Esta investigación se centró en el caso de los pagos impagos de los clientes en Taiwán y compara la precisión predictiva de la probabilidad de impago entre seis métodos de minería de datos. Desde la perspectiva de la gestión de riesgos, el resultado de la precisión predictiva de la probabilidad estimada de impago será más valioso que el resultado binario de la clasificación: clientes creíbles o no creíbles. Debido a que se desconoce la probabilidad real de impago, este estudio presentó el novedoso método de suavizado de clasificación para estimar la probabilidad real de impago. Con la probabilidad real de impago como variable de respuesta (Y) y la probabilidad predictiva de impago como variable independiente (X), el resultado de la regresión lineal simple (Y = A + BX) muestra que el modelo de pronóstico producido por la red neuronal artificial tiene el coeficiente de determinación más alto; su intersección de regresión (A) es cercana a cero y el coeficiente de regresión (B) a uno. Por lo tanto, entre las seis técnicas de minería de datos, la red neuronal artificial es la única que puede estimar con precisión la probabilidad real de impago.
La base de datos se descarga directamente de la plataforma: “UCI-Machine Learning Repository”, a continuación se comparte el link:
https://
El link de descarga de la base de datos: https://archive.ics.uci.edu/ml/machine-learning-databases/00350/default of credit card clients.xls
Tomamemos en cuenta algunos detalles:
Variable objetivo (y): BILL_AMT1 → monto de la factura de septiembre.
Variables predictoras (X): todas las demás columnas excepto:
ID (identificador irrelevante),
default payment next month (que es la variable objetivo para clasificación, pero aquí no se usa)
Librerías¶
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sbn
from scipy.stats import zscore
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler, StandardScalerurl = "https://archive.ics.uci.edu/ml/machine-learning-databases/00350/default%20of%20credit%20card%20clients.xls"
df = pd.read_excel(url, header=1)
np.set_printoptions(suppress=True)dfdf.shape(30000, 25)len(df)30000df.info()<class 'pandas.core.frame.DataFrame'>
RangeIndex: 30000 entries, 0 to 29999
Data columns (total 25 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 ID 30000 non-null int64
1 LIMIT_BAL 30000 non-null int64
2 SEX 30000 non-null int64
3 EDUCATION 30000 non-null int64
4 MARRIAGE 30000 non-null int64
5 AGE 30000 non-null int64
6 PAY_0 30000 non-null int64
7 PAY_2 30000 non-null int64
8 PAY_3 30000 non-null int64
9 PAY_4 30000 non-null int64
10 PAY_5 30000 non-null int64
11 PAY_6 30000 non-null int64
12 BILL_AMT1 30000 non-null int64
13 BILL_AMT2 30000 non-null int64
14 BILL_AMT3 30000 non-null int64
15 BILL_AMT4 30000 non-null int64
16 BILL_AMT5 30000 non-null int64
17 BILL_AMT6 30000 non-null int64
18 PAY_AMT1 30000 non-null int64
19 PAY_AMT2 30000 non-null int64
20 PAY_AMT3 30000 non-null int64
21 PAY_AMT4 30000 non-null int64
22 PAY_AMT5 30000 non-null int64
23 PAY_AMT6 30000 non-null int64
24 default payment next month 30000 non-null int64
dtypes: int64(25)
memory usage: 5.7 MB
# Definir objetivo continuo (ej: monto de la factura de septiembre)
y = df["BILL_AMT1"].astype(float)
X = df.drop(["ID", "default payment next month", "BILL_AMT1"], axis=1)Preprocesamiento de datos¶
Variables Numéricas
numeric_vars = df.select_dtypes(include=[np.number]).columns.tolist()
print("Variables numéricas:", numeric_vars)Variables numéricas: ['ID', 'LIMIT_BAL', 'SEX', 'EDUCATION', 'MARRIAGE', 'AGE', 'PAY_0', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6', 'BILL_AMT1', 'BILL_AMT2', 'BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6', 'PAY_AMT1', 'PAY_AMT2', 'PAY_AMT3', 'PAY_AMT4', 'PAY_AMT5', 'PAY_AMT6', 'default payment next month']
numeric_vars['ID',
'LIMIT_BAL',
'SEX',
'EDUCATION',
'MARRIAGE',
'AGE',
'PAY_0',
'PAY_2',
'PAY_3',
'PAY_4',
'PAY_5',
'PAY_6',
'BILL_AMT1',
'BILL_AMT2',
'BILL_AMT3',
'BILL_AMT4',
'BILL_AMT5',
'BILL_AMT6',
'PAY_AMT1',
'PAY_AMT2',
'PAY_AMT3',
'PAY_AMT4',
'PAY_AMT5',
'PAY_AMT6',
'default payment next month']Histogramas de todas las variables numéricas
plt.figure(figsize=(20, 20))
for i, col in enumerate(numeric_vars):
plt.subplot(5, 5, i + 1)
sbn.histplot(df[col], kde=True)
plt.title(col)
plt.tight_layout()
plt.show()
Boxplot para ver outliers
plt.figure(figsize=(20, 20))
for i, col in enumerate(numeric_vars):
plt.subplot(5, 5, i + 1)
sbn.boxplot(x=df[col]) # col: columna
plt.title(col)
plt.tight_layout()
plt.show()
plt.figure(figsize=(20, 20))
for i, col in enumerate(numeric_vars):
plt.subplot(5, 5, i + 1)
sbn.boxplot(y=df[col]) # Vertical
plt.title(col)
plt.tight_layout()
plt.show()
#RANGO INTERCUARTILICO
outliers_info = []
for col in numeric_vars:
Q1 = df[col].quantile(0.25) # CUARTIL DE 25%
Q3 = df[col].quantile(0.75) # CUARTIL DE 75%
IQR = Q3 - Q1
lower = Q1 - 1.5 * IQR
upper = Q3 + 1.5 * IQR
outliers = df[(df[col] < lower) | (df[col] > upper)]
count = outliers.shape[0]
prop = count / df.shape[0]
outliers_info.append(
{"Variable": col, "Outliers": count, "Proporción": round(prop, 4)}
)
outliers_df = pd.DataFrame(outliers_info)
print(outliers_df.sort_values("Outliers", ascending=False)) Variable Outliers Proporción
24 default payment next month 6636 0.2212
7 PAY_2 4410 0.1470
8 PAY_3 4209 0.1403
9 PAY_4 3508 0.1169
6 PAY_0 3130 0.1043
11 PAY_6 3079 0.1026
21 PAY_AMT4 2994 0.0998
10 PAY_5 2968 0.0989
23 PAY_AMT6 2958 0.0986
22 PAY_AMT5 2945 0.0982
18 PAY_AMT1 2745 0.0915
16 BILL_AMT5 2725 0.0908
19 PAY_AMT2 2714 0.0905
17 BILL_AMT6 2693 0.0898
15 BILL_AMT4 2622 0.0874
20 PAY_AMT3 2598 0.0866
14 BILL_AMT3 2469 0.0823
12 BILL_AMT1 2400 0.0800
13 BILL_AMT2 2395 0.0798
3 EDUCATION 454 0.0151
5 AGE 272 0.0091
1 LIMIT_BAL 167 0.0056
2 SEX 0 0.0000
0 ID 0 0.0000
4 MARRIAGE 0 0.0000
Barplot
plt.figure(figsize=(12, 6))
sbn.barplot(
data=outliers_df.sort_values("Outliers", ascending=False),
x="Variable",
y="Outliers",
hue="Variable",
palette="viridis",
legend=False,
)
plt.xticks(rotation=90)
plt.title("Cantidad de outliers por variable (IQR)")
plt.tight_layout()
plt.show()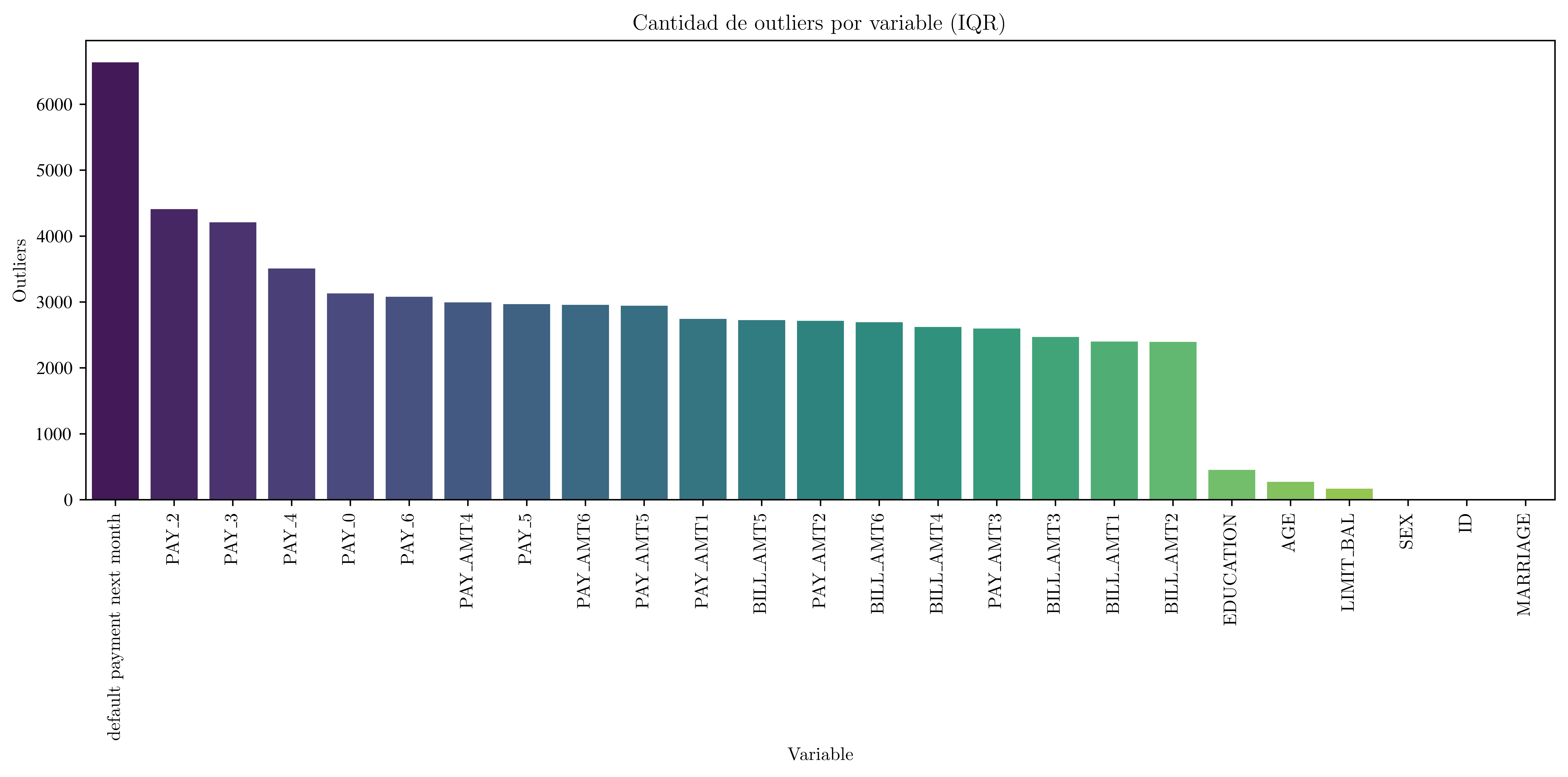
Corte de outliers: Z-Scores
df_z = df[numeric_vars].apply(zscore)
df_no_outliers_z = df[(np.abs(df_z) < 3).all(axis=1)]
print("Filas sin outliers (Z-Score):", len(df_no_outliers_z))Filas sin outliers (Z-Score): 26429
Corte de outliers con: IQR
df_no_outliers_iqr = df.copy()
for col in numeric_vars:
Q1 = df[col].quantile(0.25)
Q3 = df[col].quantile(0.75)
IQR = Q3 - Q1
lower = Q1 - 1.5 * IQR
upper = Q3 + 1.5 * IQR
df_no_outliers_iqr = df_no_outliers_iqr[
(df_no_outliers_iqr[col] >= lower) & (df_no_outliers_iqr[col] <= upper)
]
print("Filas sin outliers (IQR):", len(df_no_outliers_iqr))Filas sin outliers (IQR): 10993
Boxplots con el DataFrame limpio
import matplotlib.pyplot as plt
import seaborn as sbn
plt.figure(figsize=(20, 20))
for i, col in enumerate(numeric_vars):
plt.subplot(5, 5, i + 1)
sbn.boxplot(y=df_no_outliers_iqr[col]) # horizontal
plt.title(f"{col} (IQR)")
plt.tight_layout()
plt.show()
Detección de datos perdidos
missing = df.isnull().sum()
missing_percent = (missing / len(df)) * 100
missing_df = pd.DataFrame({"Valores perdidos": missing, "Porcentaje": missing_percent})
print(missing_df) Valores perdidos Porcentaje
ID 0 0.0
LIMIT_BAL 0 0.0
SEX 0 0.0
EDUCATION 0 0.0
MARRIAGE 0 0.0
AGE 0 0.0
PAY_0 0 0.0
PAY_2 0 0.0
PAY_3 0 0.0
PAY_4 0 0.0
PAY_5 0 0.0
PAY_6 0 0.0
BILL_AMT1 0 0.0
BILL_AMT2 0 0.0
BILL_AMT3 0 0.0
BILL_AMT4 0 0.0
BILL_AMT5 0 0.0
BILL_AMT6 0 0.0
PAY_AMT1 0 0.0
PAY_AMT2 0 0.0
PAY_AMT3 0 0.0
PAY_AMT4 0 0.0
PAY_AMT5 0 0.0
PAY_AMT6 0 0.0
default payment next month 0 0.0
Imputación de valores faltantes con la mediana
df_imputed = df.copy()
for col in numeric_vars:
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
/tmp/ipykernel_124295/3441159290.py:3: FutureWarning: A value is trying to be set on a copy of a DataFrame or Series through chained assignment using an inplace method.
The behavior will change in pandas 3.0. This inplace method will never work because the intermediate object on which we are setting values always behaves as a copy.
For example, when doing 'df[col].method(value, inplace=True)', try using 'df.method({col: value}, inplace=True)' or df[col] = df[col].method(value) instead, to perform the operation inplace on the original object.
df_imputed[col].fillna(df_imputed[col].median(), inplace=True)
Normalización: Min-Max Scaling
scaler = MinMaxScaler()
df_minmax = pd.DataFrame(
scaler.fit_transform(df_imputed[numeric_vars]), columns=numeric_vars
)
df_minmax = pd.concat([df_imputed.drop(numeric_vars, axis=1), df_minmax], axis=1)
df_minmaxEstandarización: Z-score
scaler_std = StandardScaler()
df_zscore = pd.DataFrame(
scaler_std.fit_transform(df_imputed[numeric_vars]), columns=numeric_vars
)
df_zscoreDivision de datos¶
Considerar estos detalles:
-Tamaño del dataset: Muy pequeño (< 500 muestras) División recomendada: 90% train / 10% test o usar validación cruzada (cross-validation) Detalles: No conviene perder muchos datos para prueba; mejor usar técnicas como k-fold CV.
-Tamaño del dataset: Pequeño - Medio (500 – 2000 muestras) División recomendada: 80% train / 20% test Detalles: Buen balance entre entrenamiento y evaluación; común en muchos proyectos.
-Tamaño del dataset: Grande (2000 – 50,000 muestras) División recomendada: 70% train / 30% test Detalles: Proporción estándar en machine learning. Suficiente información para evaluar rendimiento.
-Tamaño del dataset: Muy grande (> 50,000 muestras) División recomendada: 60% train / 40% test o incluso 50% / 50% Detalles: Cuando el dataset es enorme, se puede reservar más datos para prueba sin perjudicar el entrenamiento.
-Para modelos finales / tuning fino División recomendada: 60% train / 20% val / 20% test Detalles: Se separa un conjunto de validación para ajustar hiperparámetros y otro test para la evaluación final.
Fuentes bibliográficas:
Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow de Aurélien Géron.
Deep Learning Specialization de Andrew Ng.
Scikit-learn documentation.
Kaggle Notebooks / Competitions.
Blogs y best practices en empresas (Google, Microsoft, Towards Data Science).
# Usá df_minmax o df_zscore según el procesamiento elegido:
X = df_minmax.drop(columns=["BILL_AMT1", "ID", "default payment next month"])
y = df_minmax["BILL_AMT1"]X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3, random_state=2025
)print("Tamaño del dataset:", len(df_minmax))
print("Tamaño X_train:", X_train.shape)
print("Tamaño X_test:", X_test.shape)
print("Tamaño y_train:", y_train.shape)
print("Tamaño y_test:", y_test.shape)Tamaño del dataset: 30000
Tamaño X_train: (21000, 22)
Tamaño X_test: (9000, 22)
Tamaño y_train: (21000,)
Tamaño y_test: (9000,)
Alternartivo: Modelo final con tuning de hiperparámetros.
Separar los datos mediante:
Entrenamiento (60%)
Validación (20%) → tuning de hiperparámetros.
Prueba (20%) → evaluación final
X_temp, X_test, y_temp, y_test = train_test_split(
X, y, test_size=0.2, random_state=2025
)
X_train, X_val, y_train, y_val = train_test_split(
X_temp, y_temp, test_size=0.25, random_state=2025
)Modelo Machine Learning de predicción: Regresion¶
# Crear y entrenar modelo
modelo = LinearRegression()
modelo.fit(X_train, y_train)# Predicciones
y_pred = modelo.predict(X_test)
y_predarray([0.14911765, 0.18466693, 0.15172128, ..., 0.16673364, 0.15332879,
0.16662126], shape=(6000,))Metricas de la Regresión
MSE(Mean Squared Error): Promedio del cuadrado del error.
RMSE (Raíz del MSE): Error típico esperado.
MAE (Mean Absolute Error): Promedio del valor absoluto del error.
R^2(coeficiente de determinación):Qué tanto explica el modelo la variabilidad de los datos.
mae = mean_absolute_error(y_test, y_pred)
rmse = np.sqrt(mean_squared_error(y_test, y_pred))
r2 = r2_score(y_test, y_pred)Resultados:
print("Predicciones:", y_pred[:10])Predicciones: [0.14911765 0.18466693 0.15172128 0.16102159 0.15097255 0.15282459
0.19158715 0.21399728 0.17877903 0.15421744]
print("Coeficientes:", modelo.coef_)Coeficientes: [ 0.01010368 -0.00033645 0.00301687 0.00089026 -0.00030341 0.00211393
0.01708278 -0.01513309 -0.00098225 -0.00438378 -0.00126067 0.9504712
0.02662981 -0.01000361 -0.00818543 -0.02109244 -0.59057455 0.19884261
0.12683068 0.07493774 0.05683012 0.02880422]
print(" MAE:", mean_absolute_error(y_test, y_pred))
print(" MSE:", mean_squared_error(y_test, y_pred))
print(" RMSE:", np.sqrt(mean_squared_error(y_test, y_pred)))
print(" R²:", r2_score(y_test, y_pred)) MAE: 0.007222868831554046
MSE: 0.0003529968890821301
RMSE: 0.018788211439147957
R²: 0.9172951049609116
print("--- Evaluación del Modelo de Regresión Lineal ---")
print(f"MAE (Error Absoluto Medio): {mae:.2f}")
print(f"RMSE (Raíz del Error Cuadrático Medio): {rmse:.2f}")
print(f"R^2 (Coeficiente de Determinación): {r2:.4f}")--- Evaluación del Modelo de Regresión Lineal ---
MAE (Error Absoluto Medio): 0.01
RMSE (Raíz del Error Cuadrático Medio): 0.02
R^2 (Coeficiente de Determinación): 0.9173
Gráfico
# Gráfico: valores reales vs. predichos
plt.figure(figsize=(8, 6))
plt.scatter(y_test, y_pred, alpha=0.5, color="blue")
plt.plot(
[y_test.min(), y_test.max()],
[y_test.min(), y_test.max()],
color="red",
linestyle="--",
)
plt.xlabel("Valores reales (BILL_AMT1)")
plt.ylabel("Valores predichos")
plt.title("Regresión Lineal: Valores Reales vs. Predichos")
plt.grid(True)
plt.tight_layout()
plt.show()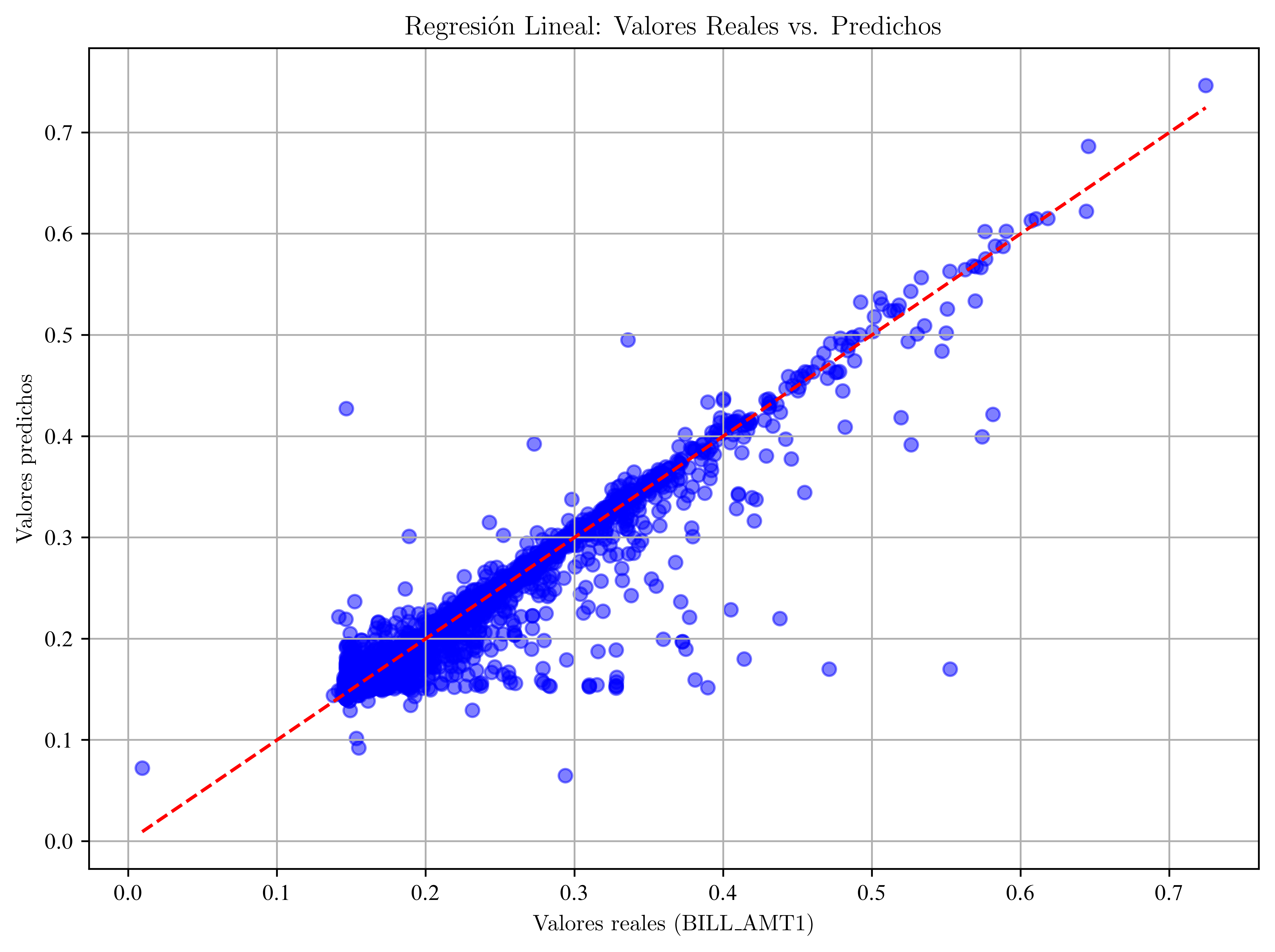
Modelo Machine Learning de predicción: Regresion con validación y búsqueda de hiperparámetros texto en negrita¶
from sklearn.linear_model import Ridge
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
from sklearn.model_selection import GridSearchCV, train_test_split# Separación 60% train / 20% val / 20% test
X_temp, X_test, y_temp, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
X_train, X_val, y_train, y_val = train_test_split(
X_temp, y_temp, test_size=0.25, random_state=42
)# Definir modelo y grid de hiperparámetros
ridge = Ridge()
param_grid = {"alpha": [0.01, 0.1, 1, 10, 100]}# Grid search usando validación cruzada dentro del set de entrenamiento
grid = GridSearchCV(
estimator=ridge, param_grid=param_grid, cv=5, scoring="neg_mean_squared_error"
)
grid.fit(X_train, y_train)# Mejor modelo
best_model = grid.best_estimator_
print("Mejor alpha encontrado:", grid.best_params_["alpha"])Mejor alpha encontrado: 0.1
# Evaluación en set de validación
y_val_pred = best_model.predict(X_val)
print("\n--- Validación ---")
print("MAE:", mean_absolute_error(y_val, y_val_pred))
print(
"RMSE:", np.sqrt(mean_squared_error(y_val, y_val_pred))
) # Calculate RMSE by taking the square root
print("R²:", r2_score(y_val, y_val_pred))
--- Validación ---
MAE: 0.006877855067917816
RMSE: 0.017155035191697938
R²: 0.926152870105252
# Evaluación en set de prueba (final)
y_test_pred = best_model.predict(X_test)
print("\n--- Prueba (Test Final) ---")
print("MAE:", mean_absolute_error(y_test, y_test_pred))
print("RMSE:", np.sqrt(mean_squared_error(y_test, y_test_pred)))
print("R²:", r2_score(y_test, y_test_pred))
--- Prueba (Test Final) ---
MAE: 0.007161325570049812
RMSE: 0.017668534680958907
R²: 0.9282777532542785
Gráfico:
plt.figure(figsize=(8, 6))
plt.scatter(y_test, y_pred, alpha=0.5, color="blue")
plt.plot([0, 1], [0, 1], "r--") # línea ideal: predicción = real
plt.xlabel("Valores reales")
plt.ylabel("Predicciones")
plt.title("Predicciones vs Valores Reales")
plt.grid(True)
plt.show()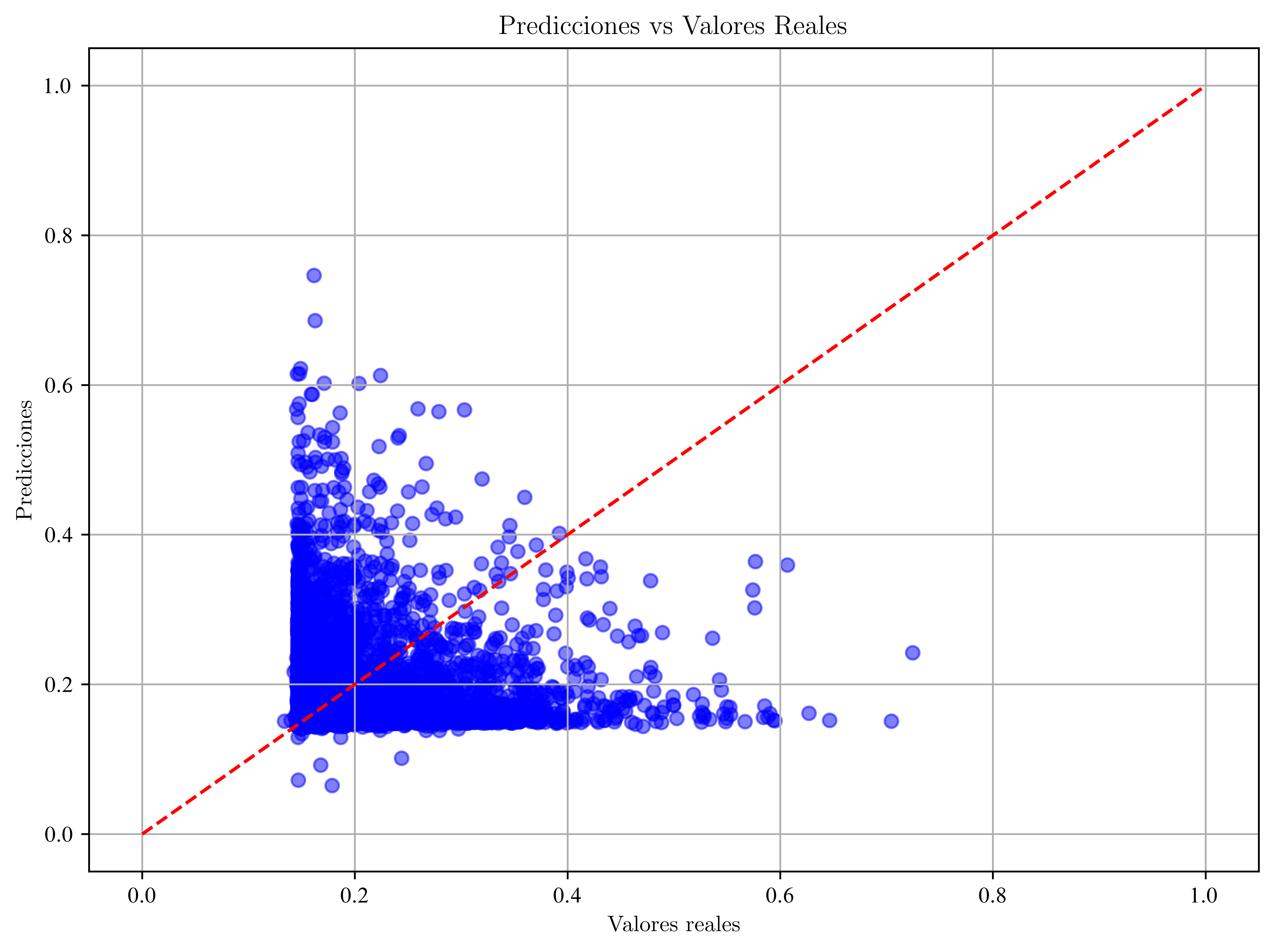
Distribución de Errores
errores = y_test - y_pred
plt.figure(figsize=(8, 6))
plt.hist(errores, bins=50, color="orange", edgecolor="black")
plt.title("Distribución de Errores de Predicción")
plt.xlabel("Error")
plt.ylabel("Frecuencia")
plt.grid(True)
plt.show()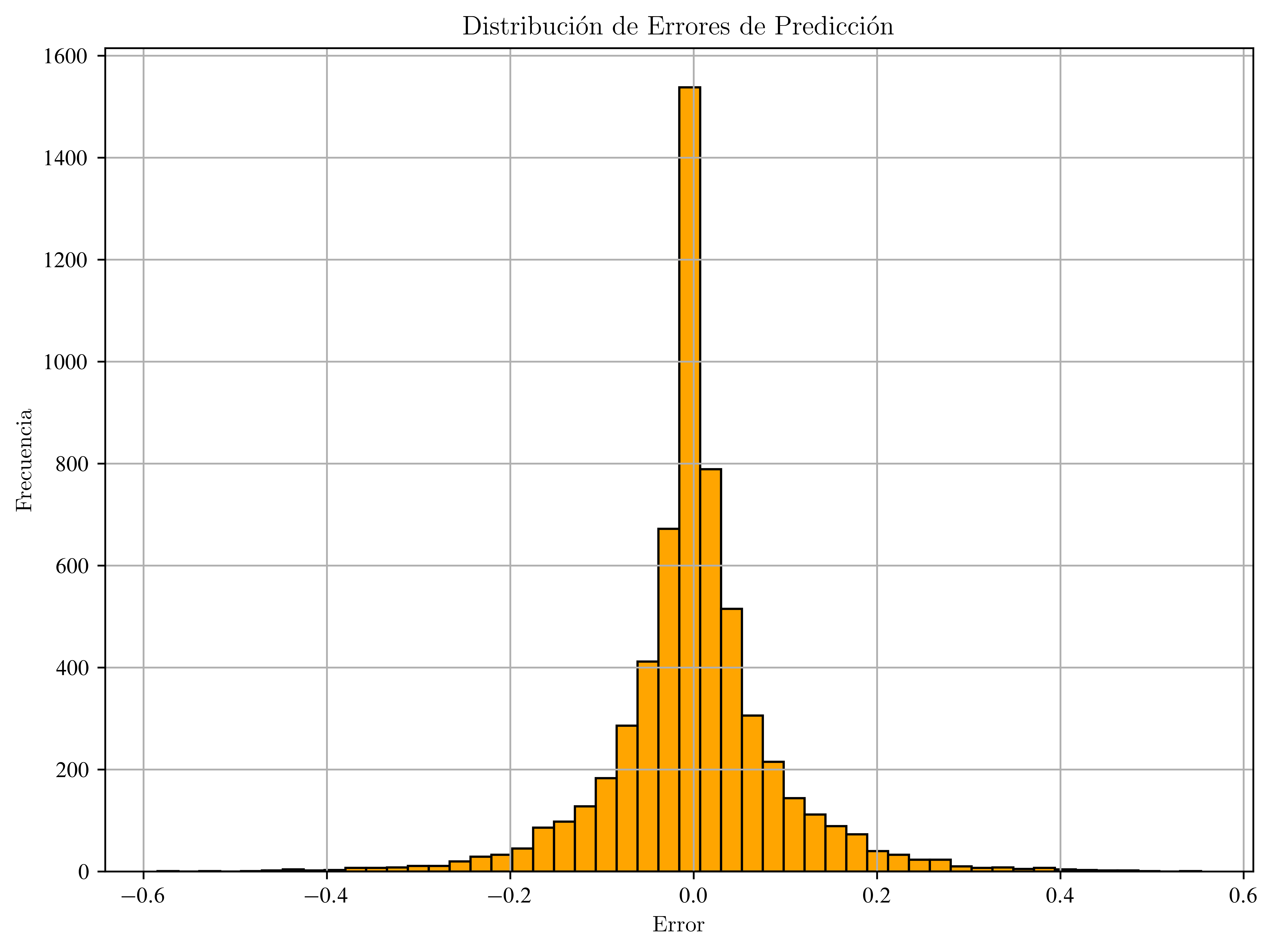
Modelo Estadístico Clásico de Regresión¶
import statsmodels.api as sm---------------------------------------------------------------------------
ImportError Traceback (most recent call last)
Cell In[46], line 1
----> 1 import statsmodels.api as sm
File /usr/lib/python3.13/site-packages/statsmodels/api.py:76
1 __all__ = [
2 "BayesGaussMI",
3 "BinomialBayesMixedGLM",
(...) 72 "__version_info__"
73 ]
---> 76 from . import datasets, distributions, iolib, regression, robust, tools
77 from .__init__ import test
78 from statsmodels._version import (
79 version as __version__, version_tuple as __version_info__
80 )
File /usr/lib/python3.13/site-packages/statsmodels/distributions/__init__.py:7
2 from .empirical_distribution import (
3 ECDF, ECDFDiscrete, monotone_fn_inverter, StepFunction
4 )
5 from .edgeworth import ExpandedNormal
----> 7 from .discrete import (
8 genpoisson_p, zipoisson, zigenpoisson, zinegbin,
9 )
11 __all__ = [
12 'ECDF',
13 'ECDFDiscrete',
(...) 21 'zipoisson'
22 ]
24 test = PytestTester()
File /usr/lib/python3.13/site-packages/statsmodels/distributions/discrete.py:5
3 from scipy.stats import rv_discrete, poisson, nbinom
4 from scipy.special import gammaln
----> 5 from scipy._lib._util import _lazywhere
7 from statsmodels.base.model import GenericLikelihoodModel
10 class genpoisson_p_gen(rv_discrete):
ImportError: cannot import name '_lazywhere' from 'scipy._lib._util' (/usr/lib/python3.13/site-packages/scipy/_lib/_util.py)# Agregar constante (intercepto) manualmente
X_train_sm = sm.add_constant(X_train)
X_test_sm = sm.add_constant(X_test)# Ajustar modelo OLS (Ordinary Least Squares)
modelo_ols = sm.OLS(y_train, X_train_sm).fit()
modelo_ols.summary()Correlación de variables numéricas:
correlation_matrix = X_train.corr()
plt.figure(figsize=(12, 10))
sbn.heatmap(correlation_matrix, annot=False, cmap="coolwarm")
plt.title("Matriz de Correlación entre Variables Predictoras")
plt.show()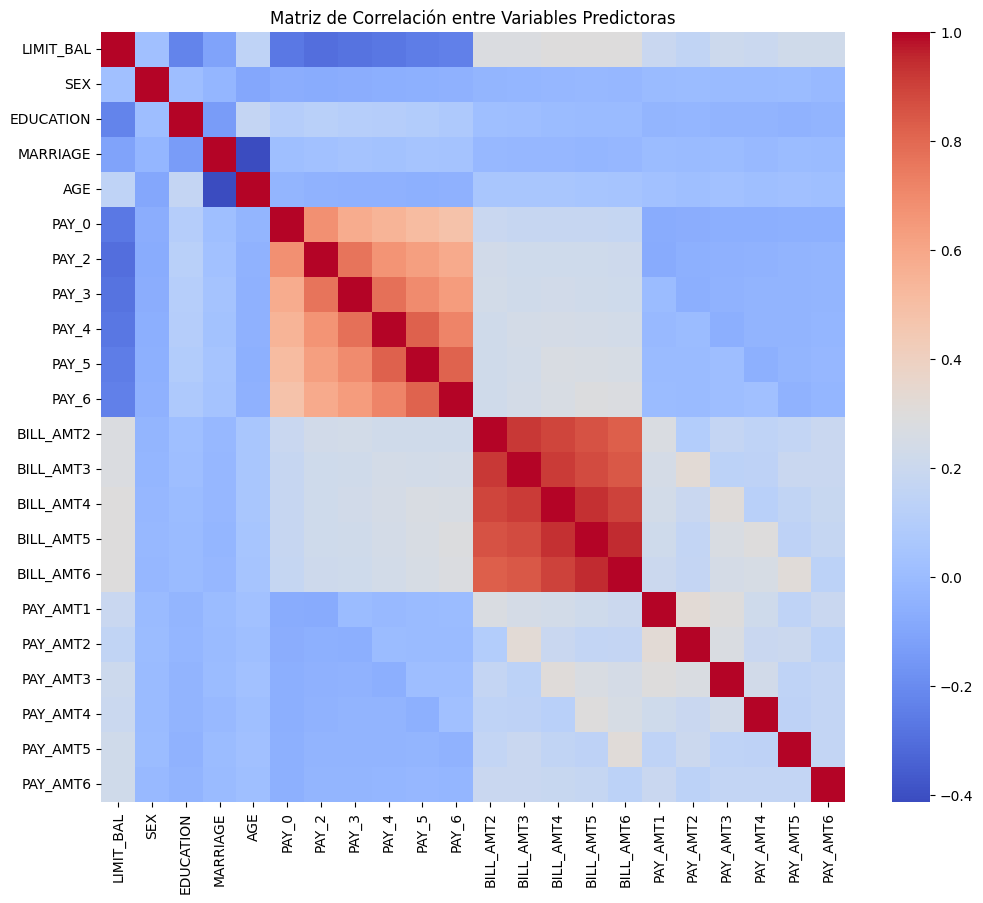
Matriz de correlación de las variables finales
plt.figure(figsize=(20, 10))
sbn.heatmap(
X_train_sm.drop(columns="const").corr(), annot=True, cmap="coolwarm", fmt=".2f"
)
plt.title("Matriz de Correlación - Variables Seleccionadas")
plt.show()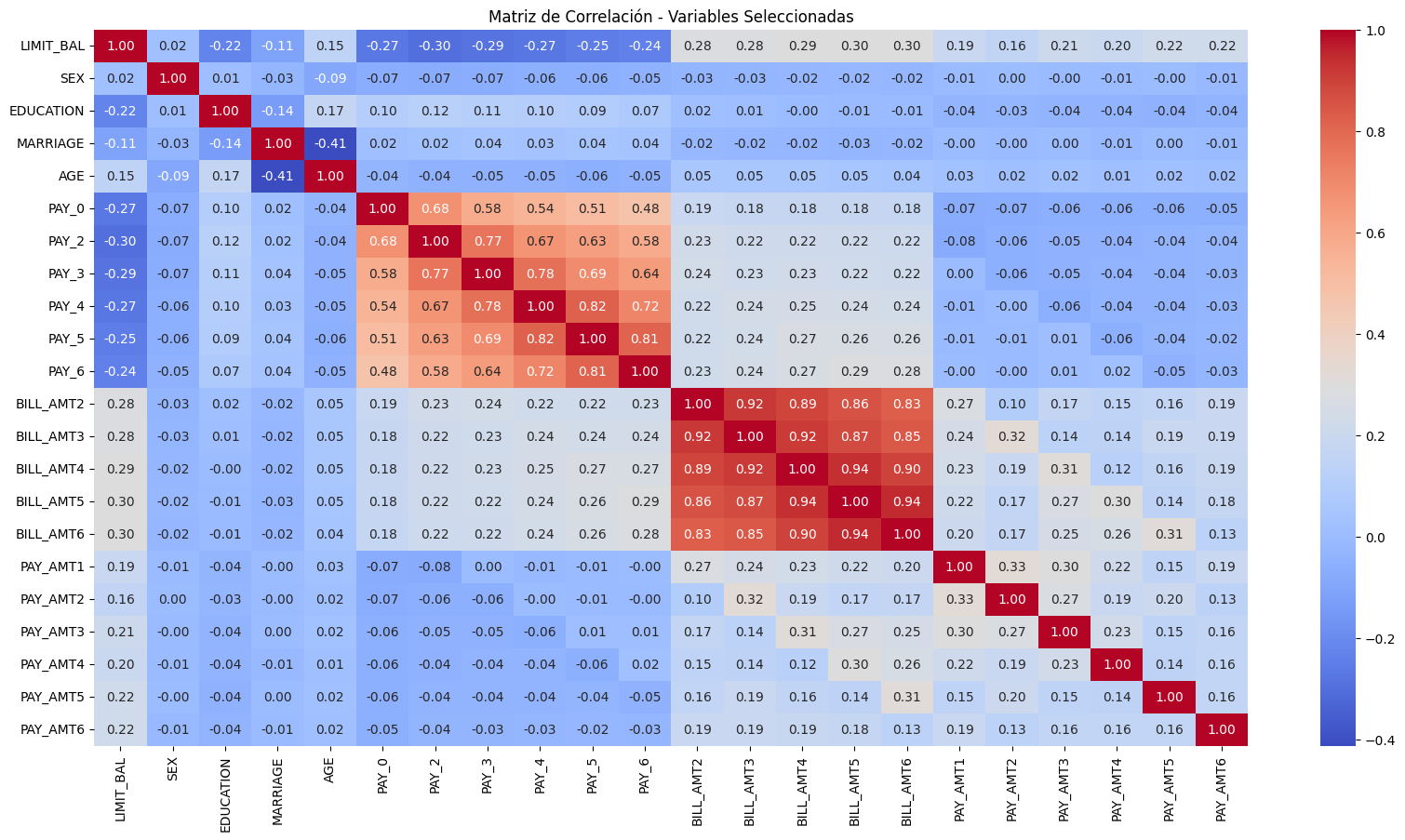
Selección de variables
import statsmodels.api as smX_opt = sm.add_constant(X_train.copy())
modelo = sm.OLS(y_train, X_opt).fit()# Iterativa: elimina variables con p > 0.05
while max(modelo.pvalues) > 0.05:
worst_pval_var = modelo.pvalues.idxmax()
print(
f"Eliminando '{worst_pval_var}' con p-valor = {modelo.pvalues[worst_pval_var]:.4f}"
)
X_opt = X_opt.drop(columns=worst_pval_var)
modelo = sm.OLS(y_train, X_opt).fit()
# Resultado final
print("\n--- Modelo final tras selección ---")
print(modelo.summary())Eliminando 'PAY_5' con p-valor = 0.9512
Eliminando 'BILL_AMT3' con p-valor = 0.8683
Eliminando 'AGE' con p-valor = 0.4280
Eliminando 'PAY_4' con p-valor = 0.3297
Eliminando 'PAY_0' con p-valor = 0.1200
Eliminando 'PAY_6' con p-valor = 0.1031
Eliminando 'MARRIAGE' con p-valor = 0.0617
--- Modelo final tras selección ---
OLS Regression Results
==============================================================================
Dep. Variable: BILL_AMT1 R-squared: 0.929
Model: OLS Adj. R-squared: 0.929
Method: Least Squares F-statistic: 1.573e+04
Date: Sun, 20 Jul 2025 Prob (F-statistic): 0.00
Time: 20:24:04 Log-Likelihood: 47336.
No. Observations: 18000 AIC: -9.464e+04
Df Residuals: 17984 BIC: -9.451e+04
Df Model: 15
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
const 0.0872 0.002 36.873 0.000 0.083 0.092
LIMIT_BAL 0.0121 0.001 10.111 0.000 0.010 0.014
SEX -0.0006 0.000 -2.375 0.018 -0.001 -0.000
EDUCATION 0.0042 0.001 4.150 0.000 0.002 0.006
PAY_2 0.0180 0.002 10.344 0.000 0.015 0.021
PAY_3 -0.0188 0.002 -10.817 0.000 -0.022 -0.015
BILL_AMT2 0.9528 0.005 202.725 0.000 0.944 0.962
BILL_AMT4 0.0295 0.009 3.318 0.001 0.012 0.047
BILL_AMT5 -0.0359 0.011 -3.354 0.001 -0.057 -0.015
BILL_AMT6 -0.0211 0.011 -1.970 0.049 -0.042 -0.000
PAY_AMT1 -0.5119 0.007 -68.611 0.000 -0.527 -0.497
PAY_AMT2 0.1023 0.009 10.827 0.000 0.084 0.121
PAY_AMT3 0.0710 0.007 9.660 0.000 0.057 0.085
PAY_AMT4 0.0940 0.006 15.249 0.000 0.082 0.106
PAY_AMT5 0.0275 0.004 6.134 0.000 0.019 0.036
PAY_AMT6 0.0326 0.004 7.749 0.000 0.024 0.041
==============================================================================
Omnibus: 22330.031 Durbin-Watson: 1.978
Prob(Omnibus): 0.000 Jarque-Bera (JB): 7771210.696
Skew: 6.438 Prob(JB): 0.00
Kurtosis: 103.974 Cond. No. 154.
==============================================================================
Notes:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.
VIF
Considerar:
VIF > 5 → posible colinealidad moderada.
VIF > 10 → colinealidad severa, considerar eliminar.
import pandas as pd
from statsmodels.stats.outliers_influence import variance_inflation_factor# Calcular VIF solo para variables numéricas finales
vif_data = pd.DataFrame()
vif_data["Variable"] = X_opt.columns
vif_data["VIF"] = [
variance_inflation_factor(X_opt.values, i) for i in range(X_opt.shape[1])
]
print("\n--- VIF (Multicolinealidad) ---")
print(vif_data)
--- VIF (Multicolinealidad) ---
Variable VIF
0 const 330.838699
1 LIMIT_BAL 1.474082
2 SEX 1.007121
3 EDUCATION 1.062764
4 PAY_2 2.585007
5 PAY_3 2.546360
6 BILL_AMT2 6.031519
7 BILL_AMT4 17.210318
8 BILL_AMT5 25.168125
9 BILL_AMT6 14.518973
10 PAY_AMT1 1.347421
11 PAY_AMT2 1.265380
12 PAY_AMT3 1.412560
13 PAY_AMT4 1.738691
14 PAY_AMT5 1.675559
15 PAY_AMT6 1.157711
Consideraciones Extras¶
Regularización (Ridge y Lasso)¶
Para evitar sobreajuste o cuando hay muchas variables correlacionadas.
from sklearn.linear_model import Lasso, Ridge# Ridge Regression (L2)
modelo_ridge = Ridge(alpha=1.0).fit(X_train, y_train)
modelo_ridge.score(X_test, y_test)0.927044875037085# Lasso Regression (L1)
modelo_lasso = Lasso(alpha=1.0).fit(X_train, y_train)
modelo_lasso.score(X_test, y_test)-0.00030956403466819715from sklearn.metrics import mean_squared_error, r2_score# Modelo Ridge:
y_pred_ridge = modelo_ridge.predict(X_test)
print("Ridge - R²:", r2_score(y_test, y_pred_ridge))
print("Ridge - MSE:", mean_squared_error(y_test, y_pred_ridge))Ridge - R²: 0.927044875037085
Ridge - MSE: 0.0003175433240174616
# Modelo Lasso:
y_pred_lasso = modelo_lasso.predict(X_test)
print("Lasso - R²:", r2_score(y_test, y_pred_lasso))
print("Lasso - MSE:", mean_squared_error(y_test, y_pred_lasso))Lasso - R²: -0.00030956403466819715
Lasso - MSE: 0.0043539316007133394
import matplotlib.pyplot as plt
import pandas as pdcoef_ridge = pd.Series(
modelo_ridge.coef_,
index=df.drop(["ID", "default payment next month", "BILL_AMT1"], axis=1).columns,
)
coef_lasso = pd.Series(
modelo_lasso.coef_,
index=df.drop(["ID", "default payment next month", "BILL_AMT1"], axis=1).columns,
)
plt.figure(figsize=(10, 6))
coef_ridge.plot(label="Ridge", alpha=0.7)
coef_lasso.plot(label="Lasso", alpha=0.7)
plt.legend()
plt.title("Comparación de Coeficientes: Ridge vs Lasso")
plt.grid(True)
plt.tight_layout()
plt.show()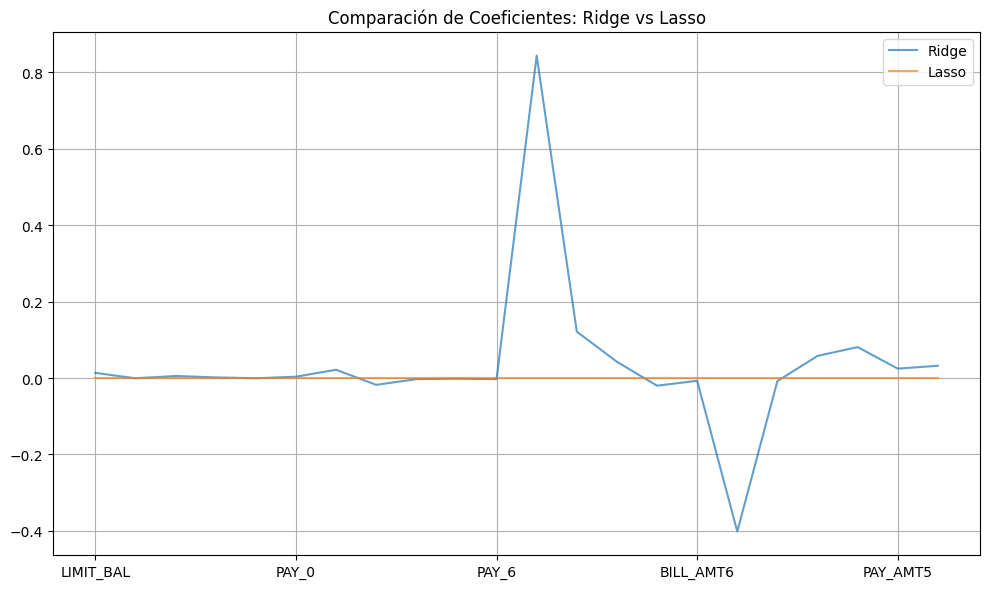
Cross-Validation¶
Para estimar el rendimiento del modelo con mayor robustez:
from sklearn.model_selection import cross_val_scorescores = cross_val_score(best_model, X, y, cv=5, scoring="r2")
print("R² promedio (CV):", scores.mean())R² promedio (CV): 0.9276842702376229
Análisis de residuos¶
Ver cómo se comportan los errores del modelo:
residuos = y_test - y_pred
plt.hist(residuos, bins=50)
plt.title("Distribución de residuos")
plt.xlabel("Error")
plt.ylabel("Frecuencia")
plt.grid(True)
plt.show()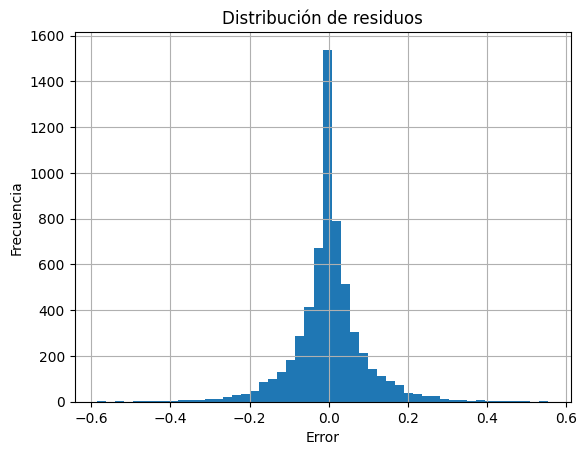
import scipy.stats as stats
import seaborn as snsNormalidad
# Histograma y Q-Q plot
sns.histplot(residuos, kde=True)
plt.title("Distribución de residuos")
plt.show()
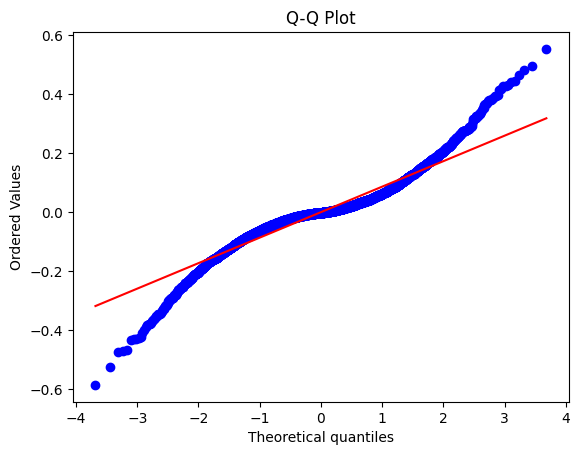
# Q-Q plot
stats.probplot(residuos, dist="norm", plot=plt)
plt.title("Q-Q Plot")
plt.show()
# Test de Shapiro-Wilk
shapiro_test = stats.shapiro(residuos)
print("Shapiro-Wilk p-value:", shapiro_test.pvalue)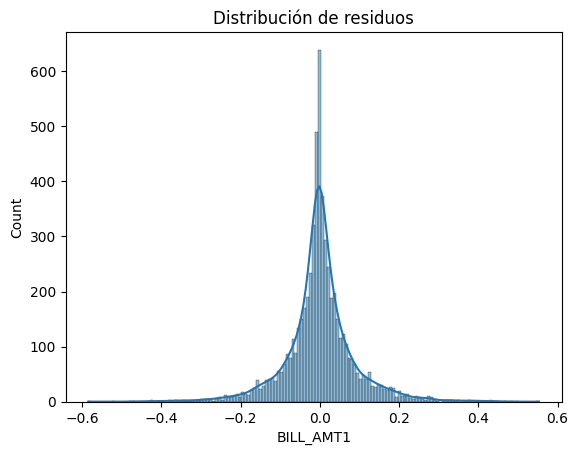
Shapiro-Wilk p-value: 1.198771088903701e-49
/usr/local/lib/python3.11/dist-packages/scipy/stats/_axis_nan_policy.py:586: UserWarning: scipy.stats.shapiro: For N > 5000, computed p-value may not be accurate. Current N is 6000.
res = hypotest_fun_out(*samples, **kwds)
Homocedasticidad
plt.scatter(y_pred, residuos)
plt.axhline(0, color="red", linestyle="--")
plt.xlabel("Valores predichos")
plt.ylabel("Residuos")
plt.title("Residuos vs Predicción")
plt.grid(True)
plt.show()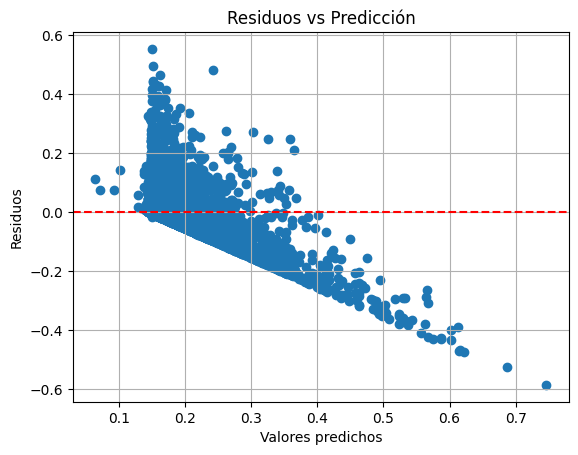
import statsmodels.api as sm
import statsmodels.stats.api as smsX_const = sm.add_constant(X_train)
modelo_ols = sm.OLS(y_train, X_const).fit()
bp_test = sms.het_breuschpagan(modelo_ols.resid, X_const)
print("Breusch-Pagan p-value:", bp_test[1])Breusch-Pagan p-value: 2.359134149746979e-243
Independencia de errores - Autocorrelación
El valor de Durbin-Watson oscila entre 0 y 4:
a) Aprox a 2 → errores no autocorrelacionados (ideal)
b) < 2 : autocorrelación positiva
c) > 2 : autocorrelación negativa
El test Durbin-Watson está diseñado para interpretación numérica y comparativa
from statsmodels.stats.stattools import durbin_watsondw = durbin_watson(residuos)
print("Durbin-Watson:", dw)Durbin-Watson: 2.0020479671683638
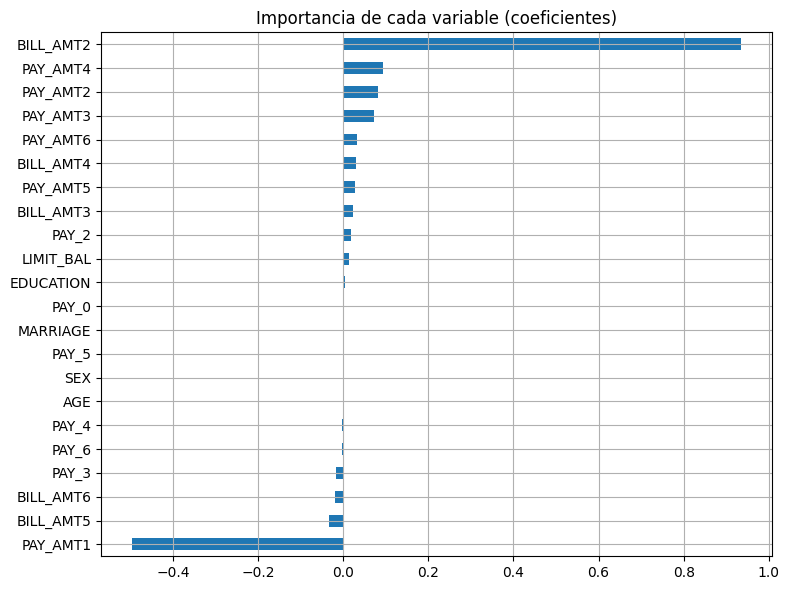
Importancia de variables¶
Interpretar cuáles variables influyen más
coef = pd.Series(best_model.coef_, index=X.columns)
coef.sort_values().plot(kind="barh", figsize=(8, 6))
plt.title("Importancia de cada variable (coeficientes)")
plt.grid(True)
plt.tight_layout()
plt.show()