Least Squares¶
Definition: Supervised learning
Given a matrix and some associated output vector , find a function that takes a vector and returns a prediction for where some “loss function” is minimized for all .
The task:
Given a vector can we find a function that accurately predicts the pregression of the disease of the patients in most cases?
To get some sense of the quality of this predictor, we gather the following numbers:
True Positives , number of correctly predicted events as positive
False Positives , number of events falsely predicted as positive which belong to negative
True Negatives , number of correctly predicted events as negative
False Negatives , number of events falsely predicted as negative which belong to positive
This is visualised in the so called confusion matrix: $$
$$ We can look at the fraction of correctly labeled observations in the data
or simply put
Now we try to find a function where the accuracy is higher than 0.5
?load_diabetesObject `load_diabetes` not found.
import pandas as pd
from sklearn.datasets import load_diabetes
# load dataset
diabetes = load_diabetes()
df = pd.DataFrame(diabetes.data, columns=diabetes.feature_names)
df["target"] = diabetes.target
n = 5
print(f"First {n} rows of the diabetes dataset:")
print(df.head(n))First 5 rows of the diabetes dataset:
age sex bmi bp s1 s2 s3 \
0 0.038076 0.050680 0.061696 0.021872 -0.044223 -0.034821 -0.043401
1 -0.001882 -0.044642 -0.051474 -0.026328 -0.008449 -0.019163 0.074412
2 0.085299 0.050680 0.044451 -0.005670 -0.045599 -0.034194 -0.032356
3 -0.089063 -0.044642 -0.011595 -0.036656 0.012191 0.024991 -0.036038
4 0.005383 -0.044642 -0.036385 0.021872 0.003935 0.015596 0.008142
s4 s5 s6 target
0 -0.002592 0.019907 -0.017646 151.0
1 -0.039493 -0.068332 -0.092204 75.0
2 -0.002592 0.002861 -0.025930 141.0
3 0.034309 0.022688 -0.009362 206.0
4 -0.002592 -0.031988 -0.046641 135.0
import numpy as np
y = diabetes.target
# min and max value
y_min, y_max = np.min(y), np.max(y)
print("Min target:", y_min)
print("Max target:", y_max)
# Use median as threshold (for classification later)
threshold = np.mean(y)
print("Median (threshold):", threshold)
print(f"""
Interpretation:
- values <= to {threshold:.1f} = disease more stable
- values > {threshold:.1f} = disease has gotten worse
""")Min target: 25.0
Max target: 346.0
Median (threshold): 152.13348416289594
Interpretation:
- values <= to 152.1 = disease more stable
- values > 152.1 = disease has gotten worse
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import accuracy_score, confusion_matrix
def plot_bars_and_confusion(
truth,
prediction,
axes=None,
vmin=None,
vmax=None,
cmap="viridis",
title=None,
bar_color=None,
):
accuracy = accuracy_score(truth, prediction)
cm = confusion_matrix(truth, prediction)
if not isinstance(truth, pd.Series):
truth = pd.Series(truth)
if not isinstance(prediction, pd.Series):
prediction = pd.Series(prediction)
correct = pd.Series(truth.values == prediction.values)
truth.sort_index(inplace=True)
prediction.sort_index(inplace=True)
if not axes:
fig, axes = plt.subplots(1, 2, figsize=(8, 2))
if not vmin:
vmin = cm.min()
if not vmax:
vmax = cm.max()
if not bar_color:
correct.value_counts().plot.barh(ax=axes[0])
else:
correct.value_counts().plot.barh(ax=axes[0], color=bar_color)
axes[0].text(150, 0.5, "Accuracy {:0.3f}".format(accuracy))
sns.heatmap(
cm,
annot=True,
fmt="d",
cmap=cmap,
xticklabels=["False", "True"],
yticklabels=["False", "True"],
ax=axes[1],
vmin=vmin,
vmax=vmax,
)
axes[1].set_ylabel("Truth")
axes[1].set_xlabel("Predicted")
if title:
plt.suptitle(title)
plt.show()import numpy as np
import pandas as pd
from sklearn.datasets import load_diabetes
def disease_prediction(truth, feature):
x = df[feature]
threshold_feature = np.median(x)
x_bin = (x < threshold_feature).astype(int)
prediction = x_bin
plot_bars_and_confusion(truth=truth, prediction=prediction)
# load data
diabetes = load_diabetes()
df = pd.DataFrame(diabetes.data, columns=diabetes.feature_names)
df["target"] = diabetes.target
# threshold for disease progression
y = diabetes.target
threshold_truth = np.median(y)
y_bin = (y > threshold_truth).astype(int) # 1 = worse, 0 = stable
# disease_prediction(y_bin, "bmi")
disease_prediction(y_bin, "bp")
# disease_prediction(y_bin, "age")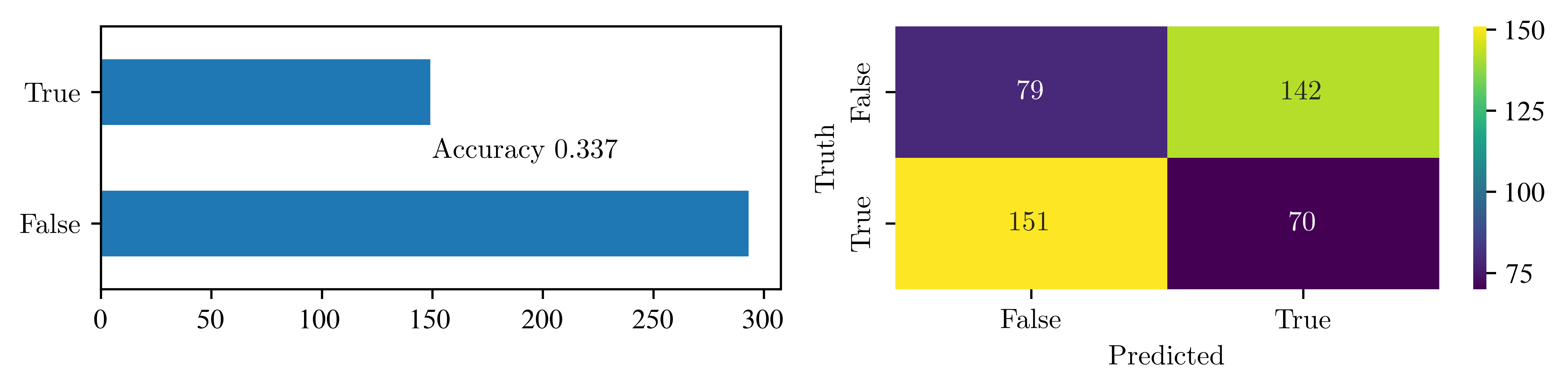
Linear Models¶
Can we improve our predictor by combining more variables into one predictor?
Lets assume a linear weighted combination of variables:
where .
For a single sample of the diabetes data we simply evaluate:
When we include a 1 as the first entry into our sample e.g. we can rewrite in matrix form
where .
How do you find those weights? Like before we choose a loss function and try to opimize it. The loss function measures the error for a single training example (i.e., how far the model’s prediction is from the actual target). In this case we choose a loss function called the residual sum of squares (RSS). We calculate it over all samples in a matrix .
Undefined control sequence: \RSS at position 16: L(\v{\beta}) = \̲R̲S̲S̲(\v{\beta}) = \…
L(\v{\beta}) = \RSS(\v{\beta}) = \sum_{i=1}^N (y_i - \v{x}_i^\top \v\beta)^2.Remark: Here is a row in , hence the transpose.
Now we rewrite the loss function in matrix form:
Undefined control sequence: \RSS at position 1: \̲R̲S̲S̲(\v\beta) = (\v…
\RSS(\v\beta) = (\v{y} - \textbf{X} \v\beta)^\top (\v{y} - \textbf{X} \v\beta )and we optimize the loss function just like we would any other function, by differentiating with respect to and setting the result equals to zero.
We need the product rule:
this yields
Solving for leads to
We just performed Linear Least Squares regression.
Now we can define a function to predict disease progression according to
Linear Regression with sklearn¶
Goal: Create a 2D dataset with two classes and use the least squares method to seperate them.
Create random points in a 2D parameter space
from sklearn.datasets import make_blobsUse scikit-learn’s linear regressor to find the parameters for .
from sklearn import linear_model regression = linear_model.LinearRegression() regression.fit(X, Y) b_1, b_2 = regression.coef_ b_0 = regression.intercept_Draw a dashed line into the plot where .
x1 = np.linspace(-2, 2) x2 = ... plt.plot(x1, x2, color='gray', linestyle='--')
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.colors import ListedColormap
from sklearn import linear_model
from sklearn.datasets import make_blobs
# create example data
X, y = make_blobs(n_samples=100, centers=2, cluster_std=1.5, random_state=0)
# train the linear regressor and save the coefficents
# y = b0 + b1x1 + b2x2
regression = linear_model.LinearRegression() # creates empty object
regression.fit(X, y)
b_1, b_2 = regression.coef_ # weights
b_0 = regression.intercept_ # bias
# solve the function y = b_0 + b_1*X_1 + b_2 * X_2 for X2
# remember: y is either > 0.5 or <= 0.5
x1 = np.linspace(-1, 4)
x2 = (0.5 - b_0 - b_1 * x1) / b_2
discrete_cmap = ListedColormap(["red", "blue"])
plt.scatter(X[:, 0], X[:, 1], s=25, c=y, cmap=discrete_cmap)
plt.plot(x1, x2, color="gray", linestyle="--") # draws line for y >= / < 0.5
plt.fill_between(x1, x2, 6, color="crimson", alpha=0.1)
plt.fill_between(x1, x2, -1, color="blue", alpha=0.1)
plt.grid()
plt.xlabel("X1")
plt.ylabel("X2")
plt.margins(x=0, y=0)
plt.show()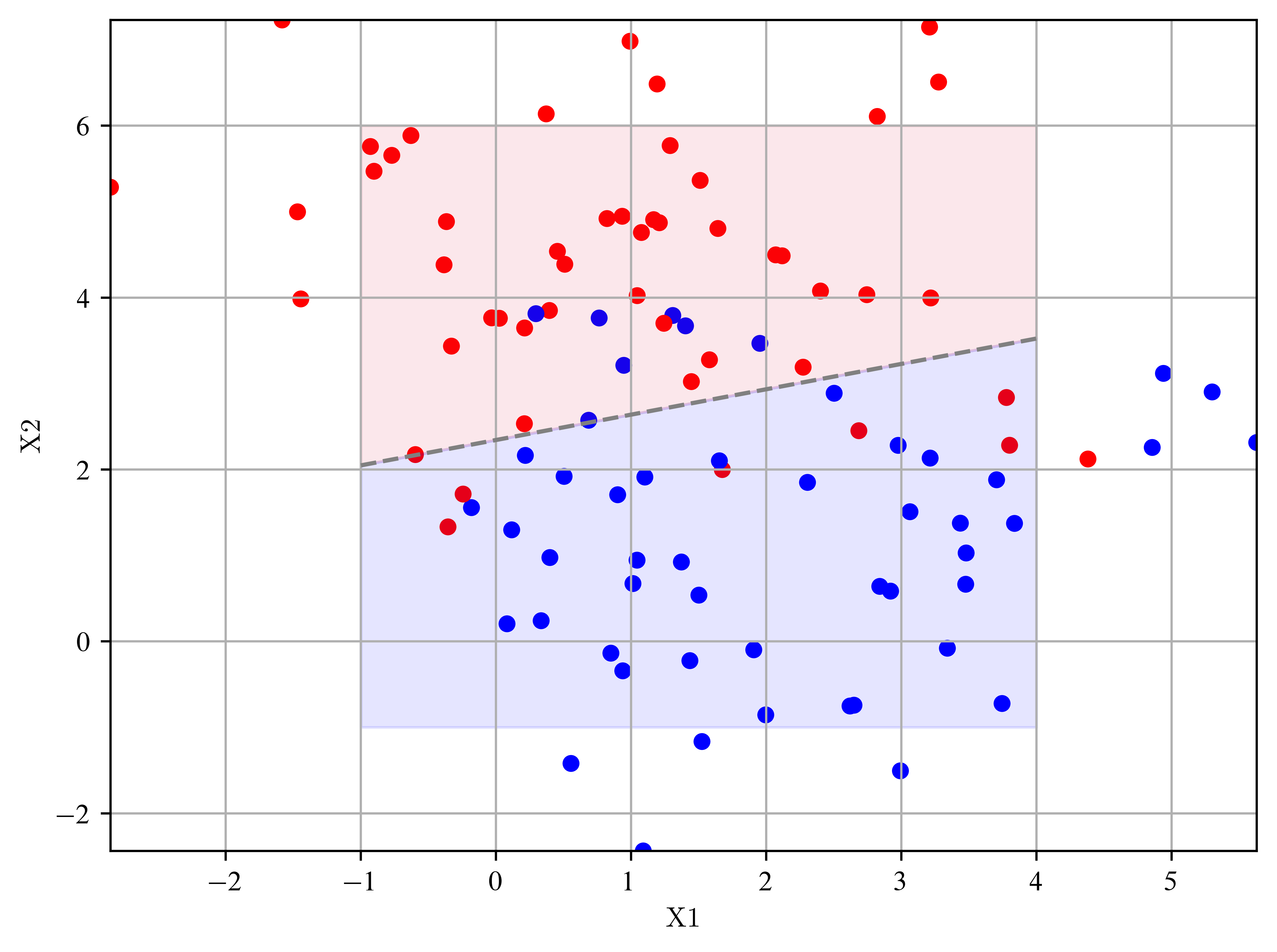
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.datasets import load_diabetes
# Load data
diabetes = load_diabetes()
df = pd.DataFrame(diabetes.data, columns=diabetes.feature_names)
df["target"] = diabetes.target
# Create progression classes (1 = worse, 0 = stable)
# target is continious, make a binary class out of it by creating a threshold
threshold = np.median(df["target"])
df["progression_class"] = (df["target"] > threshold).astype(int)
# --- Function for scatterplot ---
def plot_progression(feature_x, feature_y, data=df):
fig, ax = plt.subplots(figsize=(8, 6))
sc = ax.scatter(
data[feature_x],
data[feature_y],
c=data["progression_class"],
cmap=plt.cm.get_cmap("bwr", 2),
vmin=-0.5,
vmax=1.5,
alpha=0.7,
)
ax.set_xlabel(feature_x)
ax.set_ylabel(feature_y)
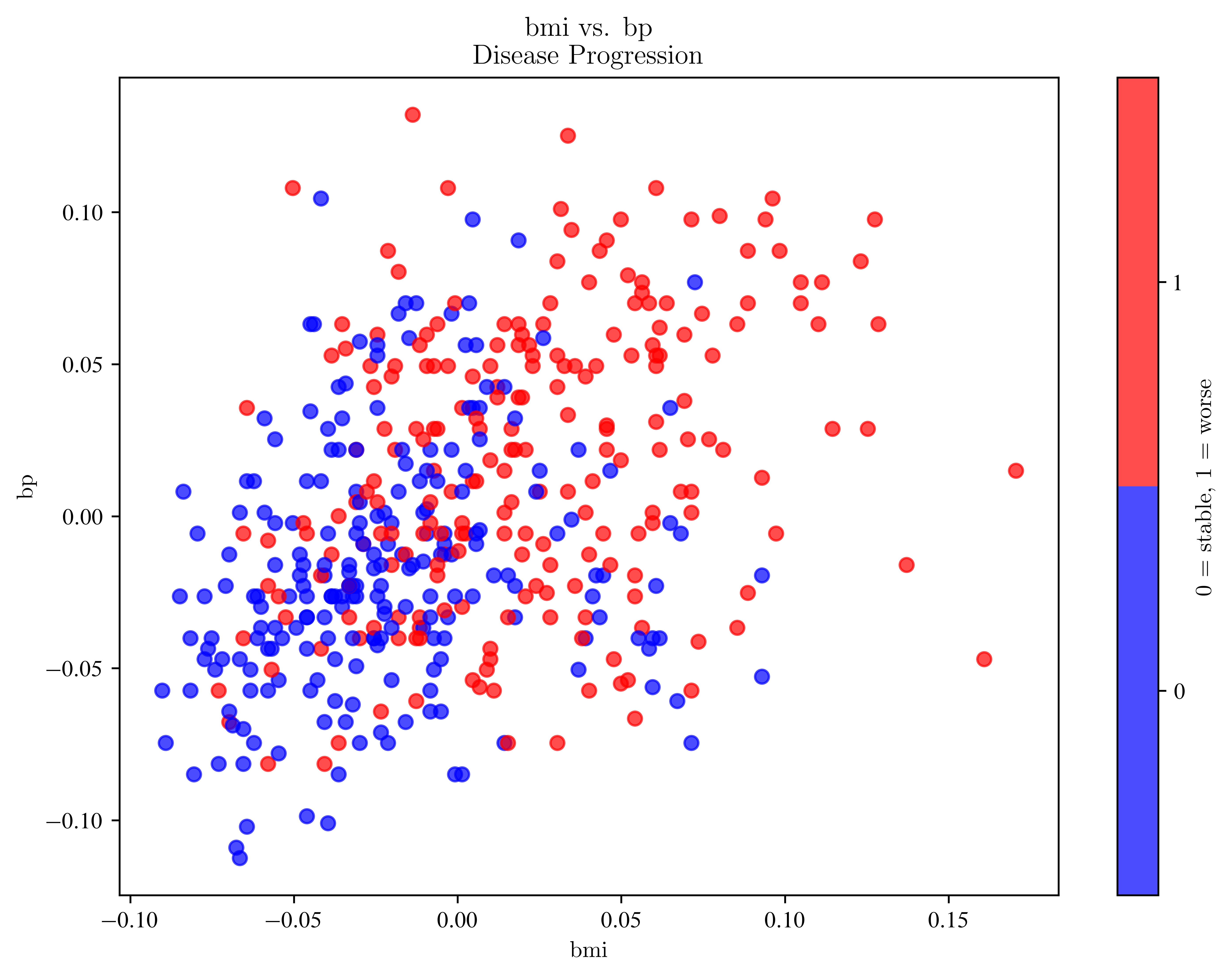
ax.set_title(f"{feature_x} vs. {feature_y}\nDisease Progression")
cbar = fig.colorbar(sc, ticks=[0, 1], label="0 = stable, 1 = worse")
plot_progression("bmi", "bp")
# plot_progression("bmi", "age")/tmp/ipykernel_123479/1633364373.py:25: MatplotlibDeprecationWarning: The get_cmap function was deprecated in Matplotlib 3.7 and will be removed in 3.11. Use ``matplotlib.colormaps[name]`` or ``matplotlib.colormaps.get_cmap()`` or ``pyplot.get_cmap()`` instead.
cmap=plt.cm.get_cmap("bwr", 2),
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from matplotlib.colors import ListedColormap
from sklearn.datasets import load_diabetes
from sklearn.linear_model import LinearRegression
from sklearn.metrics import accuracy_score, confusion_matrix
# Load data
diabetes = load_diabetes()
df = pd.DataFrame(diabetes.data, columns=diabetes.feature_names)
df["target"] = diabetes.target
# Binary disease progression (0 = stable, 1 = worse)
threshold = np.median(df["target"])
df["progression_class"] = (df["target"] > threshold).astype(int)
# --- Function: scatter + regression boundary + accuracy ---
def plot_progression_with_regression(feature_x, feature_y, data=df):
X = data[[feature_x, feature_y]].values
y = data["progression_class"].values
# Train linear regression
regression = LinearRegression()
regression.fit(X, y)
b_1, b_2 = regression.coef_
b_0 = regression.intercept_
# Predict & classify
y_pred = regression.predict(X)
y_pred_class = (y_pred >= 0.5).astype(int)
# Accuracy & Confusion Matrix
acc = accuracy_score(y, y_pred_class)
cm = confusion_matrix(y, y_pred_class)
# Decision boundary: where prediction = 0.5
x1s = np.linspace(X[:, 0].min(), X[:, 0].max(), 100)
x2s = (0.5 - b_0 - b_1 * x1s) / b_2
# Scatterplot
discrete_cmap = ListedColormap(["xkcd:blue", "xkcd:red"])
plt.figure(figsize=(8, 6))
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=discrete_cmap, s=25, alpha=0.7)
plt.plot(x1s, x2s, color="gray", linestyle="--")
plt.fill_between(x1s, x2s, X[:, 1].max(), color="crimson", alpha=0.1)
plt.fill_between(x1s, x2s, X[:, 1].min(), color="blue", alpha=0.1)
plt.xlabel(feature_x)
plt.ylabel(feature_y)
plt.title(f"{feature_x} vs {feature_y}\nAccuracy = {acc:.2f}")
plt.grid()
plt.show()
print(f"Confusion Matrix ({feature_x} vs {feature_y}):\n{cm}")
print(f"Accuracy: {acc:.3f}")
# --- Example usage ---
# plot_progression_with_regression("bmi", "bp")
plot_progression_with_regression("age", "bmi")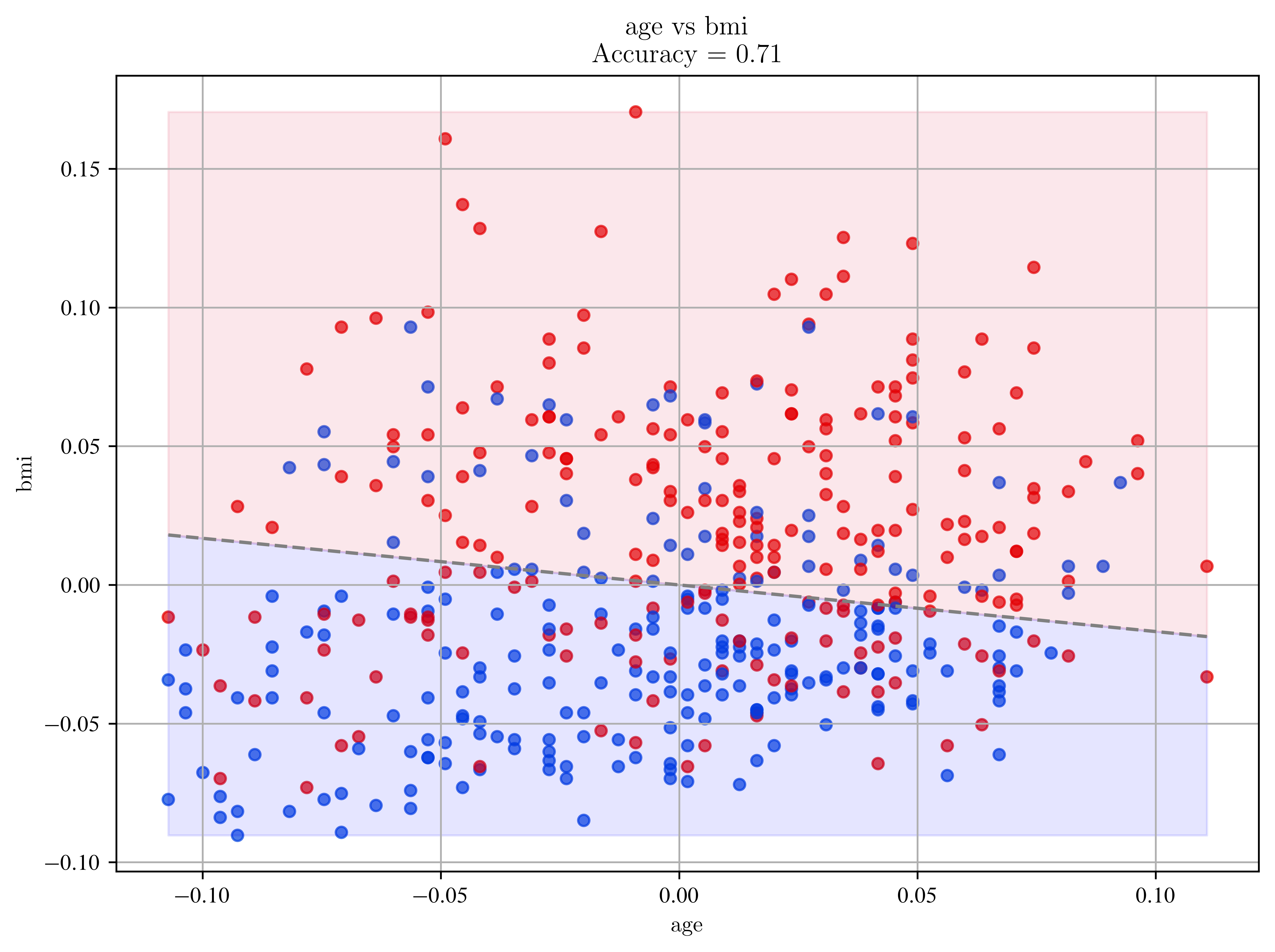
Confusion Matrix (age vs bmi):
[[168 53]
[ 75 146]]
Accuracy: 0.710
Idea of Least Squares: Minimisation of the sum of the differences between measurement and model = minimisation of the sum of the differences of the resiuals. It is assumed that individual measurements fluctuate with known variance and are unbiased (residual of measurement is zero)
General formulation of the Least Squares Method¶
Given a set of unbiased measurements , with known covariance matrix and a parametric model , an estimate for the parameters is obtained by the minimum of a function , defined as
or in matrix notation
where the weight matrix defines a metric for the difference between data and model.
Remarks
in many cases the preferred weight matrix is
here we look only at linear models , but also possible to use non-linear models
the cost function can be interpreted as a distance between data and model and the best fit parameters minimize the distance
when distances are measured in units of uncertainty the parameters have minimum variance
the derivation can be generalised from uncorrelated data points to correlated measurements with covariance matrix by the change
difference between loss and cost function: often used synonymous, but the loss function measures the error of a single training example, while the cost function is the average loss over the entire training dataset
Where to go from here?
the ‘bible’ of statistical learning by Hasties et al: https://
hastie .su .domains /ElemStatLearn/ (free eBook) toy dataset: https://
scikit -learn .org /stable /datasets /toy _dataset .html