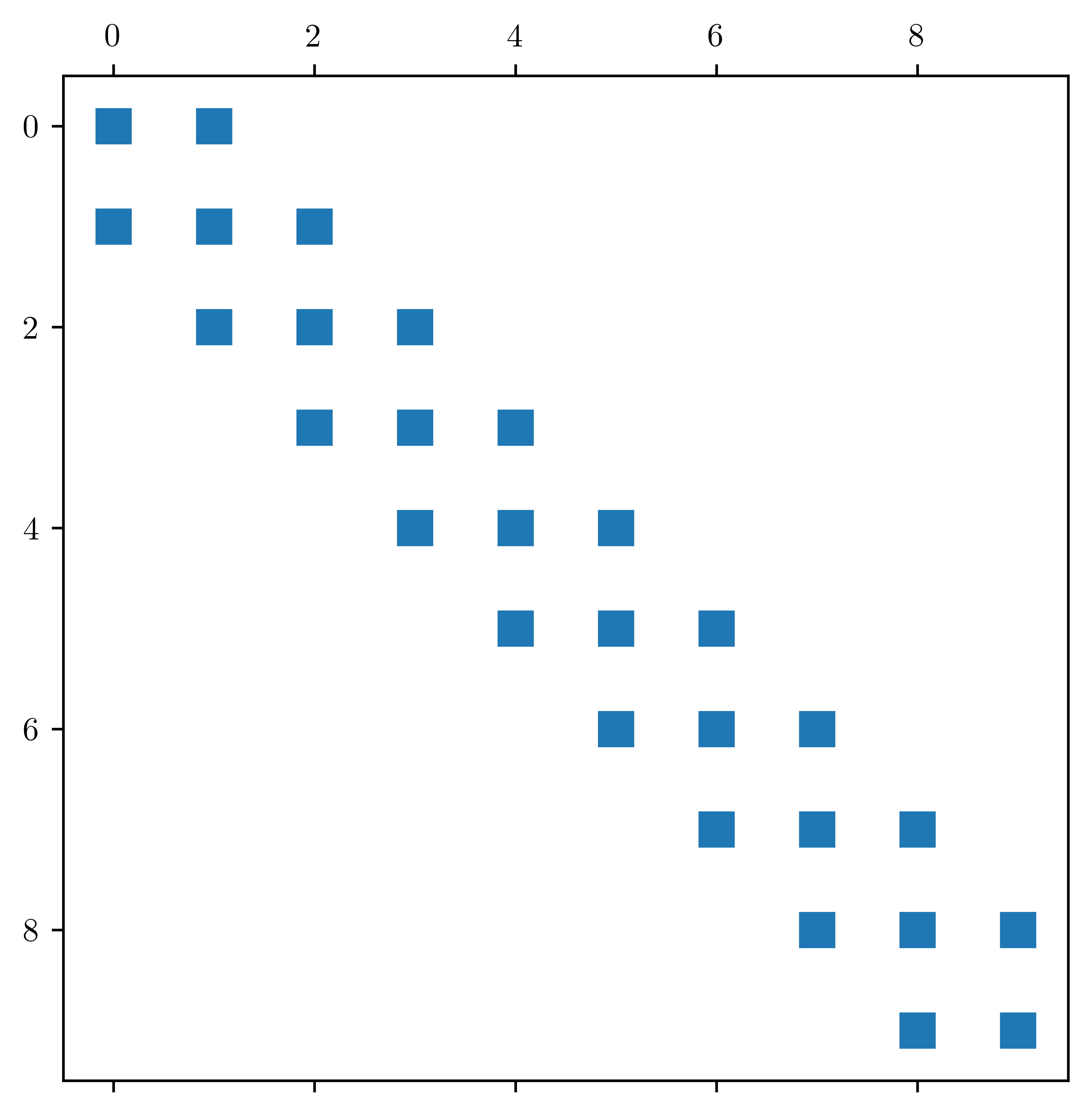
B = sp.diags([1, 1], [-1, 1], shape=[3, 3])/tmp/ipykernel_46607/1445505839.py:1: FutureWarning: Input has data type int64, but the output has been cast to float64. In the future, the output data type will match the input. To avoid this warning, set the `dtype` parameter to `None` to have the output dtype match the input, or set it to the desired output data type.
B = sp.diags([1, 1], [-1, 1], shape=[3, 3])
B<DIAgonal sparse matrix of dtype 'float64'
with 4 stored elements (2 diagonals) and shape (3, 3)>C = sp.kron(A, B, format="csr")
C<Compressed Sparse Row sparse matrix of dtype 'float64'
with 112 stored elements and shape (30, 30)>fig, (ax_A, ax_B, ax_C) = plt.subplots(1, 3, figsize=(12, 4))
ax_A.spy(A)
ax_B.spy(B)
ax_C.spy(C)
fig.tight_layout()
fig.savefig("ch10-sparse-matrix-2.pdf");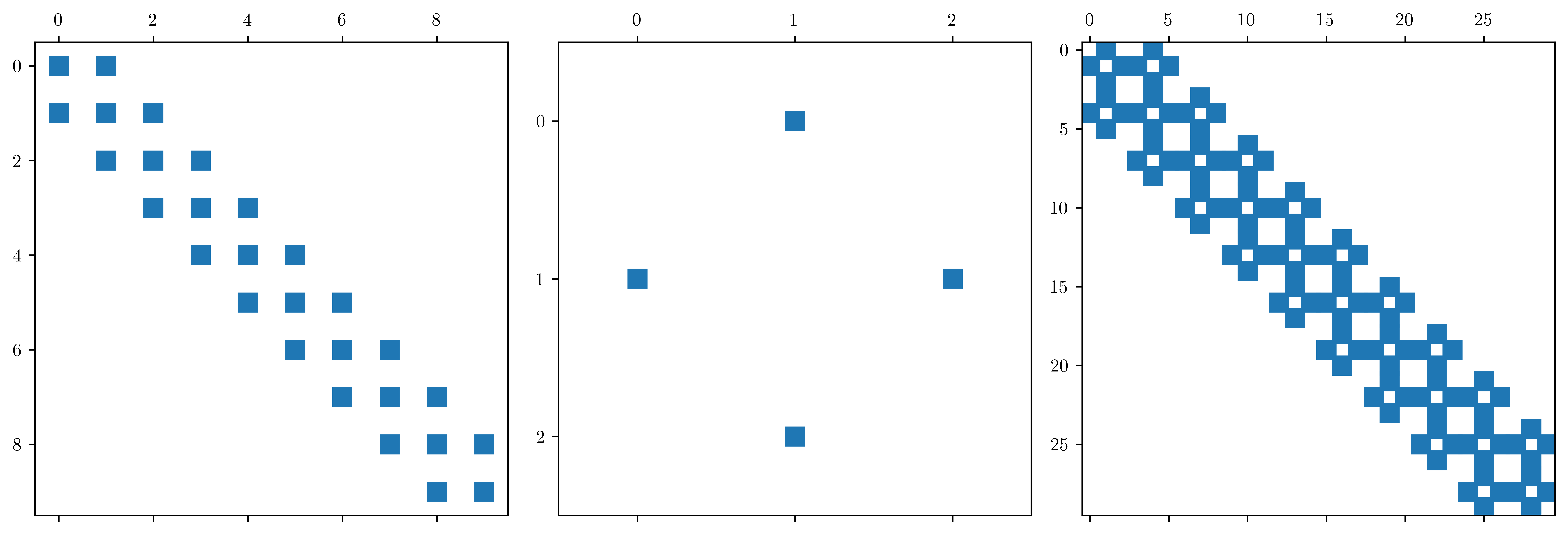
Sparse linear algebra¶
N = 10A = sp.diags([1, -2, 1], [1, 0, -1], shape=[N, N], format="csc")/tmp/ipykernel_46607/547814692.py:1: FutureWarning: Input has data type int64, but the output has been cast to float64. In the future, the output data type will match the input. To avoid this warning, set the `dtype` parameter to `None` to have the output dtype match the input, or set it to the desired output data type.
A = sp.diags([1, -2, 1], [1, 0, -1], shape=[N, N], format="csc")
b = -np.ones(N)x = sp.linalg.spsolve(A, b)xarray([ 5., 9., 12., 14., 15., 15., 14., 12., 9., 5.])np.linalg.solve(A.todense(), b)array([ 5., 9., 12., 14., 15., 15., 14., 12., 9., 5.])lu = sp.linalg.splu(A)lu.L<Compressed Sparse Column sparse array of dtype 'float64'
with 20 stored elements and shape (10, 10)>lu.perm_rarray([0, 1, 2, 3, 4, 5, 6, 8, 7, 9], dtype=int32)lu.U<Compressed Sparse Column sparse array of dtype 'float64'
with 20 stored elements and shape (10, 10)>def sp_permute(A, perm_r, perm_c):
"""permute rows and columns of A"""
M, N = A.shape
# row permumation matrix
Pr = sp.coo_matrix((np.ones(M), (perm_r, np.arange(N)))).tocsr()
# column permutation matrix
Pc = sp.coo_matrix((np.ones(M), (np.arange(M), perm_c))).tocsr()
return Pr.T * A * Pc.Tlu.L * lu.U - A<Compressed Sparse Column sparse array of dtype 'float64'
with 26 stored elements and shape (10, 10)>sp_permute(lu.L * lu.U, lu.perm_r, lu.perm_c) - A<Compressed Sparse Column sparse matrix of dtype 'float64'
with 26 stored elements and shape (10, 10)>fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(12, 4))
ax1.spy(lu.L)
ax2.spy(lu.U)
ax3.spy(A)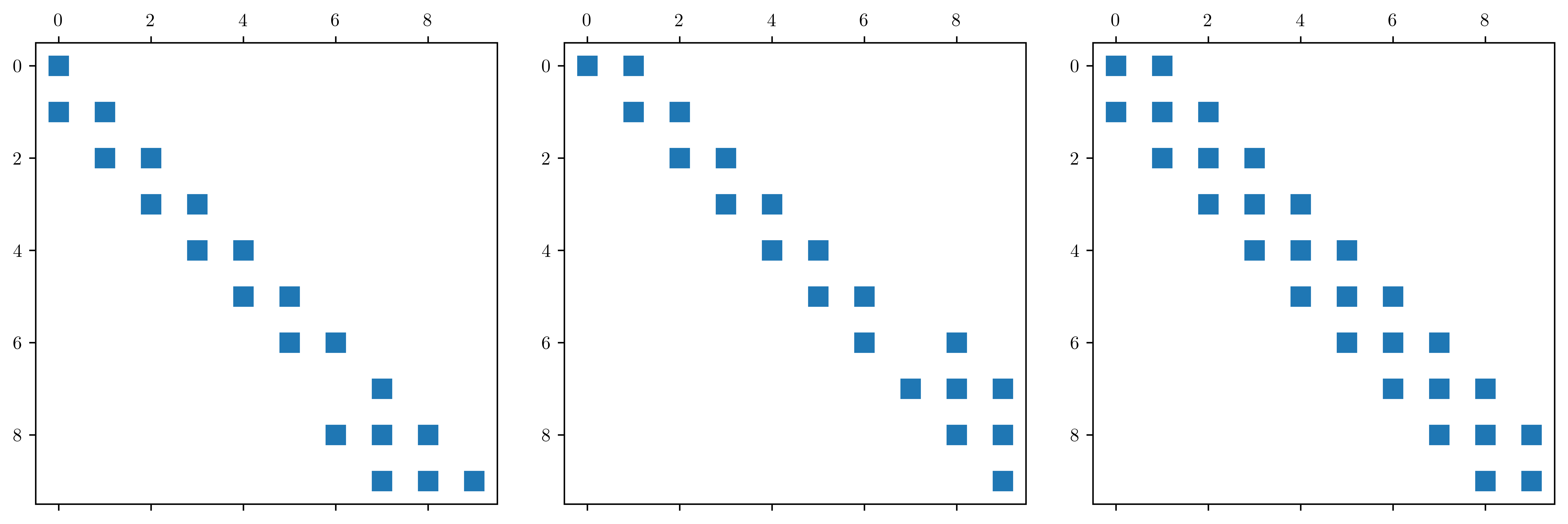
x = lu.solve(b)xarray([ 5., 9., 12., 14., 15., 15., 14., 12., 9., 5.])# use_umfpack=True is only effective if scikit-umfpack is installed
# (in which case UMFPACK is the default solver)
x = sp.linalg.spsolve(A, b, use_umfpack=True)xarray([ 5., 9., 12., 14., 15., 15., 14., 12., 9., 5.])x, info = sp.linalg.cg(A, b)xarray([ 5., 9., 12., 14., 15., 15., 14., 12., 9., 5.])x, info = sp.linalg.bicgstab(A, b)xarray([ 5., 9., 12., 14., 15., 15., 14., 12., 9., 5.])# atol argument is a recent addition
x, info = sp.linalg.lgmres(A, b, atol=1e-5)xarray([ 5., 9., 12., 14., 15., 15., 14., 12., 9., 5.])N = 25An example of a matrix reording method: Reverse Cuthil McKee¶
A = sp.diags([1, -2, 1], [8, 0, -8], shape=[N, N], format="csc")/tmp/ipykernel_46607/3160634348.py:1: FutureWarning: Input has data type int64, but the output has been cast to float64. In the future, the output data type will match the input. To avoid this warning, set the `dtype` parameter to `None` to have the output dtype match the input, or set it to the desired output data type.
A = sp.diags([1, -2, 1], [8, 0, -8], shape=[N, N], format="csc")
perm = sp.csgraph.reverse_cuthill_mckee(A)
permarray([23, 15, 7, 22, 14, 6, 21, 13, 5, 20, 12, 4, 19, 11, 3, 18, 10,
2, 17, 9, 1, 24, 16, 8, 0], dtype=int32)fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 4))
ax1.spy(A)
ax2.spy(sp_permute(A, perm, perm))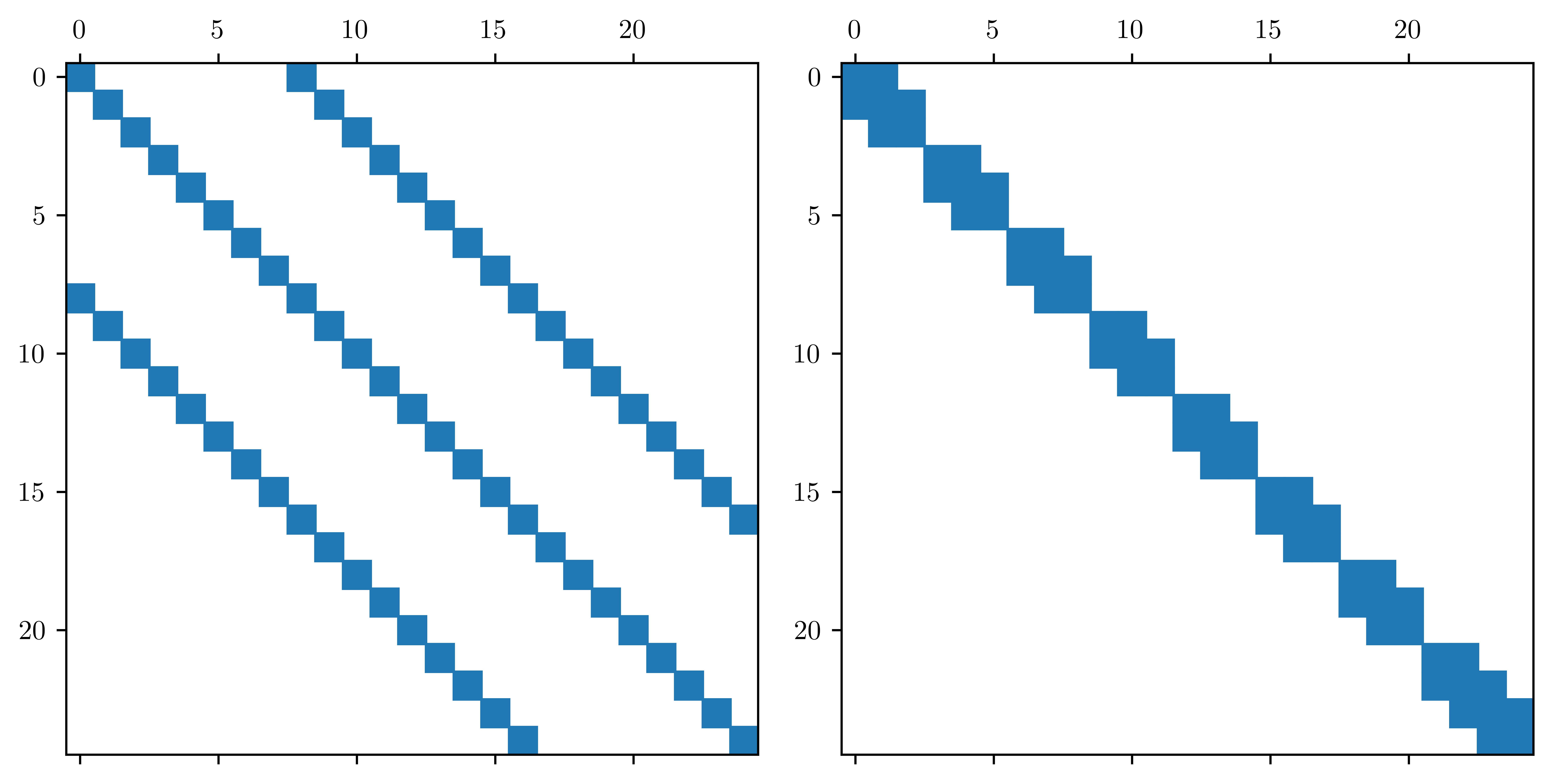
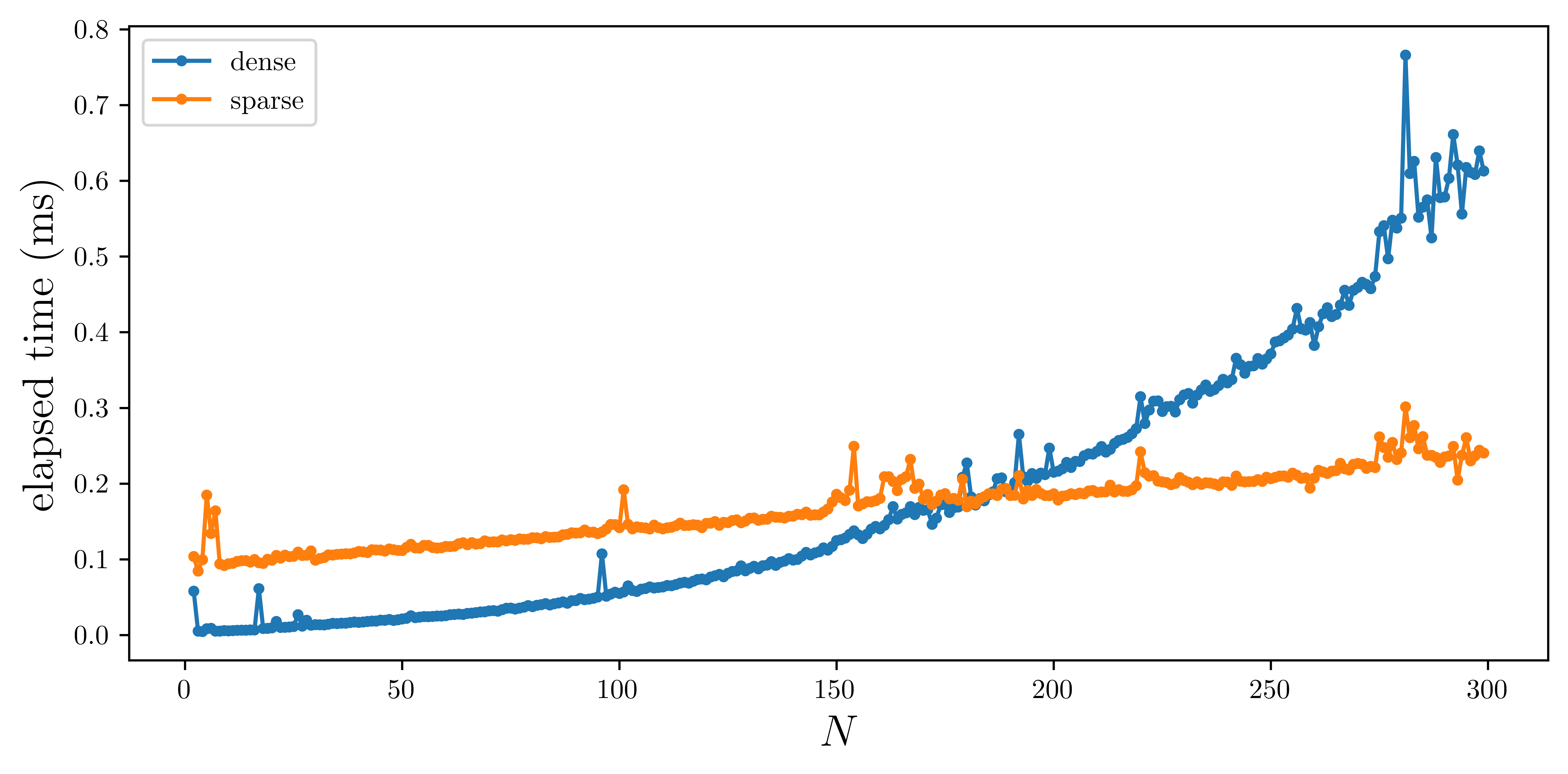
Performance comparison sparse/dense¶
# compare performance of solving Ax=b vs system size N,
# where A is the sparse matrix for the 1d poisson problem
import time
def setup(N):
A = sp.diags([1, -2, 1], [1, 0, -1], shape=[N, N], format="csr")
b = -np.ones(N)
return A, A.todense(), b
reps = 100
N_vec = np.arange(2, 300, 1)
t_sparse = np.empty(len(N_vec))
t_dense = np.empty(len(N_vec))
for idx, N in enumerate(N_vec):
A, A_dense, b = setup(N)
t = time.time()
for r in range(reps):
x = np.linalg.solve(A_dense, b)
t_dense[idx] = (time.time() - t) / reps
t = time.time()
for r in range(reps):
x = sp.linalg.spsolve(A, b, use_umfpack=True)
t_sparse[idx] = (time.time() - t) / reps
fig, ax = plt.subplots(figsize=(8, 4))
ax.plot(N_vec, t_dense * 1e3, ".-", label="dense")
ax.plot(N_vec, t_sparse * 1e3, ".-", label="sparse")
ax.set_xlabel(r"$N$", fontsize=16)
ax.set_ylabel("elapsed time (ms)", fontsize=16)
ax.legend(loc=0)
fig.tight_layout()
fig.savefig("ch10-sparse-vs-dense.pdf")/tmp/ipykernel_46607/537225260.py:7: FutureWarning: Input has data type int64, but the output has been cast to float64. In the future, the output data type will match the input. To avoid this warning, set the `dtype` parameter to `None` to have the output dtype match the input, or set it to the desired output data type.
A = sp.diags([1, -2, 1], [1, 0, -1], shape=[N, N], format="csr")
Eigenvalue problems¶
N = 10A = sp.diags([1, -2, 1], [1, 0, -1], shape=[N, N], format="csc")/tmp/ipykernel_46607/547814692.py:1: FutureWarning: Input has data type int64, but the output has been cast to float64. In the future, the output data type will match the input. To avoid this warning, set the `dtype` parameter to `None` to have the output dtype match the input, or set it to the desired output data type.
A = sp.diags([1, -2, 1], [1, 0, -1], shape=[N, N], format="csc")
evals, evecs = sp.linalg.eigs(A, k=4, which="LM")evalsarray([-3.91898595+0.j, -3.68250707+0.j, -3.30972147+0.j, -2.83083003+0.j])np.allclose(A.dot(evecs[:, 0]), evals[0] * evecs[:, 0])Trueevals, evecs = sp.linalg.eigsh(A, k=4, which="LM")evalsarray([-3.91898595, -3.68250707, -3.30972147, -2.83083003])evals, evecs = sp.linalg.eigs(A, k=4, which="SR")evalsarray([-3.91898595+0.j, -3.68250707+0.j, -3.30972147+0.j, -2.83083003+0.j])np.real(evals).argsort()array([0, 1, 2, 3])def sp_eigs_sorted(A, k=6, which="SR"):
"""compute and return eigenvalues sorted by real value"""
evals, evecs = sp.linalg.eigs(A, k=k, which=which)
idx = np.real(evals).argsort()
return evals[idx], evecs[idx]evals, evecs = sp_eigs_sorted(A, k=4, which="SM")evalsarray([-1.16916997+0.j, -0.69027853+0.j, -0.31749293+0.j, -0.08101405+0.j])Random matrix example¶
N = 100x_vec = np.linspace(0, 1, 50)# seed sp.rand with random_state to obtain a reproducible result
M1 = sp.rand(N, N, density=0.2, random_state=112312321)
# M1 = M1 + M1.conj().T
M2 = sp.rand(N, N, density=0.2, random_state=984592134)
# M2 = M2 + M2.conj().Tevals = np.array([sp_eigs_sorted((1 - x) * M1 + x * M2, k=25)[0] for x in x_vec])fig, ax = plt.subplots(figsize=(8, 4))
for idx in range(evals.shape[1]):
ax.plot(x_vec, np.real(evals[:, idx]), lw=0.5)
ax.set_xlabel(r"$x$", fontsize=16)
ax.set_ylabel(r"eig.vals. of $(1-x)M_1+xM_2$", fontsize=16)
fig.tight_layout()
fig.savefig("ch10-sparse-eigs.pdf")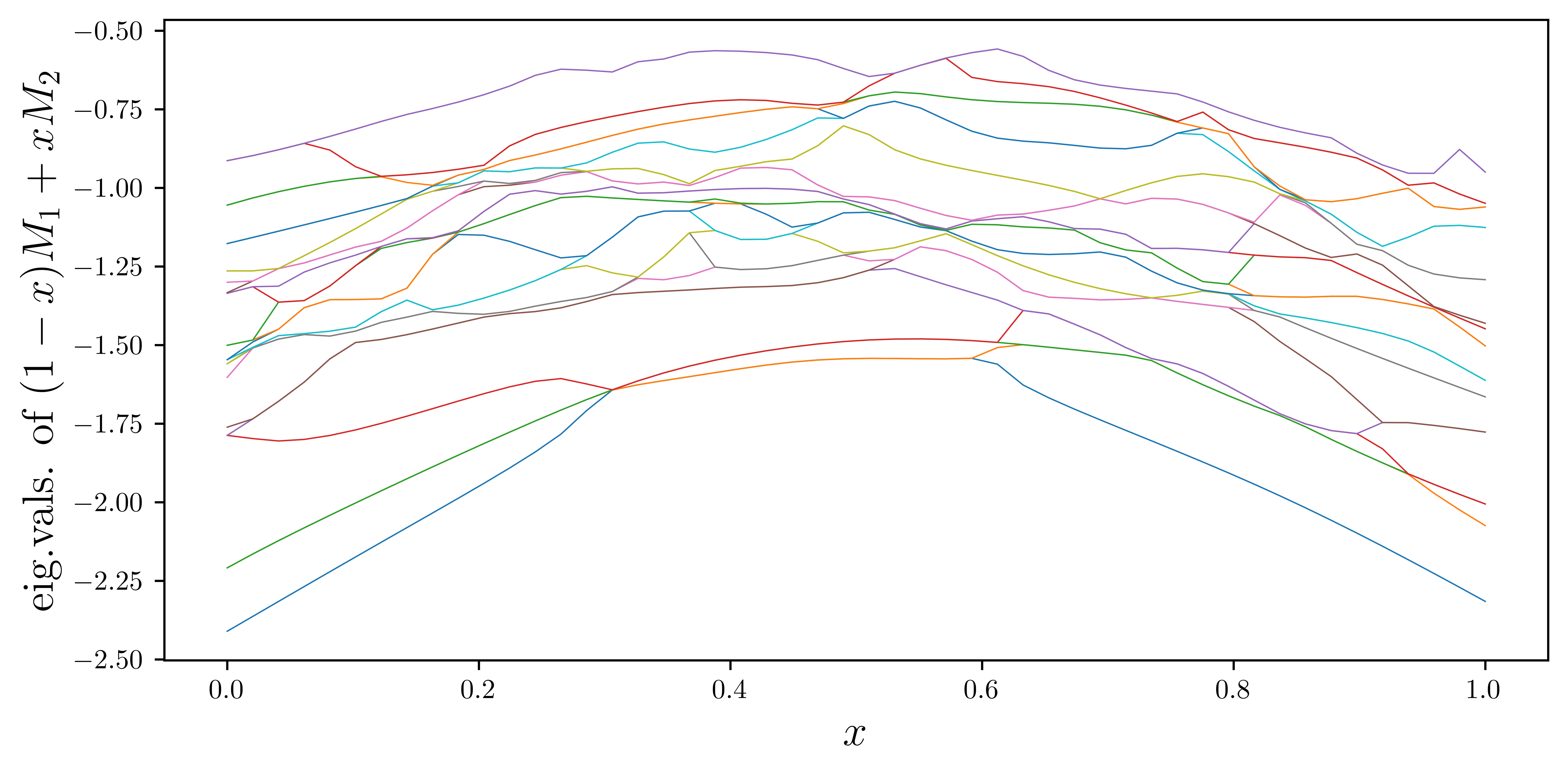
Graphs¶
g = nx.Graph()g.add_node(1)g.nodes()NodeView((1,))g.add_nodes_from([3, 4, 5])g.nodes()NodeView((1, 3, 4, 5))g.add_edge(1, 2)g.edges()EdgeView([(1, 2)])g.add_edges_from([(3, 4), (5, 6)])g.edges()EdgeView([(1, 2), (3, 4), (5, 6)])g.add_weighted_edges_from([(1, 3, 1.5), (3, 5, 2.5)])g.edges()EdgeView([(1, 2), (1, 3), (3, 4), (3, 5), (5, 6)])g.edges(data=True)EdgeDataView([(1, 2, {}), (1, 3, {'weight': 1.5}), (3, 4, {}), (3, 5, {'weight': 2.5}), (5, 6, {})])g.add_weighted_edges_from([(6, 7, 1.5)])g.nodes()NodeView((1, 3, 4, 5, 2, 6, 7))g.edges()EdgeView([(1, 2), (1, 3), (3, 4), (3, 5), (5, 6), (6, 7)])import numpy as npimport jsonwith open("tokyo-metro.json") as f:
data = json.load(f)data.keys()dict_keys(['C', 'G', 'F', 'H', 'M', 'N', 'T', 'Y', 'Z'])data["C"]{'color': '#149848',
'transfers': [['C3', 'F15'],
['C4', 'Z2'],
['C4', 'G2'],
['C7', 'M14'],
['C7', 'N6'],
['C7', 'G6'],
['C8', 'M15'],
['C8', 'H6'],
['C9', 'H7'],
['C9', 'Y18'],
['C11', 'T9'],
['C11', 'M18'],
['C11', 'Z8'],
['C12', 'M19'],
['C18', 'H21']],
'travel_times': [['C1', 'C2', 2],
['C2', 'C3', 2],
['C3', 'C4', 1],
['C4', 'C5', 2],
['C5', 'C6', 2],
['C6', 'C7', 2],
['C7', 'C8', 1],
['C8', 'C9', 3],
['C9', 'C10', 1],
['C10', 'C11', 2],
['C11', 'C12', 2],
['C12', 'C13', 2],
['C13', 'C14', 2],
['C14', 'C15', 2],
['C15', 'C16', 2],
['C16', 'C17', 3],
['C17', 'C18', 3],
['C18', 'C19', 3]]}# datag = nx.Graph()
for line in data.values():
g.add_weighted_edges_from(line["travel_times"])
g.add_edges_from(line["transfers"])for n1, n2 in g.edges():
g[n1][n2]["transfer"] = "weight" not in g[n1][n2]g.number_of_nodes()184list(g.nodes())[:5]['C1', 'C2', 'C3', 'C4', 'C5']g.number_of_edges()243list(g.edges())[:5][('C1', 'C2'), ('C2', 'C3'), ('C3', 'C4'), ('C3', 'F15'), ('C4', 'C5')]on_foot = [edge for edge in g.edges() if g.get_edge_data(*edge)["transfer"]]on_train = [edge for edge in g.edges() if not g.get_edge_data(*edge)["transfer"]]colors = [data[n[0].upper()]["color"] for n in g.nodes()]# from networkx.drawing.nx_agraph import graphviz_layoutfig, ax = plt.subplots(1, 1, figsize=(14, 10))
pos = nx.drawing.nx_agraph.graphviz_layout(g, prog="neato")
nx.draw(g, pos, ax=ax, node_size=300, node_color=colors)
nx.draw_networkx_labels(g, pos=pos, ax=ax, font_size=6)
nx.draw_networkx_edges(g, pos=pos, ax=ax, edgelist=on_train, width=2)
nx.draw_networkx_edges(g, pos=pos, ax=ax, edgelist=on_foot, edge_color="blue")
# removing the default axis on all sides:
for side in ["bottom", "right", "top", "left"]:
ax.spines[side].set_visible(False)
# removing the axis labels and ticks
ax.set_xticks([])
ax.set_yticks([])
ax.xaxis.set_ticks_position("none")
ax.yaxis.set_ticks_position("none")
fig.tight_layout()
fig.savefig("ch10-metro-graph.pdf")
fig.savefig("ch10-metro-graph.png")
g.degree()DegreeView({'C1': 1, 'C2': 2, 'C3': 3, 'C4': 4, 'C5': 2, 'C6': 2, 'C7': 5, 'C8': 4, 'C9': 4, 'C10': 2, 'C11': 5, 'C12': 3, 'C13': 2, 'C14': 2, 'C15': 2, 'C16': 2, 'C17': 2, 'C18': 3, 'C19': 1, 'F15': 3, 'Z2': 4, 'G2': 4, 'M14': 5, 'N6': 5, 'G6': 5, 'M15': 4, 'H6': 4, 'H7': 4, 'Y18': 4, 'T9': 5, 'M18': 5, 'Z8': 5, 'M19': 3, 'H21': 2, 'G1': 3, 'G3': 2, 'G4': 3, 'G5': 6, 'G7': 2, 'G8': 2, 'G9': 4, 'G10': 2, 'G11': 3, 'G12': 3, 'G13': 2, 'G14': 2, 'G15': 3, 'G16': 3, 'G17': 2, 'G18': 2, 'G19': 1, 'Z1': 3, 'F16': 3, 'Z3': 3, 'M13': 6, 'Y16': 6, 'Z4': 6, 'N7': 6, 'M16': 4, 'H8': 4, 'T10': 3, 'Z9': 3, 'H16': 3, 'H17': 3, 'F1': 2, 'F2': 3, 'F3': 3, 'F4': 3, 'F5': 3, 'F6': 3, 'F7': 3, 'F8': 3, 'F9': 4, 'F10': 2, 'F11': 2, 'F12': 2, 'F13': 3, 'F14': 2, 'Y1': 2, 'Y2': 3, 'Y3': 3, 'Y4': 3, 'Y5': 3, 'Y6': 3, 'Y7': 3, 'Y8': 3, 'Y9': 4, 'M25': 3, 'M9': 3, 'H1': 1, 'H2': 2, 'H3': 2, 'H4': 2, 'H5': 2, 'H9': 2, 'H10': 2, 'H11': 2, 'H12': 3, 'H13': 2, 'H14': 2, 'H15': 2, 'H18': 2, 'H19': 2, 'H20': 2, 'T11': 3, 'M1': 1, 'M2': 2, 'M3': 2, 'M4': 2, 'M5': 2, 'M6': 3, 'M7': 2, 'M8': 2, 'M10': 2, 'M11': 2, 'M12': 3, 'M17': 2, 'M20': 2, 'M21': 2, 'M22': 3, 'M23': 2, 'M24': 2, 'm3': 1, 'm4': 2, 'm5': 2, 'N8': 3, 'N11': 3, 'N1': 2, 'N2': 3, 'N3': 3, 'N4': 2, 'N5': 2, 'N9': 3, 'N10': 4, 'N12': 2, 'N13': 2, 'N14': 2, 'N15': 2, 'N16': 2, 'N17': 2, 'N18': 2, 'N19': 1, 'T1': 2, 'T2': 3, 'T3': 3, 'Y14': 3, 'Y13': 4, 'T6': 4, 'T4': 2, 'T5': 2, 'T7': 3, 'T8': 2, 'T12': 2, 'T13': 2, 'T14': 2, 'T15': 2, 'T16': 2, 'T17': 2, 'T18': 2, 'T19': 2, 'T20': 2, 'T21': 2, 'T22': 2, 'T23': 2, 'T24': 1, 'Z6': 3, 'Y10': 2, 'Y11': 2, 'Y12': 2, 'Y15': 2, 'Y17': 2, 'Y19': 2, 'Y20': 2, 'Y21': 2, 'Y22': 2, 'Y23': 2, 'Y24': 1, 'Z5': 2, 'Z7': 2, 'Z10': 2, 'Z11': 2, 'Z12': 2, 'Z13': 2, 'Z14': 1})d_max = max(d for (n, d) in g.degree())[(n, d) for (n, d) in g.degree() if d == d_max][('G5', 6), ('M13', 6), ('Y16', 6), ('Z4', 6), ('N7', 6)]p = nx.shortest_path(g, "Y24", "C19")np.array(p)array(['Y24', 'Y23', 'Y22', 'Y21', 'Y20', 'Y19', 'Y18', 'C9', 'C10',
'C11', 'C12', 'C13', 'C14', 'C15', 'C16', 'C17', 'C18', 'C19'],
dtype='<U3')np.sum(
[
g[p[n]][p[n + 1]]["weight"]
for n in range(len(p) - 1)
if "weight" in g[p[n]][p[n + 1]]
]
)np.int64(35)h = g.copy()for n1, n2 in h.edges():
if "transfer" in h[n1][n2]:
h[n1][n2]["weight"] = 5p = nx.shortest_path(h, "Y24", "C19")np.array(p)array(['Y24', 'Y23', 'Y22', 'Y21', 'Y20', 'Y19', 'Y18', 'C9', 'C10',
'C11', 'C12', 'C13', 'C14', 'C15', 'C16', 'C17', 'C18', 'C19'],
dtype='<U3')np.sum([h[p[n]][p[n + 1]]["weight"] for n in range(len(p) - 1)])np.int64(85)p = nx.shortest_path(h, "Z1", "H16")np.sum([h[p[n]][p[n + 1]]["weight"] for n in range(len(p) - 1)])np.int64(65)# A = nx.to_scipy_sparse_matrix(g)# A = nx.to_scipy_sparse_array(g)A<Compressed Sparse Column sparse matrix of dtype 'float64'
with 28 stored elements and shape (10, 10)>perm = sp.csgraph.reverse_cuthill_mckee(A)fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 4))
ax1.spy(A, markersize=2)
ax2.spy(sp_permute(A, perm, perm), markersize=2)
fig.tight_layout()
fig.savefig("ch12-rcm-graph.pdf")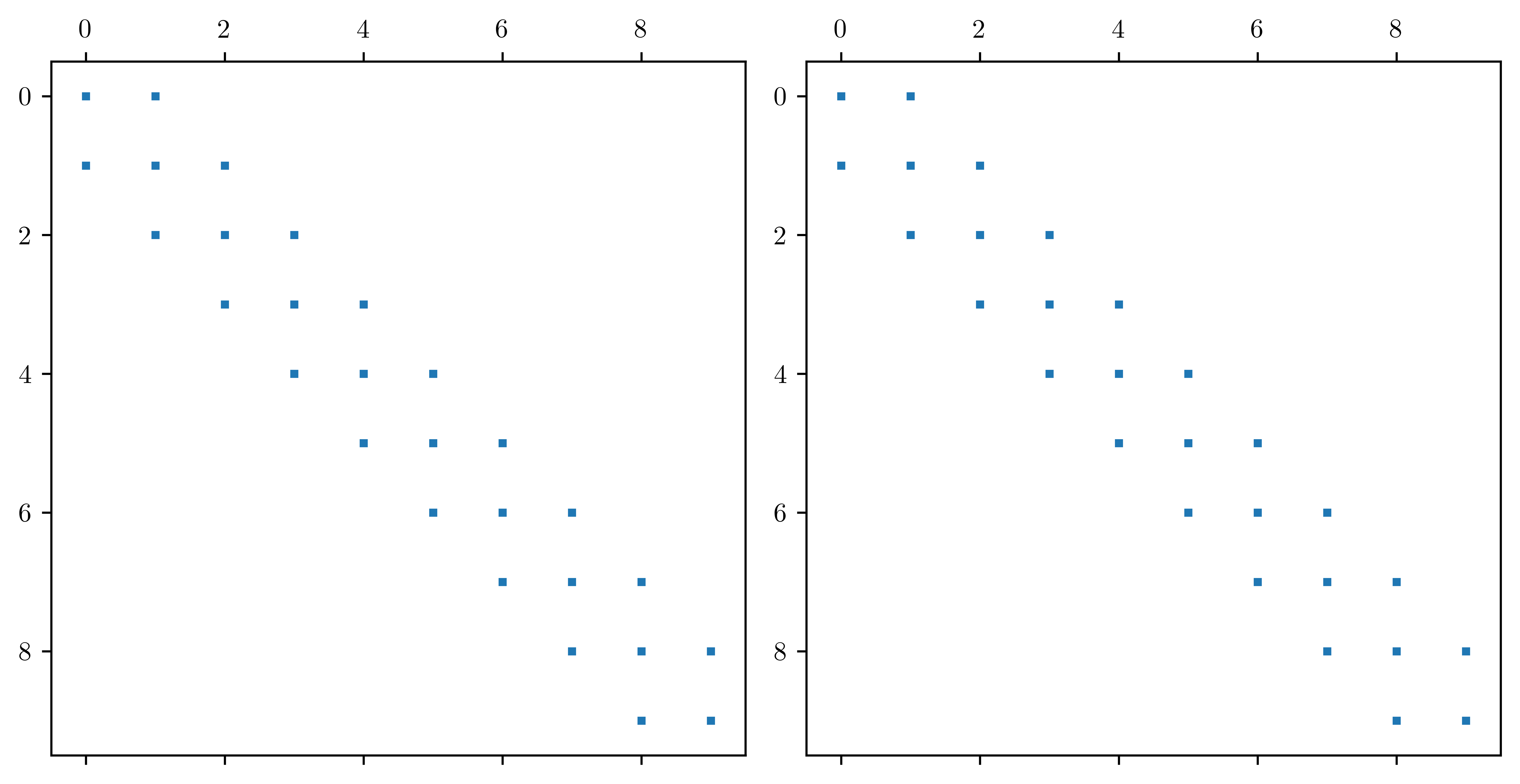
- Johansson, R. (2024). Numerical Python: Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib. Apress. 10.1007/979-8-8688-0413-7