In this notebook, we will talk about algorithms for computing Eigenvalues and Eigenvectors.
Preparation¶
Import modules
import time
import matplotlib.pyplot as plt
import numpy as np
import scipy.linalg as scl
import scipy.sparse as scspCreate random matrices¶
At first, we introduce a short notation to generate random matrices.
The dimension is a tuple (rows, cols). If rows=cols a short notation is possible such that:
(n, n) = (n) = n
def get_dimension(dim):
# converts short notion (only 1 int or 1 tuple for two times same index) into 2-tuple
# INPUT : 2-tuple, 1-tuple or int
# OUTPUT: 2-tuple
if str(type(dim)) == "<class 'tuple'>":
try:
n = dim[1]
except:
n = dim[0]
m = dim[0]
else:
m = dim
n = dim
return (m, n)
# Test function:
print(get_dimension((2, 3)), "should be: (2, 3)")
print(get_dimension(2), "should be: (2, 2)")
print(get_dimension((2)), "should be: (2, 2)")(2, 3) should be: (2, 3)
(2, 2) should be: (2, 2)
(2, 2) should be: (2, 2)
def Create_random_matrix(dim, data_type="real"):
# Returns a real or complex matrix with uniform or wide range distribution
# INPUT :- dim: number or 1-tupel or 2- tuple
# - data_type: "real or complex"
# - distribution "uniform" in [0,1) or "wide_range" in (-10^10, 10^10), approximated log distributed
# OUTPUT: Matrix of dimension dim with real or complex entries
dim = get_dimension(dim)
A = np.random.random(dim)
if data_type == "complex":
A = A + 1j * np.random.random()
if data_type == "symmetric":
A = A + A.T
if data_type == "diag_dominant":
row_sum = np.sum(A, axis=1)
A = A + A.T
for i in range(dim[0]):
A[i, i] = row_sum[i] + 0.5
return A
# test function
A = Create_random_matrix(2, "complex")
print(A)
B = Create_random_matrix((2, 1))
print(B)[[0.68696769+0.30565857j 0.37045767+0.30565857j]
[0.7581725 +0.30565857j 0.43638565+0.30565857j]]
[[0.35649781]
[0.96256572]]
Motivation¶
This can be skipped. In this example we would like to find the eigen-frequencies of a membrane on a square.
For this, we discretize the Laplace operator via a finite difference approach and determine the eigenvalues of this discretization.
The Laplace equation is
with
We assume on
Setup Matrix¶
# Some index manipulations, used later
def index_2d_to_1d(i, j, n):
"""
Convert 2D index to 1D index
Input:
i is the row index
j is the column index
n is the number of columns
Output:
1D index
"""
return i * n + j
def index_1d_to_2d(index, n):
"""
Convert 1D index to 2D index
Input:
index is the 1D index
n is the number of columns
Output:
2D index
"""
return (index // n, index % n)Just execute the next cells, it’s not important what happens inside.
def laplace_matrix_square(n):
"""
returns the Laplace operator discretized on a grid of n*n points
Input:
n is the number of points in each dimension
Output:
A is the Laplace matrix
"""
A = np.zeros((n * n, n * n), dtype=float)
for i in range(n):
for j in range(n):
# inside the domain
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j, n)] = -4.0
if i > 0 and j > 0 and i < n - 1 and j < n - 1:
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i - 1, j, n)] = 1.0
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i + 1, j, n)] = 1.0
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j - 1, n)] = 1.0
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j + 1, n)] = 1.0
continue
## on the boundary
# top boundary
if i == 0:
if j == 0: # top left corner
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i + 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j + 1, n)] = 1
elif j == n - 1: # top right corner
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i + 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j - 1, n)] = 1
else:
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i + 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j - 1, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j + 1, n)] = 1
continue
# bottom boundary
if i == n - 1:
if j == 0: # bottom left corner
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i - 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j + 1, n)] = 1
continue
elif j == n - 1: # bottom right corner
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i - 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j - 1, n)] = 1
continue
else:
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i - 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j - 1, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j + 1, n)] = 1
continue
# left boundary
if j == 0:
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i - 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i + 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j + 1, n)] = 1
# right boundary
if j == n - 1:
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i - 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i + 1, j, n)] = 1
A[index_2d_to_1d(i, j, n), index_2d_to_1d(i, j - 1, n)] = 1
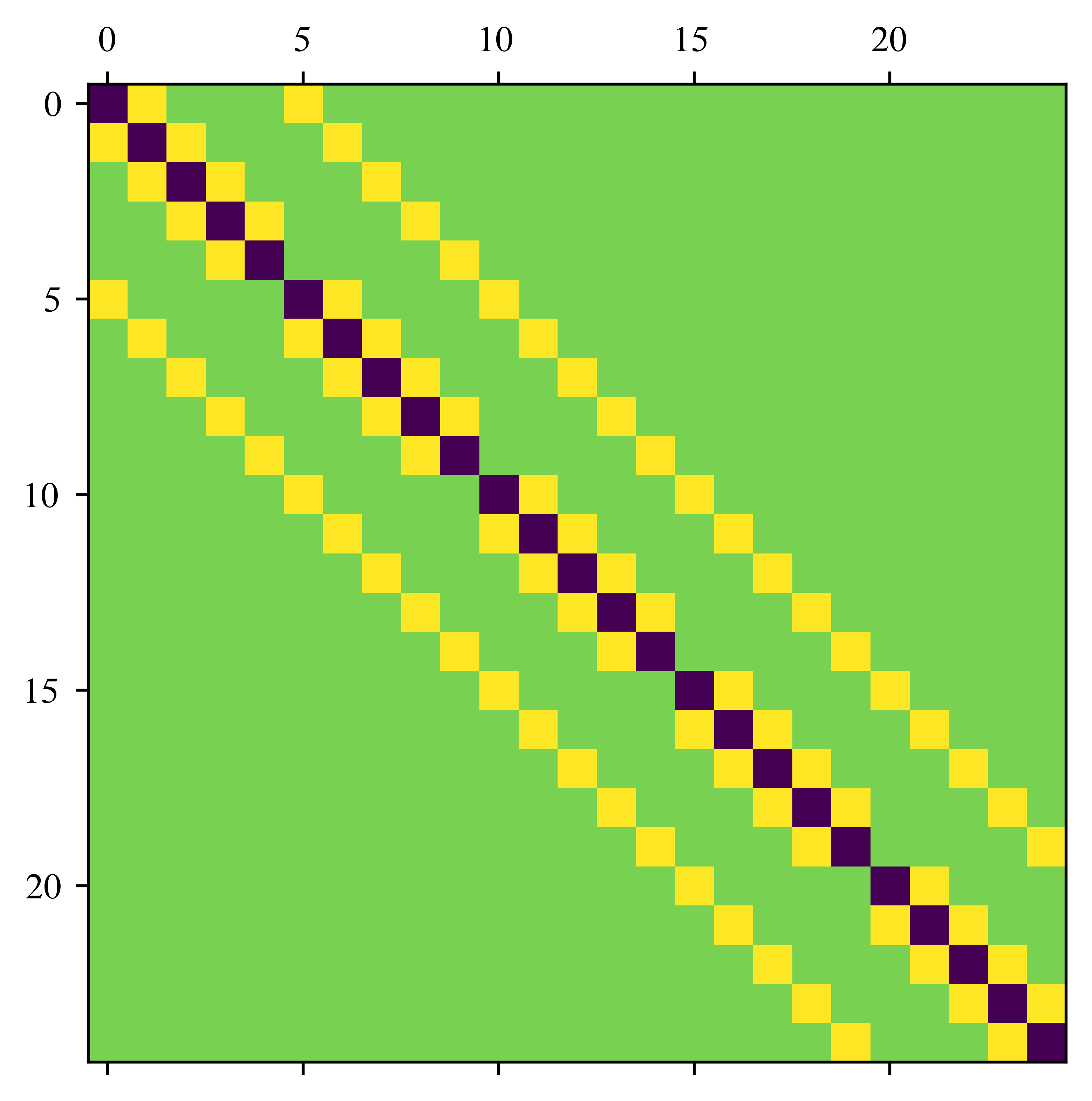
return A / n**2#### test
n = 5
A = laplace_matrix_square(n)
plt.matshow(A)/usr/lib/python3.13/site-packages/IPython/core/events.py:82: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
func(*args, **kwargs)
/usr/lib/python3.13/site-packages/IPython/core/pylabtools.py:170: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
fig.canvas.print_figure(bytes_io, **kw)
def plot_eigenfunctions(A, n_max=6):
plot_row = n_max // 3 + np.min([n_max % 3, 1])
print(plot_row)
fig, ax = plt.subplots(plot_row, 3, figsize=(15, 5 * plot_row))
n = int(np.sqrt(A.shape[0]))
for i in range(n_max):
B = np.zeros((n, n))
for j in range(n**2):
coords = index_1d_to_2d(j, n)
B[coords[0], coords[1]] = A[j, i]
# plot
row = i // 3
col = i % 3
im = ax[row, col].matshow(B)
ax[row, col].set_title("Eigenfunction " + str(i + 1))
fig.colorbar(im, ax=ax[row, col])
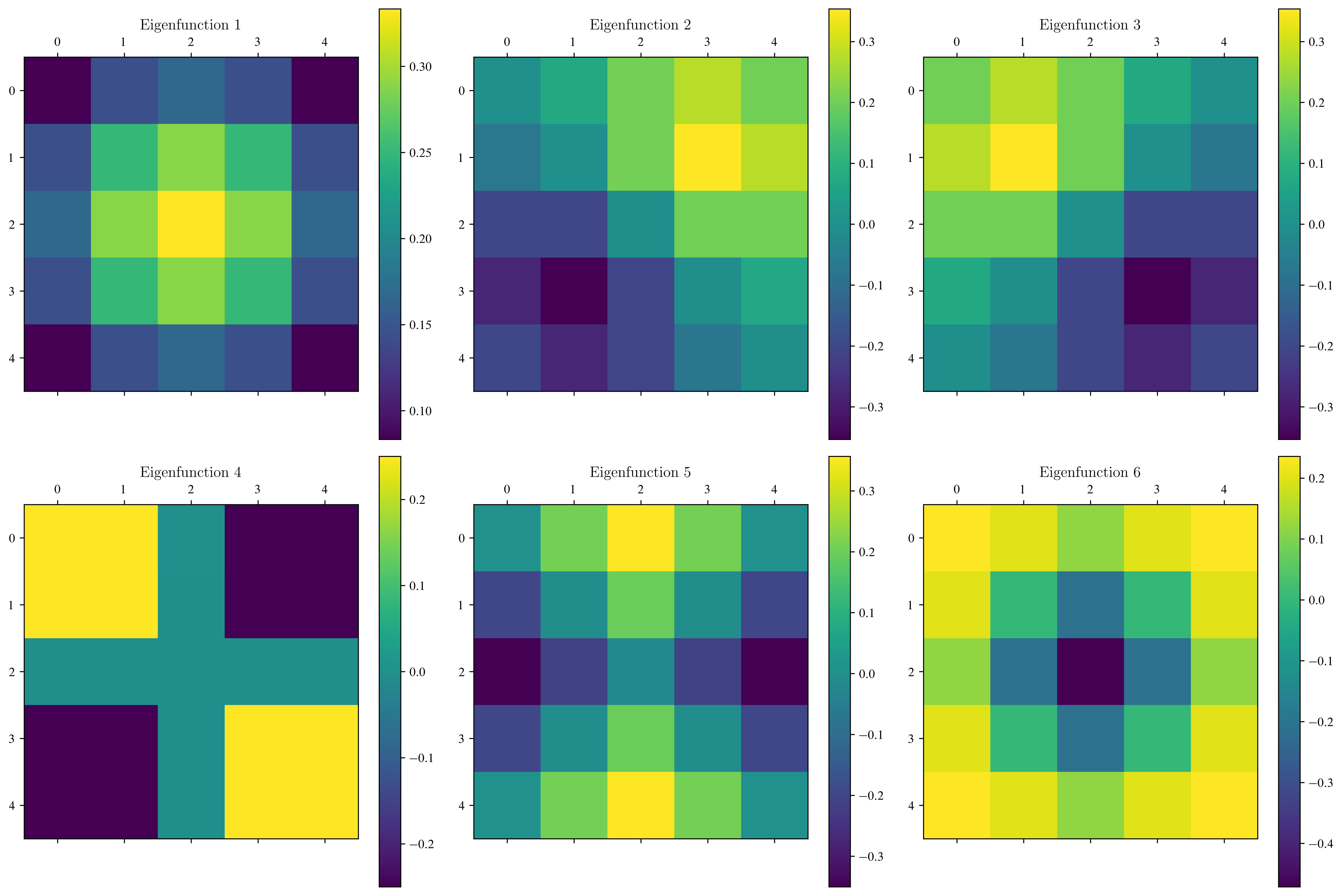
plt.show()n = 5
A = laplace_matrix_square(n)
eigenvalues, eigenvectors = np.linalg.eig(A)
mask = np.argsort(eigenvalues)
eigenvalues = eigenvalues[mask[::-1]]
eigenvectors = eigenvectors[:, mask[::-1]]
print("Eigenvalues: \n", np.real(eigenvalues))
plot_eigenfunctions(eigenvectors)Eigenvalues:
[-0.02143594 -0.05071797 -0.05071797 -0.08 -0.09071797 -0.09071797
-0.12 -0.12 -0.13071797 -0.13071797 -0.16 -0.16
-0.16 -0.16 -0.16 -0.18928203 -0.18928203 -0.2
-0.2 -0.22928203 -0.22928203 -0.24 -0.26928203 -0.26928203
-0.29856406]
2
Warning!
Please save all progress that you’ve made so far. Do save your results stored in other programs too.
The next cell is computationally expensive, so your computer could crash. After you saved everything, you can change test_capacity to True.
test_capacity = True
n = 4
times = []
while test_capacity:
time_start = time.time()
A = laplace_matrix_square(n)
eigenvalues, eigenvectors = scl.eig(A)
time_end = time.time()
times.append(time_end - time_start)
if (time_end - time_start) > 10:
test_capacity = False
n = n * 2
n_Max = n // 2
print("What??? Only " + str(n_Max))What??? Only 64
sizes = np.arange(2, len(times) + 2, 1)
plt.title("Time to calculate eigenvalues")
plt.plot(2 ** (sizes), times)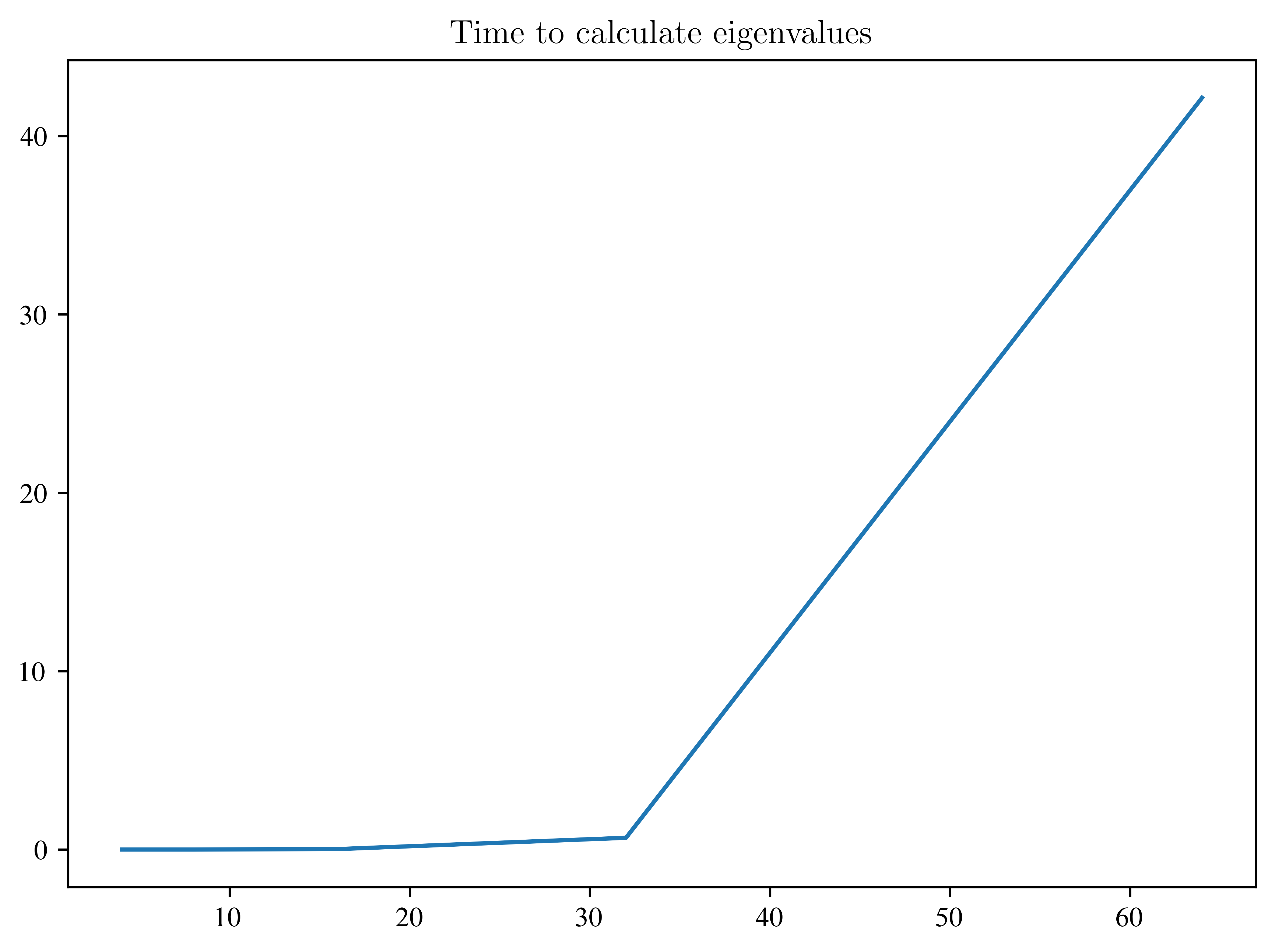
Let’s beat this!
Hessenberg form¶
Recall: A matrix can be transformed to Hessenberg form via similarity transformations. In the case of a symmetric matrix, the resulting matrix is tridiagonal. One kind of transformation is the Householder reflection. To apply this, we define the Householder matrices. (See lessons on Monday, too)
Householder matrix¶
def Householder_matrix(v):
# Create Matrix Q_v as described above.
# INPUT: v 1d-array or 2d-array with one column or one row
# OUPUT: Q Householdermatrix of v
v = v.copy() # do not overwrite original matrix
n = len(v)
v = v.reshape(
(n)
) # reshape, because tensorprod gives different results for (n) and (n,1) dimension
if np.linalg.norm(v) == 0:
return np.eye(n)
# modify first component
if v[0] == 0:
sign = 1
else:
sign = v[0] / np.abs(v[0])
v[0] += sign * np.linalg.norm(v) # linalg.norm calculates the l2 norm of a vector
v = v / np.linalg.norm(v)
# calc Q
Q = np.eye(n) - 2.0 * np.tensordot(
v, np.conj(v), axes=0
) # tensorprod does w w^T multiplication
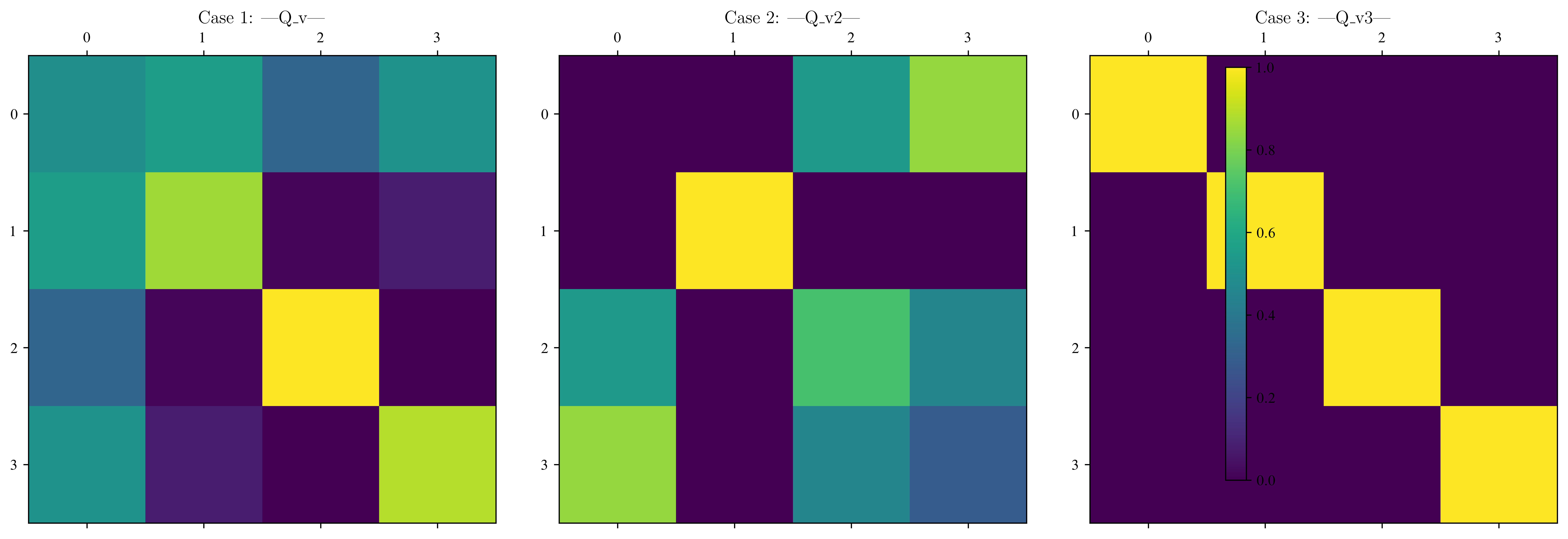
return Qfig, ax = plt.subplots(1, 3, figsize=(15, 5))
# test case 1
print("test case 1")
v = Create_random_matrix((4, 1))
# print("v=\n", v)
Q = Householder_matrix(v)
im = ax[0].matshow(np.abs(Q))
ax[0].set_title("Case 1: |Q_v| ")
# fig.colorbar(im, ax=ax[0])
# check errors
error_sym = np.max(np.abs(Q - np.conj(Q.T)))
error_id = np.max(np.abs(Q @ Q - np.eye(v.shape[0])))
error_refl = np.max(np.abs((Q @ v)[1:]))
print("maximal error: |Q-Q^*| = \t", error_sym)
print("maximal error: |QQ-id|=\t", error_id)
print("zero after reflection |Qv|=\t", error_refl)
assert error_sym < 1e-10
assert error_id < 1e-10
assert error_refl < 1e-10
# test case 2
print("\ntest case 2")
v = Create_random_matrix((4, 1))
v[0:2] = np.array([0, 0]).reshape((2, 1))
# print("\n\n\nv_2=\n", v)
Q = Householder_matrix(v)
im = ax[1].matshow(np.abs(Q))
ax[1].set_title("Case 2: |Q_v2|")
# fig.colorbar(im, ax=ax[1])
print("maximal error: |Q-Q^*| = \t", np.max(np.abs(Q - np.conj(Q.T))))
print("maximal error |QQ-id|=\t", np.max(np.abs(Q @ Q - np.eye(v.shape[0]))))
print("2nd column ofQ under diagonal: \t", Q[2:, 1], "should be zero")
assert np.max(np.abs(Q[2:, 1])) <= 1e-10
# test case 3
print("\ntest case 3")
v = np.zeros(4)
Q = Householder_matrix(v)
im = ax[2].matshow(np.abs(Q))
ax[2].set_title("Case 3: |Q_v3|")
fig.colorbar(im, ax=ax)
# print("\n\nv_3= \n", v)
print("maximal error Q-id=\t", np.max(np.abs(Q - np.eye(4))))test case 1
maximal error: |Q-Q^*| = 0.0
maximal error: |QQ-id|= 5.551115123125783e-17
zero after reflection |Qv|= 5.551115123125783e-17
test case 2
maximal error: |Q-Q^*| = 0.0
maximal error |QQ-id|= 3.885780586188048e-16
2nd column ofQ under diagonal: [0. 0.] should be zero
test case 3
maximal error Q-id= 0.0
/usr/lib/python3.13/site-packages/IPython/core/events.py:82: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
func(*args, **kwargs)
Hessenberg function¶
def Hessenberg_form(A):
# This function calculates the Hessenberg form via Housholder transformations of a complex matrix.
# The entries of the inputmatrix will be overwritten by the Hessenbergform.
# INPUT : real/complex squared Matrix A
# OUTPUT: transfomation matrix
n = A.shape[0]
# store full transformation in matrix
Q_res = np.eye(n, dtype=A.dtype)
for i in range(n - 2):
Q = Householder_matrix(A[i + 1 :, i])
A[i + 1 :, :] = Q @ A[i + 1 :, :]
A[:, i + 1 :] = A[:, i + 1 :] @ Q # Q = Q* for householder matrices
Q_res[i + 1 :, :] = Q @ Q_res[i + 1 :, :]
return np.conj(Q_res.T)fig, ax = plt.subplots(1, 2, figsize=(10, 5))
A = Create_random_matrix(5)
ax[0].matshow(A)
ax[0].set_title("A before")
B = A.copy()
Q = Hessenberg_form(A)
im = ax[1].matshow(A)
ax[1].set_title("A in Hessenbergform")
fig.colorbar(im, ax=ax)
plt.show()
print("maximal error of Q*AQ-A_h\t", np.max(np.abs(np.conj(Q.T) @ B @ Q - A)))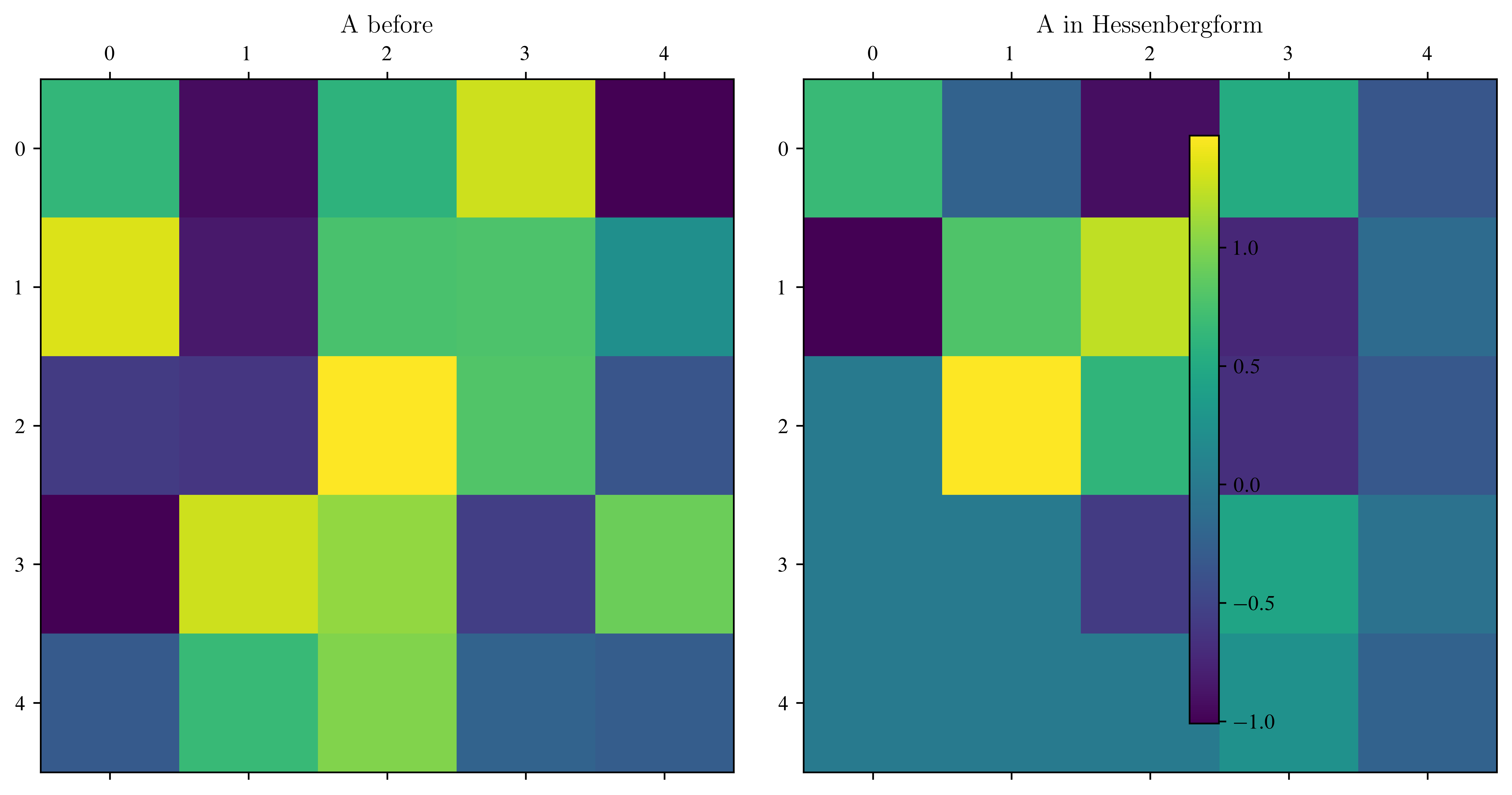
maximal error of Q*AQ-A_h 2.8098456932654787e-16
n = 5
A = laplace_matrix_square(n)
Hessenberg_form(A)
plt.matshow(A)
plt.title("A in Hessenbergform")
plt.colorbar()
plt.show()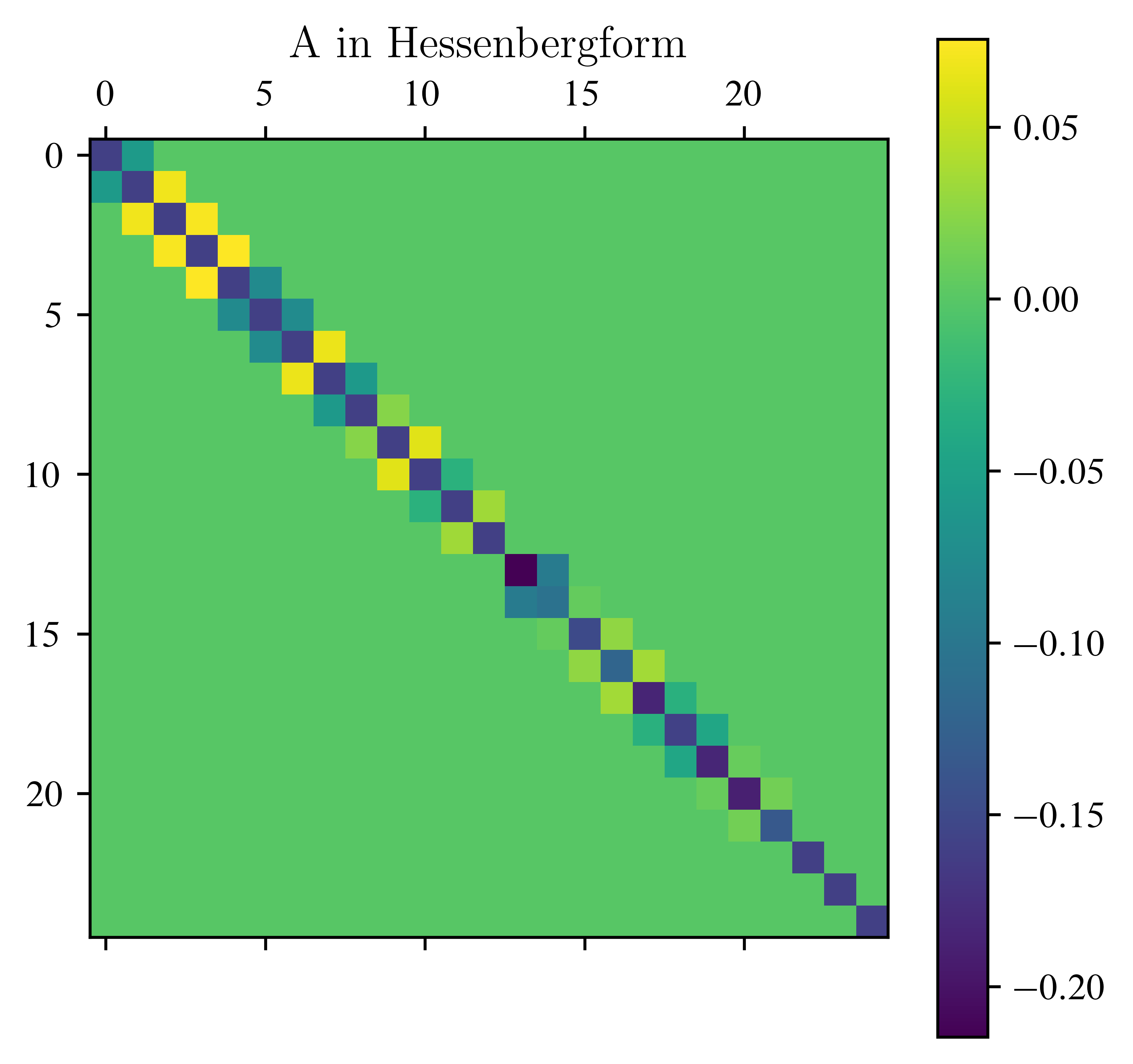
Now we have a good matrix to apply the QR-method. But first, we need the Givens rotation to efficiently calculate a QR decomposition.
QR decomposition¶
Givens rotations¶
Theory:
Due to readability, we omit the second index of the matrix for the rest of the calculation. It is always
such that
(source: inspired by “On Computing Givens Rotations Reliably and Efficiently” from DAVID BINDEL, JAMES DEMMEL, WILLIAM KAHAN and OSNI MARQUES, https://
modified eq 1, because in the written way it gives wrong results)
def Givens_rotation(v):
# This function calculates a rotation matrix,
# such that the lower component of a 2-dimensional vector is 0
# This is the complex version
# INPUT : 2-dimensional vector
# OUTPUT: 2x2-matrix
v = v.copy() # not overwrite the matrix elements
v = v.reshape((2)) # such that entries are callable
a = v[0]
b = v[1]
r = np.sqrt(a * np.conj(a) + b * np.conj(b))
Q = np.array([[np.conj(a), np.conj(b)], [-b, a]]) / r
return Qv = Create_random_matrix((2, 1))
print("v=\n", v)
Q = Givens_rotation(v)
print("Q=\n", Q)
print("Qv = \n", Q @ v)v=
[[0.25286711]
[0.25297524]]
Q=
[[ 0.70695561 0.70725792]
[-0.70725792 0.70695561]]
Qv =
[[0.35768456]
[0. ]]
The next milestone to a QR-iteration is the QR-decomposition of a Hessenberg matrix.
def QR_decomposition_Hess(A):
# This function calculates the QR decomposition of a matrix in Hessenberg form and store the values of R in A
# INPUT : A, nxn matrix in Hessenberg form
# OUTPUT: Q*, nxn matrix, orthogonal
n = A.shape[0]
Q_res = np.eye(n, dtype=A.dtype)
for i in range(n - 1):
Q = Givens_rotation(A[i : i + 2, i])
A[i : i + 2, i:] = Q @ A[i : i + 2, i:]
Q_res[i : i + 2, :] = Q @ Q_res[i : i + 2, :]
return np.conj(Q_res.T)fig, ax = plt.subplots(1, 3, figsize=(15, 5))
A = laplace_matrix_square(5)
Q_1 = Hessenberg_form(A)
B = A.copy()
# original matrix
ax[0].matshow(np.abs(A))
ax[0].set_title("A in Hessenberg form")
Q_2 = QR_decomposition_Hess(A)
# Q
ax[1].matshow(np.abs(Q_2))
ax[1].set_title("Q")
# R
im = ax[2].matshow(np.abs(A))
ax[2].set_title("R")
fig.colorbar(im, ax=ax)
error_qr = np.max(np.abs(Q_2 @ A - B))
print("maximal error |QR-A|: \t", error_qr)
assert error_qr < 1e-10maximal error |QR-A|: 5.551115123125783e-17
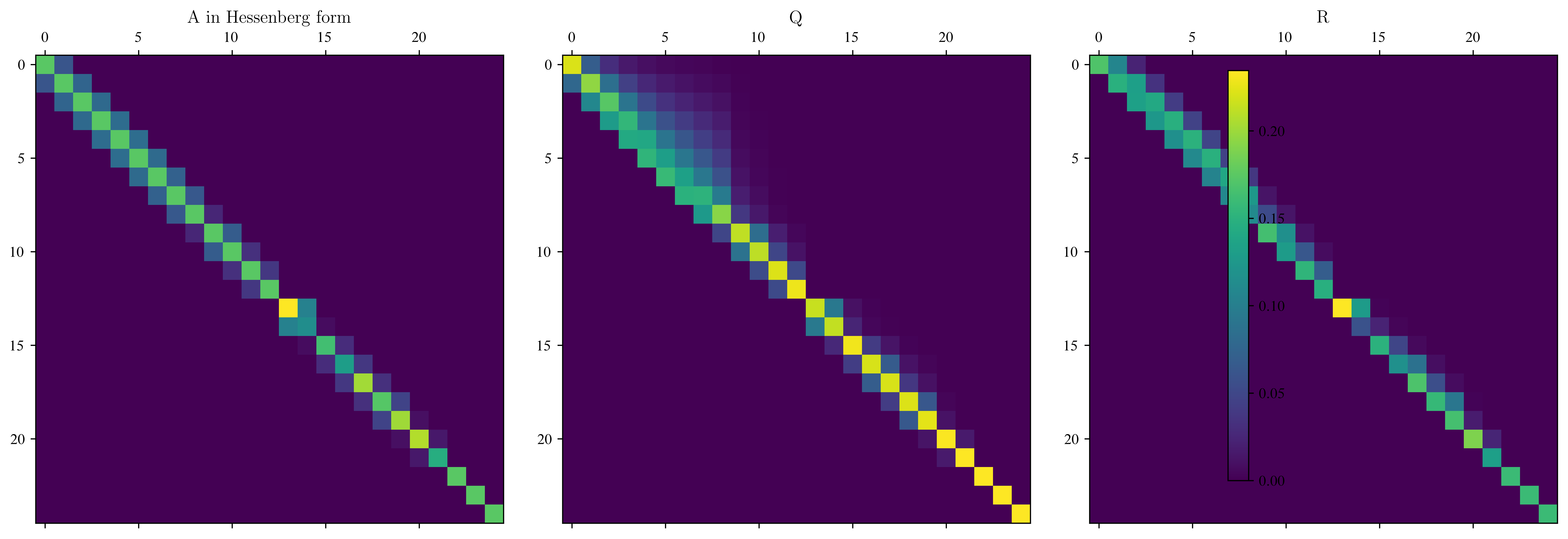
Now we have all parts together to do the QR-iteration. It will be a bit slow, but it will work.
QR Iteration¶
(QR decomposition)
Define
(QR decomposition) and define
...
This scheme is called QR iteration.
def QR_iteration_slow(A, eps=1e-6, max_Iterations=10000):
# This function iterates the basic QR algorithm, until either the entries on the first sub diagonal
# are small enough or Max iter reached The matrix A will be overwritten.
# INPUT : - A: squared Matrix in hessenberg form
# - eps: if no diag entry changes more than eps, than the algorithm is converged
# - max_iterations: If this number is reached, it aborts the iteration and gives a warning
# OUTPUT: - Q, nxn Matrix, similarity transformation
# - Iter : number of Iterations
# define values, for the computation
n = A.shape[0]
Iter = 0
error_ev = 1
diagvals = np.diag(A).copy()
Q_res = np.eye(n)
# start QR iteration
while error_ev > eps and Iter < max_Iterations:
Q_2 = QR_decomposition_Hess(A)
A[:, :] = A @ Q_2
Q_res[:, :] = Q_res @ Q_2
Iter += 1
# preparation for next iteration
error_ev = np.max(np.abs(diagvals - np.diag(A)))
diagvals = np.diag(A).copy()
if Iter == max_Iterations:
print("Warning: Algorithm may not converged.")
return Q_res, IterA = laplace_matrix_square(5)
Q_Hess = Hessenberg_form(A)
B = A.copy()
Q, Iter = QR_iteration_slow(A, max_Iterations=10000)
plt.matshow(A)
plt.title("A in Schur form")
plt.colorbar()
plt.show()
w, v = np.linalg.eig(A)
print("Iterations:\t\t\t", Iter)
print("eigenvalues\t\t", np.diag(A))
print("true eigenvalues:\t", np.sort(w))
error_eigenval = np.max(np.abs(np.sort(w) - np.sort(np.diag(A))))
print("Max error of eigenvalues:\t", error_eigenval)
assert error_eigenval < 1e-4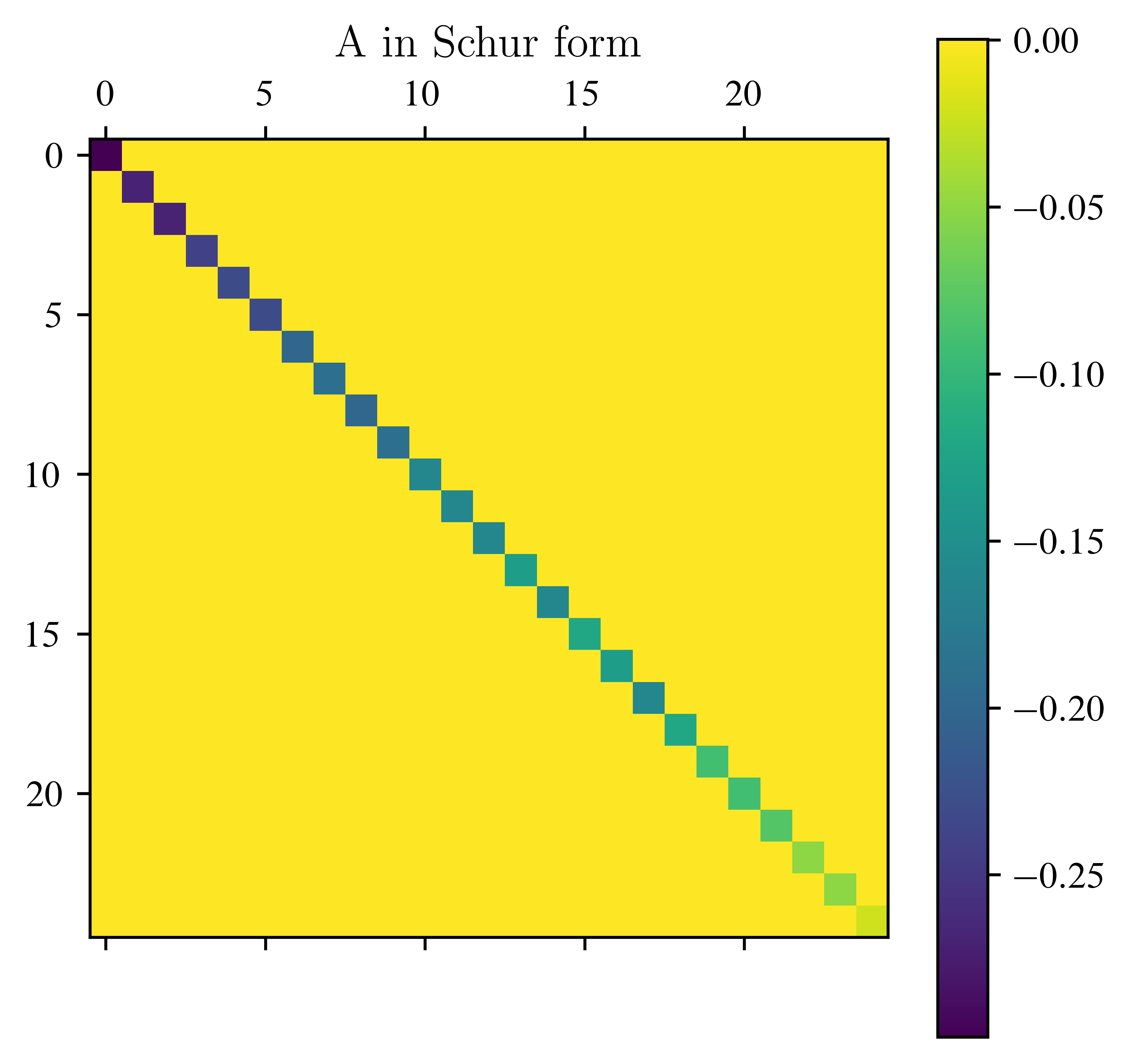
Iterations: 395
eigenvalues [-0.29856406 -0.26928203 -0.26928203 -0.24 -0.22928203 -0.22928203
-0.2 -0.18928203 -0.2 -0.18928203 -0.16 -0.16
-0.15999997 -0.130718 -0.15999902 -0.12000124 -0.13071771 -0.15999999
-0.12000001 -0.09071797 -0.09071797 -0.08 -0.05071797 -0.05071797
-0.02143594]
true eigenvalues: [-0.29856406 -0.26928203 -0.26928203 -0.24 -0.22928203 -0.22928203
-0.2 -0.2 -0.18928203 -0.18928203 -0.16 -0.16
-0.16 -0.16 -0.16 -0.13071797 -0.13071797 -0.12
-0.12 -0.09071797 -0.09071797 -0.08 -0.05071797 -0.05071797
-0.02143594]
Max error of eigenvalues: 1.2412057576044466e-06
Speed test¶
Warning: Again be cautious. The Cell is a runtime test.
test_capacity = True
n = 4
times = []
while test_capacity:
print(n)
time_start = time.time()
A = laplace_matrix_square(n)
Hessenberg_form(A)
Q, iter = QR_iteration_slow(A)
time_end = time.time()
times.append(time_end - time_start)
if (time_end - time_start) > 10:
test_capacity = False
n = n * 2
n_Max_selfwritten = n // 2
print(
"Our self written code is just " + str(n_Max // n_Max_selfwritten) + "times slower."
)4
8
16
Our self written code is just 4times slower.
This is not bad! We have still some space for improvement
Quality of Convergence¶
Maybe, we can learn something from “How the Eigenvalues converge”. For this, we plot diagonal entries over time. Ignore all the warnings.
# prepare experiment:
n = 8
A = Create_random_matrix(n, "diag_dominant")
# for comparison:
w, v = np.linalg.eig(A.copy())
w = np.sort(w)
Q_Hess = Hessenberg_form(A)
max_Iterations = 20
convergence_eigenvals = {} # dict
B = A.copy()
improvement_diag = np.zeros((n, max_Iterations))
for j in range(max_Iterations):
Q, Iter = QR_iteration_slow(B, max_Iterations=2)
improvement_diag[:, j] = np.sort(np.diag(B.copy()))
# plot over iterations
fig, ax = plt.subplots()
fig.suptitle("Convergence of eigenvalues")
ax.plot(np.arange(max_Iterations), improvement_diag.T)
ax.hlines(w, xmin=0, xmax=max_Iterations, colors="k", linestyles="dashed")
ax.set_xlabel("iterations")
ax.set_ylabel("eigenvalues")
# ax.set_xscale("log")
plt.show()Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
Warning: Algorithm may not converged.
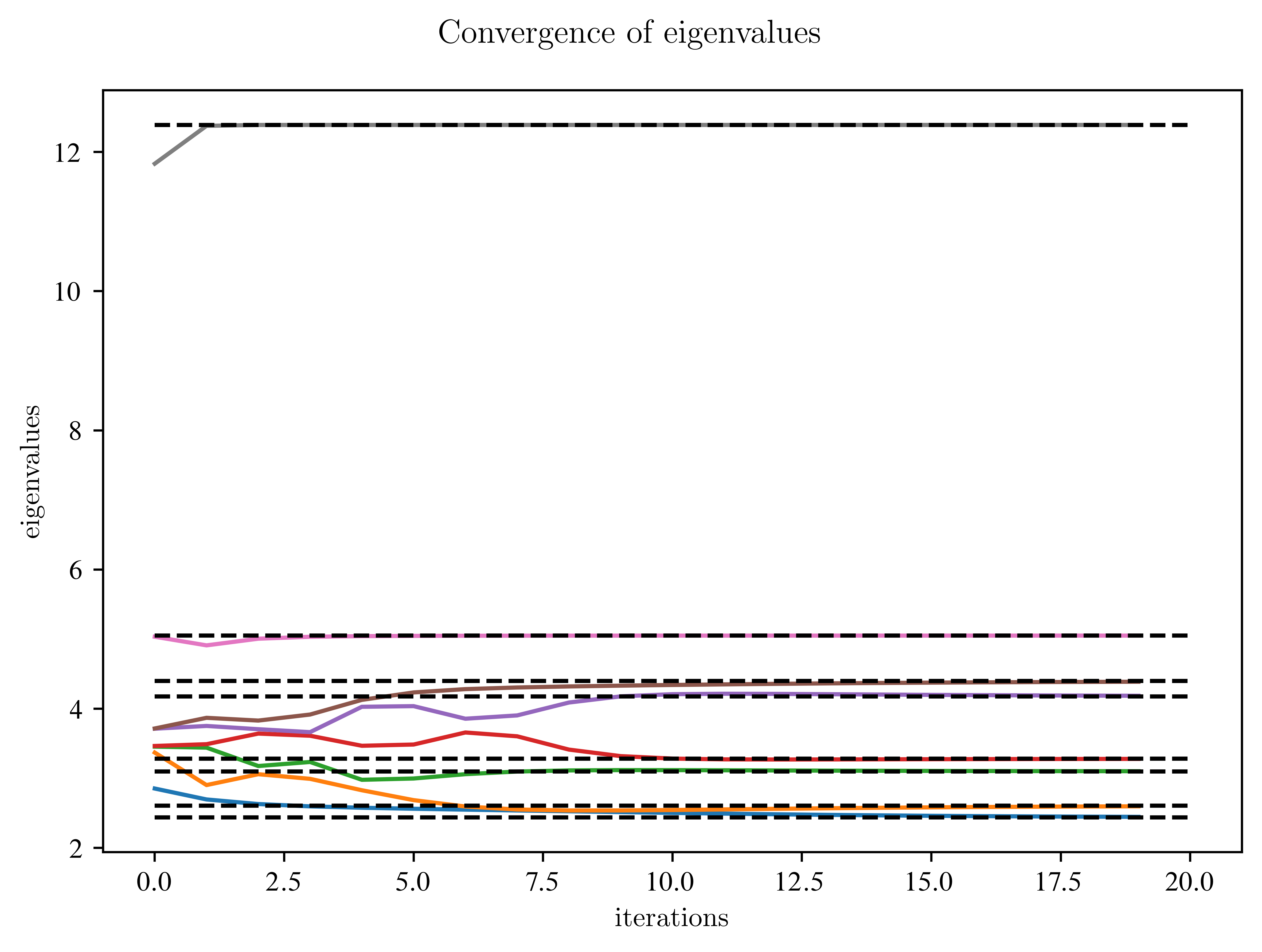
What do you observe?
Which eigenvalues converge the fastest?
Which need more time?
Is there a relation between the distance to the top and bottom of the figure?
Solution:¶
The largest and smallest eigenvalues converge faster than the neighbouring ones.
On Optimization¶
But we aim for more efficiency! We realize:
In lecture this morning, Celine told something about shifts? Can we use them?
Deflation...? Was there something?
Shifts¶
The convergence rate of the QR method is
This ratio can be very small. If we guess , we can speed up the convergence by shifting
But how do we get a good guess for ? Different people found different answers:
Rayleigh:
Wilkinson: , i.e. the smallest eigenvalue of lower right 2x2 Matrix. This can be calculated analytically.
Further shifts are possible, including multi-shifts.
Zero Shift: No shift at all.
Note: It’s totally fine to just implement the Rayleigh shift.
def evaluate_shift(A, shift_type="Zero"):
# This function returns different types of shifts.
# INPUT : A: submatrix of size 2x2
# : Type of shift(Zero, Rayleigh, Wilkinson(?))
# OUTPUT: a number which approximates an eigenvalue.
if shift_type == "Zero":
return 0
if shift_type == "Rayleigh":
return A[1, 1]
if shift_type == "Wilkinson":
# pq formula, see Schur form
p_2 = (A[0, 0] + A[1, 1]) / 2
q = A[0, 0] * A[1, 1] - A[0, 1] * A[1, 0]
lambda_1 = p_2 + np.sqrt(p_2 * p_2 - q + 0j)
lambda_2 = p_2 - np.sqrt(p_2 * p_2 - q + 0j)
if np.abs(A[1, 1] - lambda_1) < np.abs(A[1, 1] - lambda_2):
return lambda_1
else:
return lambda_2
else:
print("Warning: No valid shift type, no shift used")
return 0Next, test your shifts. Do they behave as expected?
Solution¶
The shifts behave as expected, if they are close to eigenvalues of 2x2 Matrix. At least the real part.
A = Create_random_matrix(2)
print(A)
sigma1 = evaluate_shift(A)
print("No shift:\n sigma =", sigma1)
sigma2 = evaluate_shift(A, "Rayleigh")
print("Raiyleigh shift:\n sigma=", sigma2)
sigma3 = evaluate_shift(A, "Wilkinson")
print("Wilkonson shift:\n sigma=", sigma3)
w, v = np.linalg.eig(A)
print("compare to eigenvalues:", w)
print("No valid input:\n")
sigma5 = evaluate_shift(A, "blabla")[[0.86789127 0.43466432]
[0.1479929 0.88211951]]
No shift:
sigma = 0
Raiyleigh shift:
sigma= 0.882119510150282
Wilkonson shift:
sigma= (1.128733284603029+0j)
compare to eigenvalues: [0.62127749 1.12873328]
No valid input:
Warning: No valid shift type, no shift used
Let us test our first improvement. For this, we have to modify the QR algorithm.
def QR_iteration_shift(
A, eps=1e-6, max_Iterations=10000, shift="Zero", suppress_warning=False
):
# This function iterates the QR algorithm using shifts and a first glance of deflation.
# I titerates, until either Max iter reached or
# the values on the diagonal are
# converged. The matrix A will be overwritten.
# INPUT : - A: squared Matrix in hessenberg form
# - eps: if no diag entry changes more than eps, than the algorithm is converged
# - max_iterations: If this number is reached, it aborts the iteration and gives a warning
# - shift: accelerate convergence, types: Zero, Wilkinson, Francis, Rayleigh, None,
# if None Wilkinson or Francis will be selected
# OUTPUT: - Q, nxn Matrix, similarity transformation
# - Iter : number of Iterations
# define values, for the computation
n = A.shape[0]
Iter = 0
error_ev = 1
diagvals = np.diag(A).copy()
Q_res = np.eye(n, dtype=A.dtype)
# the next two values may be used to accelerate the computations
# (based on observations made before)
min_pos_not_converged = 0
max_pos_not_converged = n
# start QR iteration
while error_ev > eps and Iter < max_Iterations:
# print(Iter, min_pos_not_converged, max_pos_not_converged)
# evaluate shifts
l = min_pos_not_converged
m = max_pos_not_converged # for readability
sigma = evaluate_shift(A[m - 2 : m, m - 2 : m], shift)
# QR iteration including shifts
A[l:m, l:m] = A[l:m, l:m] - sigma * np.eye(m - l, dtype=A.dtype)
Q_2 = QR_decomposition_Hess(A[l:m, l:m])
A[l:m, l:m] = A[l:m, l:m] @ Q_2 + sigma * np.eye(m - l, dtype=A.dtype)
Q_res[l:m, l:m] = Q_res[l:m, l:m] @ Q_2
Iter += 1
# preparation for next iteration
error_ev = np.max(np.abs(diagvals - np.diag(A)))
indices_not_converged = np.where(np.abs(diagvals - np.diag(A)) > eps)[0]
if len(indices_not_converged) == 0:
break
min_pos_not_converged = indices_not_converged[0]
max_pos_not_converged = indices_not_converged[-1] + 1
diagvals = np.diag(A).copy()
if not suppress_warning and Iter == max_Iterations:
print("Warning: Algorithm may not converged.")
return Q_res, Itern = 5
A = Create_random_matrix(n, "diag_dominant")
Q_Hess = Hessenberg_form(A)
w, v = np.linalg.eig(A)
B = A.copy()
Q, Iter = QR_iteration_shift(A, max_Iterations=10000, shift="Rayleigh")
eigenvalues = np.diag(A)
mask = np.argsort(eigenvalues)
eigenvalues = eigenvalues[mask[::-1]]
eigenvectors = eigenvectors[:, mask[::-1]]
plt.matshow(A)
plt.title("check result visually")
plt.colorbar()
plt.show()
print("Iterations:\t\t\t", Iter)
print("true eigenvalues:\t", w)
print("eigenvalues\t\t", eigenvalues)
error_eigenval = np.max(np.abs(np.sort(w) - np.sort(eigenvalues)))
print("Max error of eigenvalues:\t", error_eigenval)
assert error_eigenval < 1e-4/usr/lib/python3.13/site-packages/IPython/core/pylabtools.py:170: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
fig.canvas.print_figure(bytes_io, **kw)
Iterations: 9
true eigenvalues: [7.76051061 3.4018155 2.29909318 1.68528222 1.11668061]
eigenvalues [7.76051061 3.40181551 2.29909318 1.68528222 1.11668061]
Max error of eigenvalues: 7.423446746201989e-09
Test different shifts:
# prepare experiment:
n = 15
A = Create_random_matrix(n, "diag_dominant")
# for comparison:
w, v = np.linalg.eig(A.copy())
w = np.sort(w)
Q_Hess = Hessenberg_form(A)
list_shifts = ["Zero", "Rayleigh", "Wilkinson"]
print("shifts", " iterartions", " error eigenvalues")
for shift in list_shifts:
B = A.copy()
Q, Iter = QR_iteration_shift(B, max_Iterations=10000, shift=shift)
eigenvalues = np.sort(np.diag(B))
error_eigenval = np.max(np.abs(w - eigenvalues))
print(shift, Iter, error_eigenval)shifts iterartions error eigenvalues
Zero 218 1.972414224482577e-05
Rayleigh 33 1.5847918053424337e-06
Wilkinson 27 2.1925407835965416e-06
/tmp/ipykernel_122996/153652883.py:36: ComplexWarning: Casting complex values to real discards the imaginary part
A[l:m, l:m] = A[l:m, l:m] - sigma * np.eye(m - l, dtype=A.dtype)
/tmp/ipykernel_122996/153652883.py:38: ComplexWarning: Casting complex values to real discards the imaginary part
A[l:m, l:m] = A[l:m, l:m] @ Q_2 + sigma * np.eye(m - l, dtype=A.dtype)
Watch the convergence of the eigenvalues:
# prepare experiment:
n = 15
A = Create_random_matrix(n, "diag_dominant")
# for comparison:
w, v = np.linalg.eig(A.copy())
w = np.sort(w)
Q_Hess = Hessenberg_form(A)
list_shifts = ["Zero", "Rayleigh", "Wilkinson"]
print("shifts", " iterartions", " error eigenvalues")
max_Iterations = 100
convergence_eigenvals = {} # dict
for i, shift in enumerate(list_shifts):
B = A.copy()
improvement_diag = np.zeros((n, max_Iterations))
for j in range(max_Iterations):
Q, Iter = QR_iteration_shift(
B, max_Iterations=2, shift=shift, suppress_warning=True
)
improvement_diag[:, j] = np.sort(np.diag(B.copy()))
convergence_eigenvals[shift] = improvement_diag
############
# plotting #
############
# plot over iterations
fig, ax = plt.subplots(1, len(list_shifts), figsize=(5 * len(list_shifts), 5))
fig.suptitle("Convergence of eigenvalues")
for i, shift in enumerate(list_shifts):
ax[i].plot(np.arange(max_Iterations), convergence_eigenvals[shift].T)
ax[i].hlines(w, xmin=0, xmax=max_Iterations, colors="k", linestyles="dashed")
ax[i].set_xlabel("iterations")
ax[i].set_ylabel("eigenvalues")
ax[i].set_title(shift)
ax[i].set_xscale("log")
plt.show()
fig, ax = plt.subplots(
1, len(list_shifts), figsize=(5 * len(list_shifts), 5), sharey=True
)
fig.suptitle("Error of eigenvalues")
for i, shift in enumerate(list_shifts):
for j in [0, 4, 8, 12, 14]:
ax[i].plot(
np.arange(max_Iterations),
np.abs(convergence_eigenvals[shift][j, :] - w[j]),
label=str(np.round(w[j], 2)),
)
ax[i].hlines(
1e-6, xmin=0, xmax=max_Iterations, colors="k", linestyles="dashed", label="eps"
)
ax[i].set_xlabel("iterations")
ax[i].set_ylabel("eigenvalues")
ax[i].set_title(shift)
ax[i].set_xscale("log")
ax[i].set_yscale("log")
ax[i].legend()
plt.show()shifts iterartions error eigenvalues
/tmp/ipykernel_122996/153652883.py:36: ComplexWarning: Casting complex values to real discards the imaginary part
A[l:m, l:m] = A[l:m, l:m] - sigma * np.eye(m - l, dtype=A.dtype)
/tmp/ipykernel_122996/153652883.py:38: ComplexWarning: Casting complex values to real discards the imaginary part
A[l:m, l:m] = A[l:m, l:m] @ Q_2 + sigma * np.eye(m - l, dtype=A.dtype)
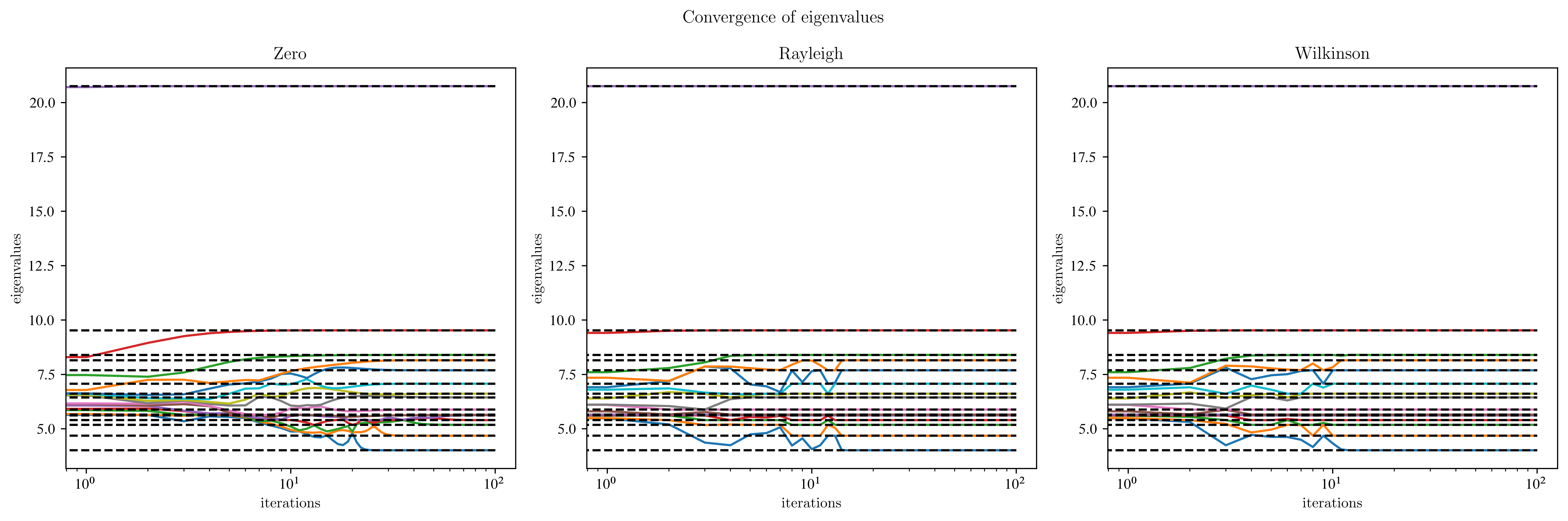
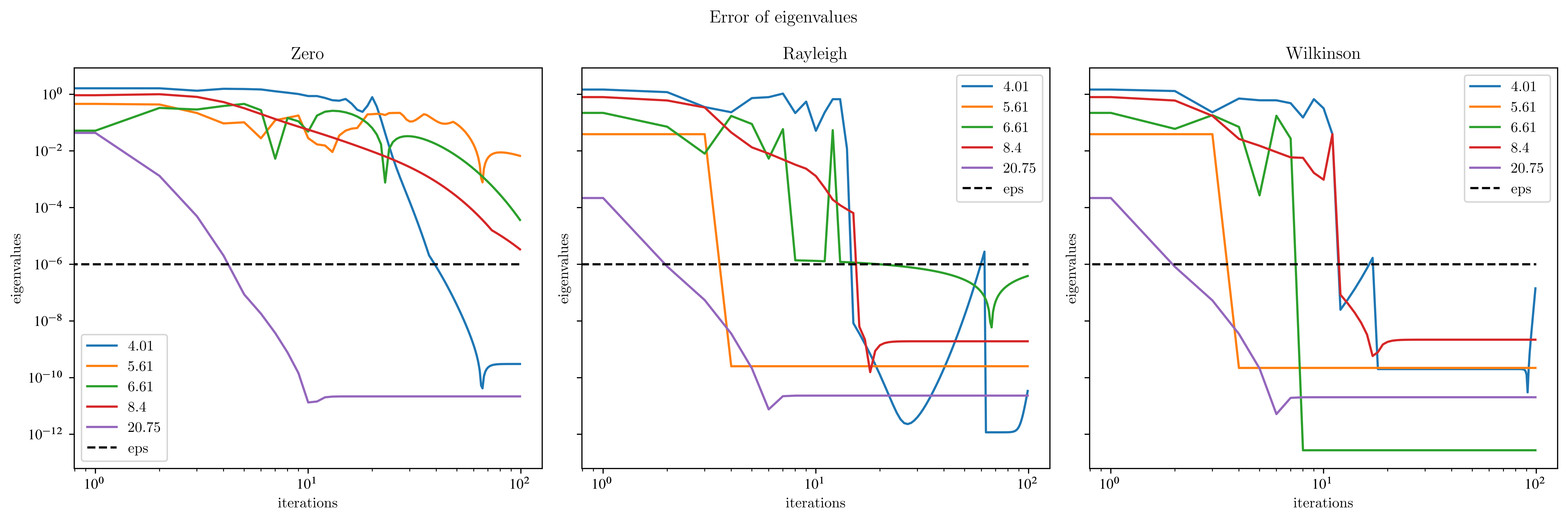
The x scale is logarithmic ! What do you observe?
Solution:¶
Zero shift: polynomial convergence, some EVs do not cross the eps -line
Rayleigh: you see sharp edges, the convergence is not monotonic, there are variations of order eps
Here, eigenvalues jump back from the converged regime to the not converged regime
Deflation¶
Recall: The QR-iteration is another way of the power method. For a matrix
with the submatrices , both square matrices and a rectangular matrix, we calculate
This can easily be generalized to powers. If we split up the problem into two or more subproblems, we will save a lot of work.
It’s possible to neglect values on the subdiagonal, if
A zero on the subdiagonal splits the matrix in two submatrices, which can be iterated independent of each other.
In the next function, we try to find the indices i, at which the above condition is fulfilled. Please try to understand the next functions. Hint: start at the bottom (eigenvalue_QR) and go upwards. What has changed? Why were the changes done?
Solution¶
The function deflation finds subdiagonal entries, which can be neglected.
The split function takes such a list of subdiag entries and returns a list of squared matrices. The matrices are blocks of the diagonal and exclude the given entries
def Deflation(A, eps=1e-10):
# returns list with indices, at which the subdiag of A is zero or neglectable
# INPUT : -A ,nxn matrix
# - eps, a relativ threashold indicating, if an entry vanishes
# OUTPUT: list of columns of the entries on the first subdiagonal, that vanish
# get values of diag and subdiag
diag = np.abs(np.diag(A)).copy()
subdiag = np.abs(np.diag(A, k=-1)).copy()
# values, at which deflation makes sense
compare_value = eps * (diag[:-1] + diag[1:])
# indexarray to return the positions
n = len(subdiag)
positions = np.arange(n)
# checks, if deflation should be applied,
pos_zeros_long = np.where(subdiag < compare_value, positions, -1)
# just give the values of the deflation
pos_zeros = pos_zeros_long[pos_zeros_long > -1]
return pos_zerosA = Create_random_matrix(5)
A[2, 1] = 0
plt.matshow(np.abs(A))
plt.colorbar()
plt.plot(1, 2, "rs")
plt.show()
x = Deflation(A)
print("Deflations in column", x, "(red point)")/usr/lib/python3.13/site-packages/IPython/core/pylabtools.py:170: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
fig.canvas.print_figure(bytes_io, **kw)
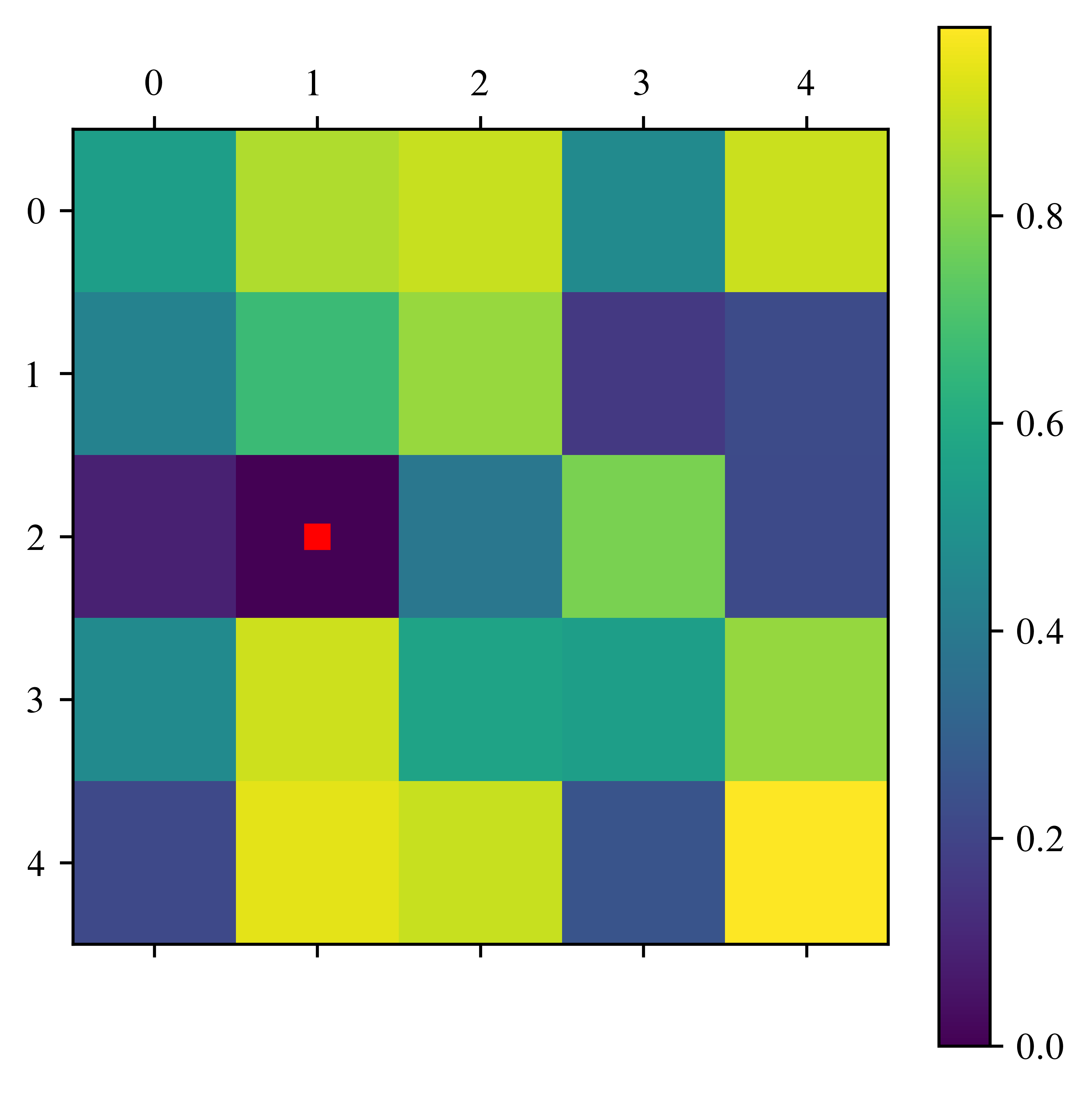
Deflations in column [1] (red point)
Now, we can cut the matrix into pieces.
def Split(A, col_zeros, offset=0):
# this function splits a matrix into smaller matrices at the position of vanishing elements, such that
# QR iteration gives the same result. It ignores 1 dimensional blocks in the complex case and 2 dim, real
# blocks, if the eigenvalues of this block are complex
# INPUT : - A: nxn matrix
# - col_zeros: list with the columns of the subdiagonal, which vanish
# - offset: the row (and the column) from the position of the original matrix
# OUTPUT: list of tuples with first part is the submatrix and the second the offset
lst = []
n = A.shape[0]
recent_zero = 0
# switch from column to row entries of the zeros
row_zeros = [i + 1 for i in col_zeros]
for i in range(len(col_zeros)):
if row_zeros[i] - recent_zero > 1: # can't be eigenvalue block
lst.append(
(
A[recent_zero : row_zeros[i], recent_zero : row_zeros[i]],
recent_zero + offset,
)
)
recent_zero = row_zeros[i]
# last block of matrix
if n - recent_zero > 1:
lst.append((A[recent_zero:, recent_zero:], recent_zero + offset))
return lstimport matplotlib.patches as patches
# insert postions ,where the splitting will take place
pos_zeros = [0, 2, 5]
print("Assume zeros at positions :", pos_zeros)
A = Create_random_matrix(7)
for i in pos_zeros:
A[i + 1, i] = 0
# plot original matrix
plt.matshow(np.abs(A))
# plot zeros (red dots)
for i in pos_zeros:
plt.plot(i, i + 1, "rs")
# plot frame around submatrices
rect0 = patches.Rectangle(
(-0.5, -0.5), 1, 1, linewidth=6, edgecolor="b", facecolor="none"
)
rect1 = patches.Rectangle(
(0.5, 0.5), 2, 2, linewidth=4, edgecolor="k", facecolor="none"
)
rect2 = patches.Rectangle(
(2.5, 2.5), 3, 3, linewidth=4, edgecolor="k", facecolor="none"
)
rect3 = patches.Rectangle(
(5.5, 5.5), 1, 1, linewidth=6, edgecolor="b", facecolor="none"
)
ax = plt.gca()
ax.add_patch(rect0)
ax.add_patch(rect1)
ax.add_patch(rect2)
ax.add_patch(rect3)
plt.title("A")
plt.colorbar()
plt.show()
print(" red dots are set to 0, blue boxes are eigenvalues. The two black boxes remain.")
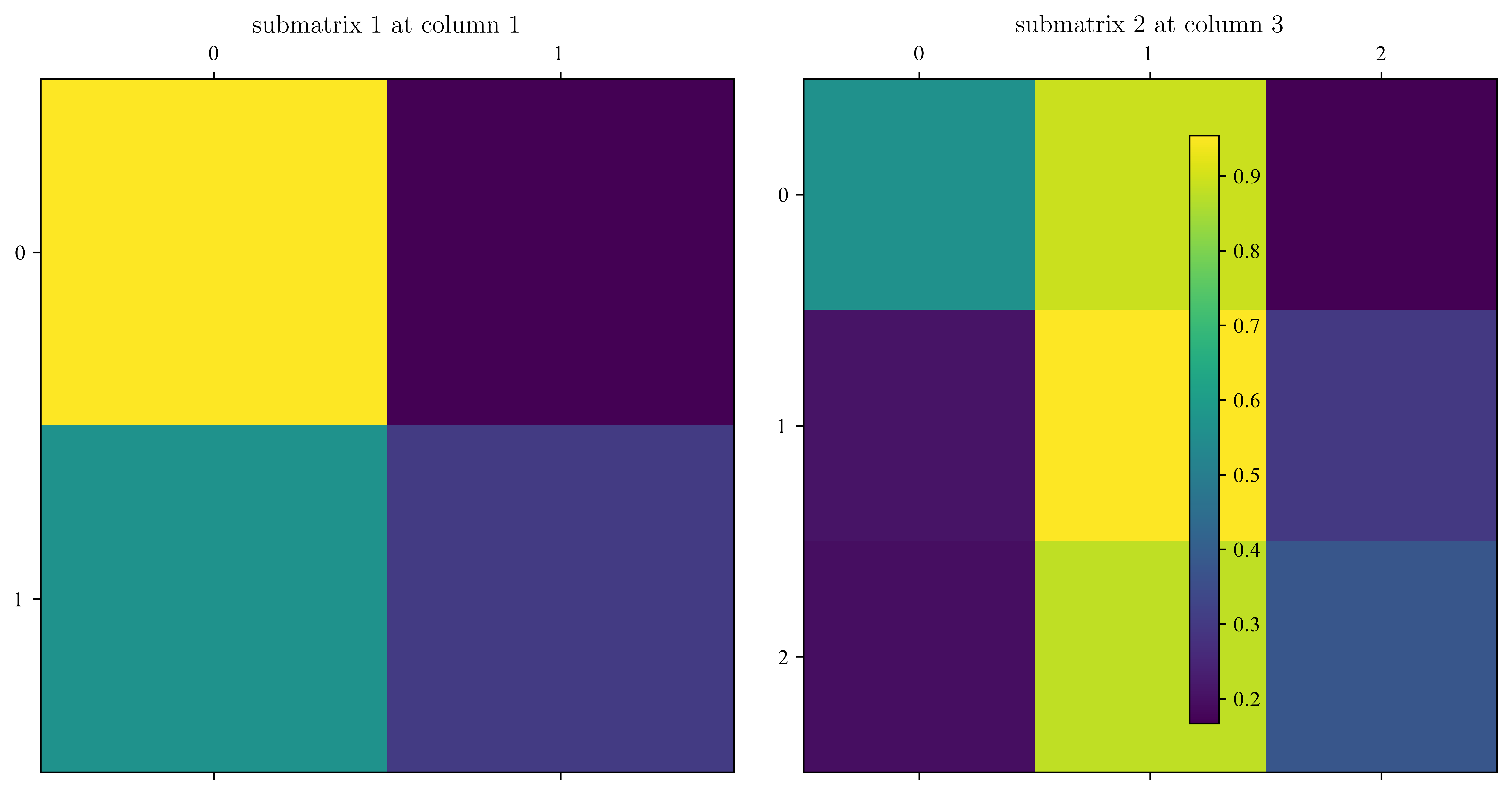
# split the matrix in submatrices
lst = Split(A, pos_zeros)
# plot submatrices
fig, ax = plt.subplots(1, 2, figsize=(10, 5))
for i, mat in enumerate(lst):
im = ax[i].matshow(np.abs(mat[0]))
ax[i].set_title("submatrix " + str(i + 1) + " at column " + str(mat[1]))
plt.colorbar(im, ax=ax)
plt.show()Assume zeros at positions : [0, 2, 5]
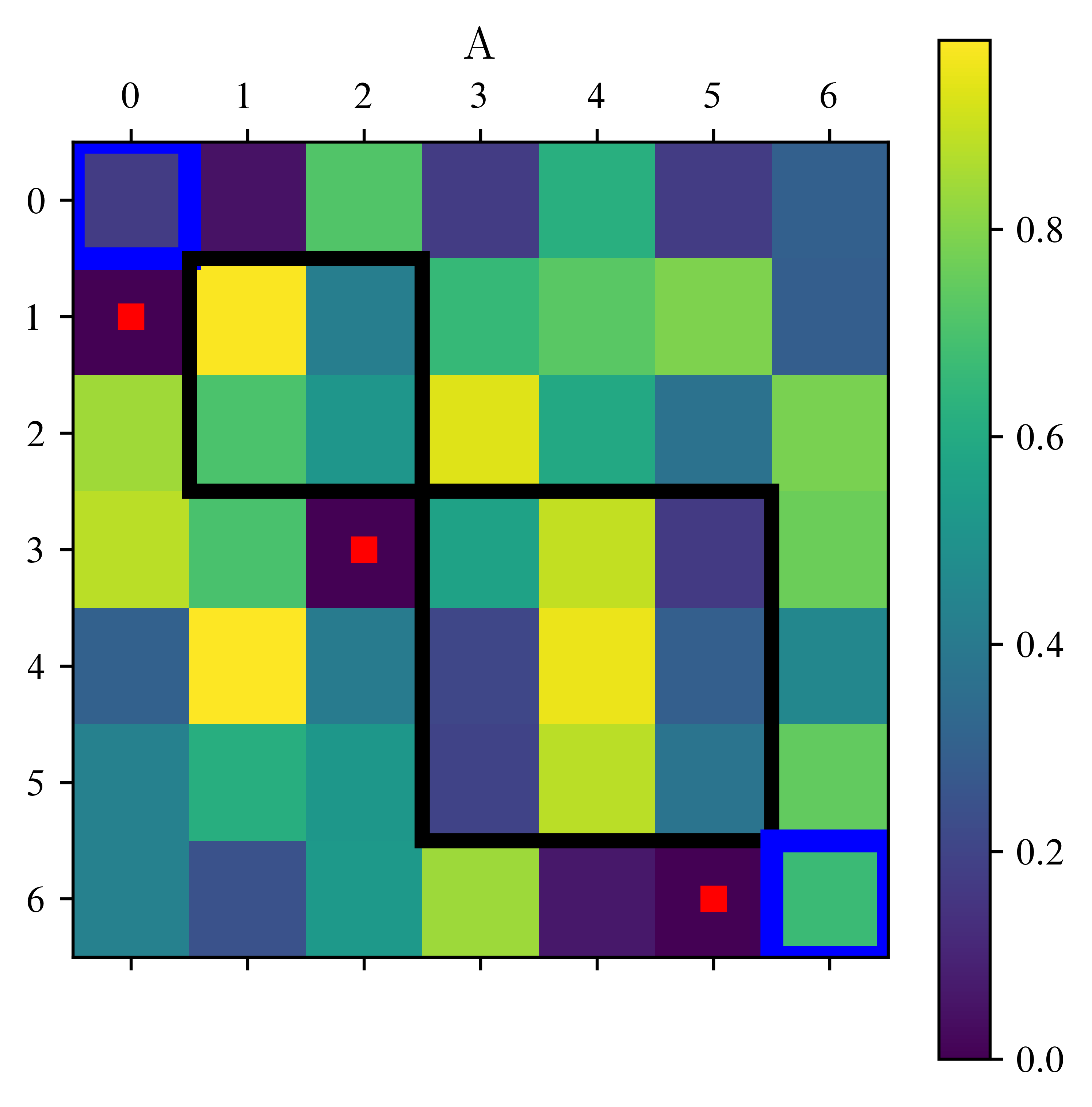
red dots are set to 0, blue boxes are eigenvalues. The two black boxes remain.
This function you already know quite well. What was modified?
Solution:¶
The “check deflation” part was added. In the case of deflation, the interaction breaks. The positions of the possible deflations are returned too.
def QR_iteration(
A,
eps=1e-6,
max_Iterations=10000,
shift="None",
check_deflation=True,
eps_deflation=1e-10,
):
# This function iterates the QR algorithm, until either Max iter reached, the values on the diagonal
# converged or you could deflate the matrix. The matrix A will be overwritten.
# INPUT : - A: squared Matrix in hessenberg form
# - eps: if no diag entry changes more than eps, than the algorithm is converged
# - max_iterations: If this number is reached, it aborts the iteration and gives a warning
# - shift: accelerate convergence, types: Zero, Wilkinson, Francis, Rayleigh, None,
# if None Wilkinson or Francis will be selected
# - check_deflation: checks, if one can deflate the matrix
# - eps_deflation: threshhold for deflation
# OUTPUT: - Q, nxn Matrix, similarity transformation
# - positions of the vanishing entries for the deflation
# - Iter : number of Iterations
# define values, for the computation
if shift == "None":
shift = "Wilkinson"
n = A.shape[0]
Iter = 0
error_ev = 1
diagvals = np.diag(A).copy()
Q_res = np.eye(n, dtype=A.dtype)
pos_zeros = []
# start QR iteration
while error_ev > eps and Iter < max_Iterations:
# evaluate shifts
sigma = evaluate_shift(A[-2:, -2:], shift)
if shift == "Francis": # not important for mos timplementations
Q_2 = Francis(A, sigma)
else:
if not ("complex" in str(A.dtype)):
sigma = np.real(sigma)
# QR iteration
A[:, :] = A - sigma * np.eye(n, dtype=A.dtype)
Q_2 = QR_decomposition_Hess(A)
A[:, :] = A @ Q_2 + sigma * np.eye(n, dtype=A.dtype)
Q_res[:, :] = Q_res @ Q_2
if check_deflation:
pos_zeros = Deflation(A, eps_deflation)
if len(pos_zeros) > 0:
break
Iter += 1
# preparation for next iteration
error_ev = np.max(np.abs(diagvals - np.diag(A)))
diagvals = np.diag(A).copy()
if Iter == max_Iterations:
print("Warning: Algorithm may not converged.")
return Q_res, pos_zeros, Iterdef eigenvalue_QR(
A, shift="None", eps_deflation=1e-10, eps_convergence=1e-6, max_Iterations=1000
):
# calculates the eigenvalues and eigenvectors of matrix A using deflation and the QR iteration method
# INPUT : - A squared Matrix
# - shift, eps_deflation, eps_convergence, max_Iterations see QR_iteration
# OUTPUT: - eigenvalues
# - Q similarity transformation, it contains the eigenvectors if A is normal
Q_res = Hessenberg_form(A)
zero_pos = Deflation(A, eps_deflation)
matrixList = Split(A, zero_pos)
while matrixList != []: # iterate until no matrix in the list
B = matrixList[0][0]
n_start = matrixList[0][1]
n = B.shape[0]
Q, pos_zeros, steps = QR_iteration(
B,
eps=eps_convergence,
max_Iterations=max_Iterations,
shift=shift,
eps_deflation=eps_deflation,
)
if steps < max_Iterations:
splitted_matrices = Split(B, pos_zeros, offset=n_start)
matrixList += splitted_matrices
matrixList.pop(0) # remove matrix B from list
Q_res[:, n_start : n_start + n] = Q_res[:, n_start : n_start + n] @ Q
return np.diag(A), Q_resThe code above may be overwhelming at the first glance. Now it’s on you to verify the code and write some test. Here is a start:
A = Create_random_matrix(5, "diag_dominant")
w, v = np.linalg.eig(A)
B = A.copy()
eigenvals, Q = eigenvalue_QR(A, shift="Wilkinson")
eigenvals = np.sort(eigenvals)
w = np.sort(w)
print("diff ev:\n", np.abs(eigenvals - w) / np.abs(w))diff ev:
[4.20380869e-15 1.84642454e-15 3.35807487e-16 0.00000000e+00
0.00000000e+00]
Comparison of different speed-ups¶
Now compare the different methods for different matrix sizes. We do the first comparison for small matrices.
EXERCISE: Modify the list of shifts.
# enter HERE the type of shift you implemented
shifts = ["Zero", "Rayleigh", "Wilkinson"]
# the following is a dictionary of functions,
# prepare every function in such a way, that it only takes a matrix as input
# use lambda functions as helpers (see next line)
algorithms_slow = {"slow QR": lambda A: QR_iteration_slow(A)}
algorithms_shift = {
"QR_shift" + shifts[i]: lambda A: QR_iteration(
A,
shift=shifts[i],
)
for i in range(3)
}
algorithms_deflation = {
"QR_deflation with " + shifts[i]: lambda A: QR_iteration(
A, shift=shifts[i], check_deflation=True
)
for i in range(3)
}
algortihms_python = {"python": lambda A: np.linalg.eig(A)}
algortigms = {
**algorithms_slow,
**algorithms_shift,
**algorithms_deflation,
**algortihms_python,
}
max_size = 7
times = {alg: np.zeros(max_size - 3) for alg in algortigms.keys()}
for i in range(3, max_size):
A = laplace_matrix_square(i)
Q = Hessenberg_form(A)
for alg in algortigms:
start = time.time()
algortigms[alg](A.copy())
end = time.time()
times[alg][i - 3] = end - start
for alg in algortigms:
plt.plot(times[alg], label=alg)
plt.legend()
plt.yscale("log")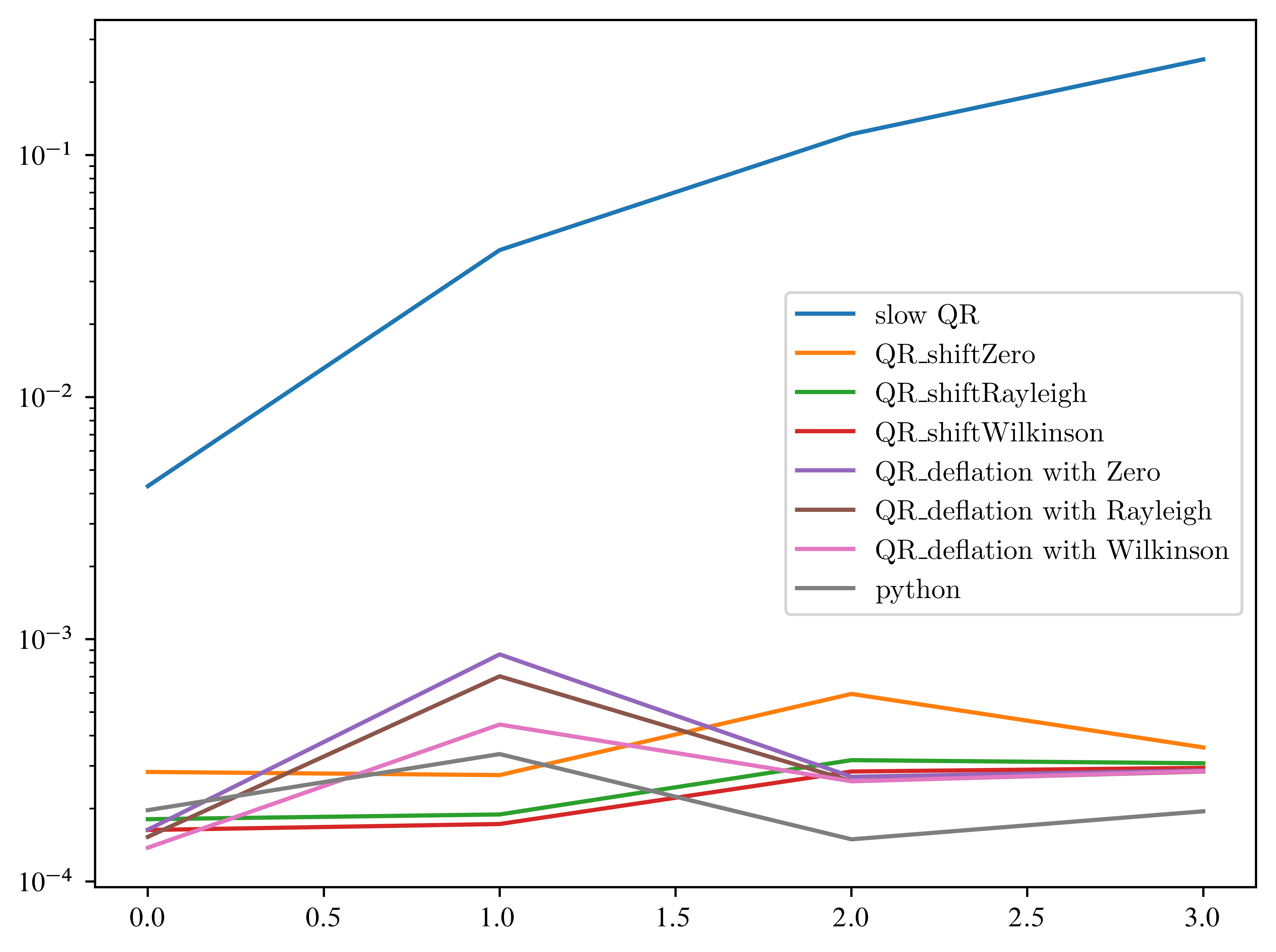
Let’s continue the competition without the slow version ...
Let’s copy the cell from above and remove the slow version (because it’s really slow...)
At this point: Again a Runtime Warning. All your unsaved data maybe lost, if your pc breaks down.
EXERCISE:
Modify the list of shifts first. Please increase the max_size slowly !
First set up all algorithms and then change test_all_implementations to True.
test_all_implementations = True
shifts = ["Zero", "Rayleigh", "Wilkinson"]
n_shifts = len(shifts)
algorithms_shift = {
"QR_shift" + shifts[i]: lambda A: QR_iteration(
A,
shift=shifts[i],
)
for i in range(n_shifts)
}
algorithms_deflation = {
"QR_deflation with " + shifts[i]: lambda A: QR_iteration(
A, shift=shifts[i], check_deflation=True
)
for i in range(n_shifts)
}
algortihms_python = {
"python_np": lambda A: np.linalg.eig(A),
"python_scipy": lambda A: scl.eig(A),
}
algortigms = {**algorithms_shift, **algorithms_deflation, **algortihms_python}
max_size = 18
times = {alg: np.zeros(max_size - 3) for alg in algortigms.keys()}
if test_all_implementations:
for i in range(3, max_size):
A = laplace_matrix_square(i)
B = A.copy()
for alg in algortigms:
if "shift" in alg:
start = time.time()
Q = Hessenberg_form(B)
else:
start = time.time()
algortigms[alg](B)
end = time.time()
times[alg][i - 3] = end - start
plt.figure()
for alg in algortigms:
plt.plot(times[alg], label=alg)
plt.legend()
plt.yscale("log")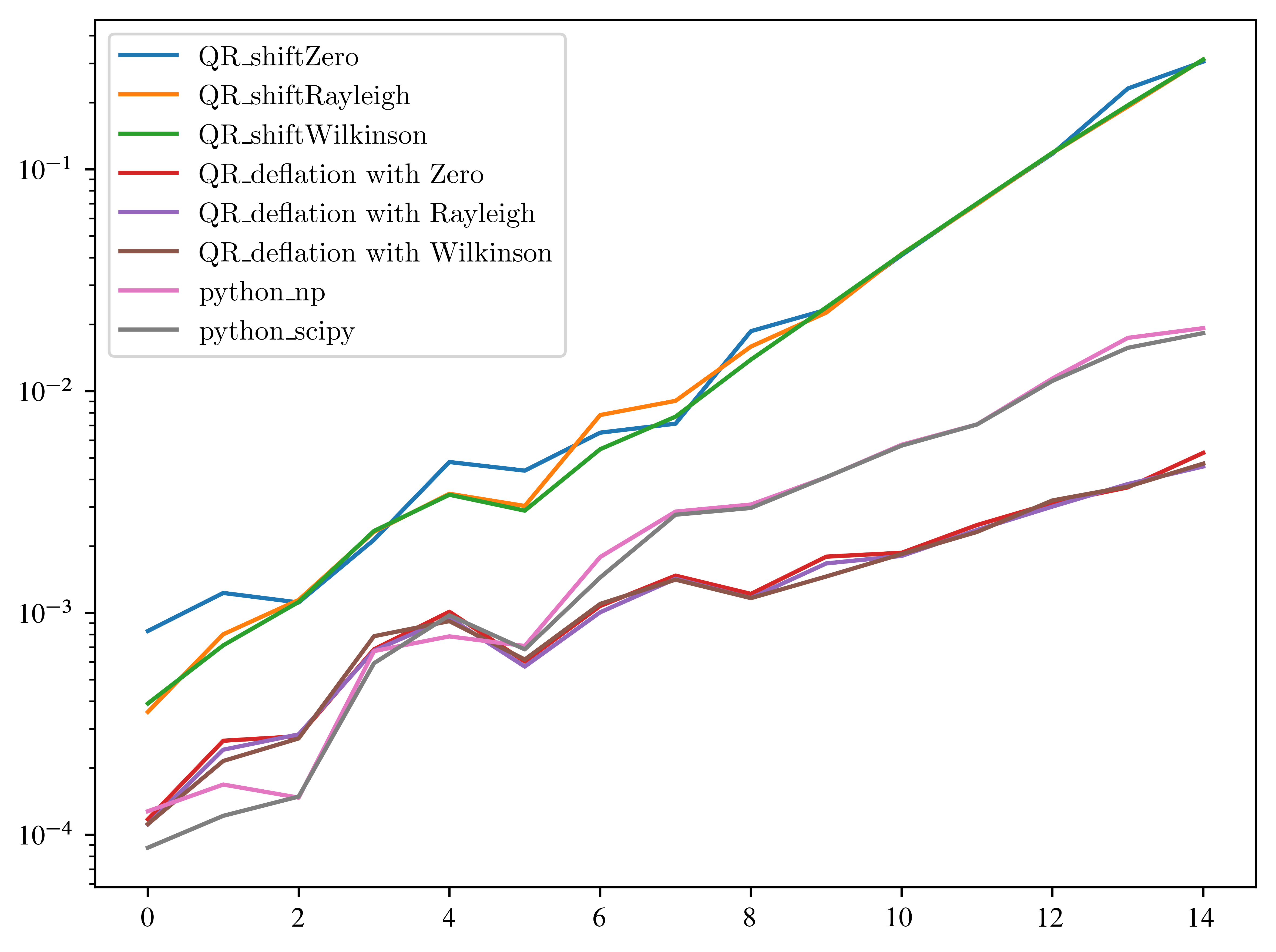
Yuppie! Great, we beat python! What about other matrices? To get a better impression, we run 100 experiments and average the time. So the next one can take some time...
EXERCISE: Modify the list of shifts.
shifts = ["Zero", "Rayleigh", "Wilkinson"]
n_shifts = len(shifts)
algorithms_shift = {
"QR_shift" + shifts[i]: lambda A: QR_iteration(
A,
shift=shifts[i],
)
for i in range(n_shifts)
}
algorithms_deflation = {
"QR_deflation with " + shifts[i]: lambda A: QR_iteration(
A, shift=shifts[i], check_deflation=True
)
for i in range(n_shifts)
}
algortihms_python = {"python": lambda A: np.linalg.eig(A)}
algortigms = {**algorithms_shift, **algorithms_deflation, **algortihms_python}
max_size = 6
times = {alg: np.zeros(max_size - 3) for alg in algortigms.keys()}
repetitions = 100
for i in range(3, max_size):
for j in range(repetitions): # average over different random matrices
A = Create_random_matrix(i * i, "diag_dominant")
for alg in algortigms:
B = A.copy() # all algortigms use the same random matrix
if "shift" in alg:
start = time.time()
Q = Hessenberg_form(B)
else:
start = time.time()
algortigms[alg](B)
end = time.time()
times[alg][i - 3] += end - start
plt.figure()
for alg in algortigms:
plt.plot(times[alg] / repetitions, label=alg)
plt.legend()
plt.yscale("log")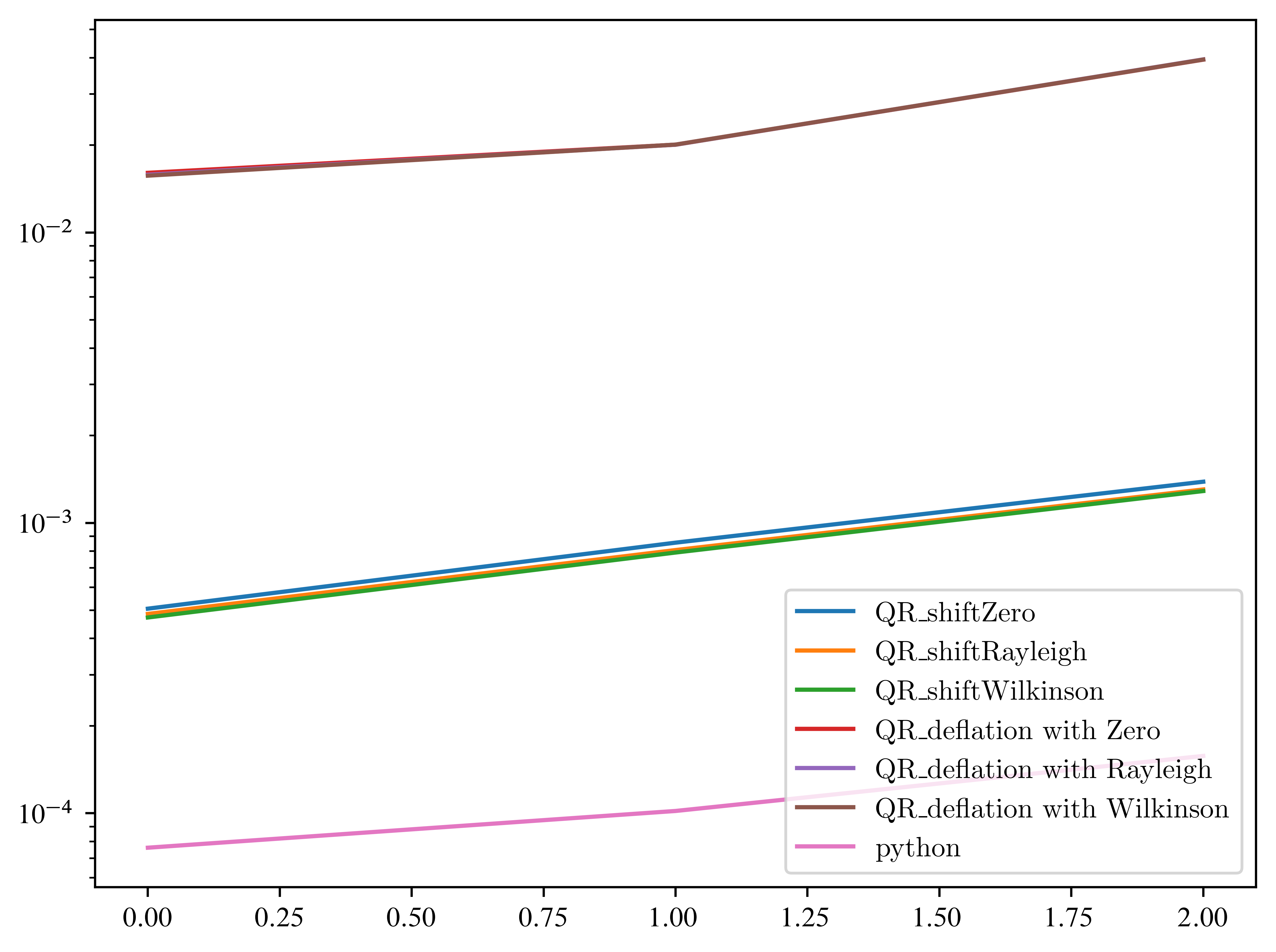
Further progress¶
You can choose to
Calculate the Eigenvectors
Apply the algorithms above to more complicated problems, e.g. singular value decomposition (SVD), inverse of matrix,
Think about, what changes, if we use arbitrary matrices with possible complex eigenvalues? What do we have to change, if input a complex matrix? Hint: for real matrices, complex eigenvalues appear in complex conjugated pairs only.
Which steps can be parallelized? What is good way to parallelize it? (independent tasks, SIMD, ...)
Implement a sparse algorithm. This maybe a lot of work, because almost every function has to be rewritten, such that sparsity can be used. Try to beat the python implementation.
Solution¶
More complicated problems:
SVD:
want ,
calculate eigenvals, and vecs of to get and . (Be carful with the dimensions.)
Do the same for to get . Maybe it’s easier to on a different way
Inverse:
Complex Eigenvals:
Real entries:
appeare in pairs, so a -blocks remain on the diag, from which you can calculate the eigenvals analytically
Changes: Deflation, Shifts to implicit double shifts
Complex entries:
complex givens rotations
complex conjugate in Q , hermitian conjugate (np.conj(Q.T))
some np.abs() somewhere, for deflation criteria,...
Parallizations:
Householder trafo can be performed in SIMD (like matrix mult of dense matrices)
The blocks can be iterated in parallel (task level parallelism)
multiple shifts “at once” (Multishift) “algorithmic optimization”
Some Ideas for optimization: 0. Calculate only eigenvalues and the coresponding vector later via the power iteration - reduce storage (ignore Q) - maybe not faster
Householde Trafo:
Householder vector contains a lot of zeros, just apply the nonzero entries.
Be aware, the matrix can become dense during the transformation process (usually at half matrix size)
a sparse and a dense matrix version should be availiable and the transition between should be without any additional time
set small entries to 0, (ignore them in calculation)
an idea: masking
Givens rotation
implement a matrix-free application (do not assemble the matrix and aplly it). Instead calculate the entries directly and change them inside the matrix
here the most time is spent, so an efficient
apply all trafsormations only on the subblocks.
for spd matatrices, the Hesserberg-form is trigiagonal.
you can do all the calculations with 2 vectors (diagonal entries and subdiagonal entries)
use implicit shifts
import cProfile
A = laplace_matrix_square(24)
cProfile.run("eigenvalue_QR(A)") 1073187 function calls in 12.241 seconds
Ordered by: standard name
ncalls tottime percall cumtime percall filename:lineno(function)
853 0.021 0.000 0.035 0.000 1575710025.py:1(Deflation)
852 1.058 0.001 2.399 0.003 1990452498.py:1(QR_decomposition_Hess)
243909 1.030 0.000 1.319 0.000 2273048651.py:1(Givens_rotation)
548 3.558 0.006 6.128 0.011 2284605642.py:1(QR_iteration)
574 0.109 0.000 0.207 0.000 2652707984.py:1(Householder_matrix)
1 3.399 3.399 3.550 3.550 3911630699.py:1(Hessenberg_form)
852 0.013 0.000 0.013 0.000 3966089958.py:1(evaluate_shift)
1 0.180 0.180 0.983 0.983 584013956.py:1(eigenvalue_QR)
549 0.005 0.000 0.005 0.000 604879709.py:1(Split)
1 0.000 0.000 10.993 10.993 <string>:1(<module>)
852 0.001 0.000 0.001 0.000 _dtype.py:23(_kind_name)
852 0.003 0.000 0.014 0.000 _dtype.py:32(__str__)
852 0.001 0.000 0.006 0.000 _dtype.py:326(_name_includes_bit_suffix)
852 0.004 0.000 0.011 0.000 _dtype.py:342(_name_get)
1722 0.000 0.000 0.001 0.000 _linalg.py:187(isComplexType)
1722 0.000 0.000 0.000 0.000 _linalg.py:2618(_norm_dispatcher)
1722 0.006 0.000 0.011 0.000 _linalg.py:2622(norm)
3679 0.024 0.000 0.147 0.000 _twodim_base_impl.py:176(eye)
2863 0.000 0.000 0.000 0.000 _twodim_base_impl.py:254(_diag_dispatcher)
2863 0.006 0.000 0.015 0.000 _twodim_base_impl.py:258(diag)
852 0.000 0.000 0.000 0.000 _type_check_impl.py:80(_real_dispatcher)
852 0.001 0.000 0.001 0.000 _type_check_impl.py:84(real)
2 0.000 0.000 0.000 0.000 base_events.py:1965(_timer_handle_cancelled)
3 1.128 0.376 10.010 3.337 base_events.py:1970(_run_once)
8 0.000 0.000 0.000 0.000 base_events.py:2068(get_debug)
2 0.000 0.000 0.000 0.000 base_events.py:457(create_future)
4 0.000 0.000 0.000 0.000 base_events.py:554(_check_closed)
8 0.000 0.000 0.000 0.000 base_events.py:772(time)
2 0.000 0.000 0.000 0.000 base_events.py:781(call_later)
2 0.000 0.000 0.000 0.000 base_events.py:805(call_at)
2 0.000 0.000 0.000 0.000 base_events.py:823(call_soon)
2 0.000 0.000 0.000 0.000 base_events.py:852(_call_soon)
1 0.000 0.000 0.000 0.000 contextlib.py:145(__exit__)
2 0.000 0.000 0.000 0.000 events.py:113(__init__)
2 0.000 0.000 0.000 0.000 events.py:157(cancel)
4 0.000 0.000 0.000 0.000 events.py:36(__init__)
2 0.000 0.000 0.000 0.000 events.py:73(cancel)
4 0.000 0.000 0.000 0.000 events.py:87(_run)
2863 0.000 0.000 0.000 0.000 fromnumeric.py:1691(_diagonal_dispatcher)
2863 0.003 0.000 0.008 0.000 fromnumeric.py:1695(diagonal)
304 0.000 0.000 0.000 0.000 fromnumeric.py:3047(_max_dispatcher)
304 0.001 0.000 0.005 0.000 fromnumeric.py:3052(max)
304 0.002 0.000 0.004 0.000 fromnumeric.py:69(_wrapreduction)
2 0.000 0.000 0.000 0.000 futures.py:310(_set_result_unless_cancelled)
1 0.000 0.000 0.000 0.000 history.py:1016(_writeout_output_cache)
1 0.000 0.000 0.000 0.000 history.py:1065(hold)
2 0.000 0.000 0.000 0.000 iostream.py:118(_run_event_pipe_gc)
2 0.000 0.000 0.000 0.000 iostream.py:127(_event_pipe_gc)
853 0.000 0.000 0.000 0.000 multiarray.py:391(where)
574 0.000 0.000 0.000 0.000 multiarray.py:769(dot)
574 0.052 0.000 0.058 0.000 numeric.py:1002(tensordot)
574 0.000 0.000 0.000 0.000 numeric.py:1199(<genexpr>)
1148 0.000 0.000 0.000 0.000 numeric.py:1200(<genexpr>)
574 0.000 0.000 0.000 0.000 numeric.py:1205(<genexpr>)
1148 0.000 0.000 0.000 0.000 numeric.py:1206(<genexpr>)
574 0.000 0.000 0.000 0.000 numeric.py:998(_tensordot_dispatcher)
1704 0.001 0.000 0.003 0.000 numerictypes.py:293(issubclass_)
852 0.002 0.000 0.005 0.000 numerictypes.py:475(issubdtype)
4 0.000 0.000 0.000 0.000 selector_events.py:740(_process_events)
3 0.000 0.000 0.001 0.000 selectors.py:435(select)
4 0.000 0.000 0.000 0.000 tasks.py:703(sleep)
6 0.000 0.000 0.000 0.000 threading.py:1136(is_alive)
1 0.000 0.000 0.000 0.000 threading.py:303(__enter__)
1 0.000 0.000 0.000 0.000 threading.py:312(_release_save)
1 0.000 0.000 0.000 0.000 threading.py:318(_is_owned)
6 0.000 0.000 0.000 0.000 threading.py:605(is_set)
1 0.000 0.000 0.000 0.000 traitlets.py:1512(_notify_trait)
1 0.000 0.000 0.000 0.000 traitlets.py:1523(notify_change)
1 0.000 0.000 0.000 0.000 traitlets.py:1527(_notify_observers)
2 0.000 0.000 0.000 0.000 traitlets.py:2304(validate)
2 0.000 0.000 0.000 0.000 traitlets.py:3474(validate)
2 0.000 0.000 0.000 0.000 traitlets.py:3486(validate_elements)
2 0.000 0.000 0.000 0.000 traitlets.py:3624(validate_elements)
2 0.000 0.000 0.000 0.000 traitlets.py:3631(set)
1 0.000 0.000 0.000 0.000 traitlets.py:629(get)
1 0.000 0.000 0.000 0.000 traitlets.py:676(__get__)
2 0.000 0.000 0.000 0.000 traitlets.py:689(set)
2 0.000 0.000 0.000 0.000 traitlets.py:708(__set__)
2 0.000 0.000 0.000 0.000 traitlets.py:718(_validate)
2 0.000 0.000 0.000 0.000 traitlets.py:727(_cross_validate)
12 0.000 0.000 0.000 0.000 typing.py:2371(cast)
2 0.000 0.000 0.000 0.000 {built-in method _asyncio.get_running_loop}
2 0.000 0.000 0.000 0.000 {built-in method _contextvars.copy_context}
2 0.000 0.000 0.000 0.000 {built-in method _heapq.heappop}
2 0.000 0.000 0.000 0.000 {built-in method _heapq.heappush}
7358 0.001 0.000 0.001 0.000 {built-in method _operator.index}
1 0.000 0.000 0.000 0.000 {built-in method _thread.allocate_lock}
1 1.190 1.190 12.183 12.183 {built-in method builtins.exec}
5 0.000 0.000 0.000 0.000 {built-in method builtins.hasattr}
2870 0.001 0.000 0.001 0.000 {built-in method builtins.isinstance}
7704 0.003 0.000 0.003 0.000 {built-in method builtins.issubclass}
574 0.002 0.000 0.002 0.000 {built-in method builtins.iter}
9716 0.002 0.000 0.002 0.000 {built-in method builtins.len}
1 0.000 0.000 0.000 0.000 {built-in method builtins.next}
2 0.000 0.000 0.000 0.000 {built-in method math.ceil}
2 0.000 0.000 0.000 0.000 {built-in method math.isnan}
2296 0.002 0.000 0.002 0.000 {built-in method math.prod}
853 0.002 0.000 0.002 0.000 {built-in method numpy.arange}
243909 0.159 0.000 0.159 0.000 {built-in method numpy.array}
5726 0.001 0.000 0.001 0.000 {built-in method numpy.asanyarray}
2870 0.000 0.000 0.000 0.000 {built-in method numpy.asarray}
3679 0.121 0.000 0.121 0.000 {built-in method numpy.zeros}
8 0.000 0.000 0.000 0.000 {built-in method time.monotonic}
1 0.000 0.000 0.000 0.000 {method '__enter__' of '_thread.lock' objects}
4 0.000 0.000 0.000 0.000 {method '__exit__' of '_thread.lock' objects}
1 0.000 0.000 0.000 0.000 {method '__exit__' of 'sqlite3.Connection' objects}
2 0.000 0.000 0.000 0.000 {method 'acquire' of '_thread.lock' objects}
5 0.000 0.000 0.000 0.000 {method 'append' of 'collections.deque' objects}
548 0.000 0.000 0.000 0.000 {method 'append' of 'list' objects}
2 0.000 0.000 0.000 0.000 {method 'cancelled' of '_asyncio.Future' objects}
247041 0.077 0.000 0.077 0.000 {method 'copy' of 'numpy.ndarray' objects}
2863 0.003 0.000 0.003 0.000 {method 'diagonal' of 'numpy.ndarray' objects}
1 0.000 0.000 0.000 0.000 {method 'disable' of '_lsprof.Profiler' objects}
1722 0.003 0.000 0.003 0.000 {method 'dot' of 'numpy.ndarray' objects}
2 0.000 0.000 0.000 0.000 {method 'extend' of 'list' objects}
4 0.000 0.000 0.000 0.000 {method 'get' of 'dict' objects}
6 0.000 0.000 0.000 0.000 {method 'is_done' of '_thread._ThreadHandle' objects}
306 0.000 0.000 0.000 0.000 {method 'items' of 'dict' objects}
3 0.000 0.000 0.001 0.000 {method 'poll' of 'select.epoll' objects}
548 0.000 0.000 0.000 0.000 {method 'pop' of 'list' objects}
4 0.000 0.000 0.000 0.000 {method 'popleft' of 'collections.deque' objects}
1722 0.001 0.000 0.001 0.000 {method 'ravel' of 'numpy.ndarray' objects}
304 0.002 0.000 0.002 0.000 {method 'reduce' of 'numpy.ufunc' objects}
1 0.000 0.000 0.000 0.000 {method 'release' of '_thread.lock' objects}
246205 0.060 0.000 0.060 0.000 {method 'reshape' of 'numpy.ndarray' objects}
4 0.000 0.000 0.000 0.000 {method 'run' of '_contextvars.Context' objects}
2 0.000 0.000 0.000 0.000 {method 'set_result' of '_asyncio.Future' objects}
1148 0.001 0.000 0.001 0.000 {method 'transpose' of 'numpy.ndarray' objects}