1.1. Setup¶
First, we import the necessary libraries. We’ll be using numpy for numerical operations and matplotlib for plotting.
import time
import matplotlib.pyplot as plt
import numpy as np
# Set up plotting style
plt.style.use("seaborn-v0_8-whitegrid")1.2. Generate an Example Problem¶
As mentioned before, the CG method requires the matrix to be symmetric and positive-definite. A simple way to generate such a matrix is to create a random matrix and compute . This construction guarantees that is symmetric and positive semi-definite. If is non-singular, will be positive-definite.
We will set up a linear system where we know the true solution x_true beforehand. This allows us to easily verify the correctness of our implementations.
def generate_spd_matrix(n):
"""Generates a symmetric positive-definite matrix of size n x n."""
M = np.random.rand(n, n)
# Ensure M is well-conditioned enough to be non-singular
M += np.eye(n)
A = M.T @ M
return A
# Define the size of our problem. Feel free to try a few different values during the course of this exercise.
n = 1000
A = generate_spd_matrix(n)
x_true = np.random.rand(n)
# Calculate the right-hand side b
b = A @ x_true
print(
f"Generated a {A.shape[0]}x{A.shape[1]} symmetric, positive-definite matrix A and vector b."
)Generated a 1000x1000 symmetric, positive-definite matrix A and vector b.
1.3. The “Naive” Approach: Direct Solver¶
Before diving into iterative methods, let’s solve the system using a standard direct solver from NumPy, np.linalg.solve. This function typically uses an LU decomposition, which is very accurate and robust for small to moderately sized dense matrices.
print("Solving Ax = b using the direct method (np.linalg.solve)...")
# Time the direct solver
start_time = time.time()
x_naive = np.linalg.solve(A, b)
end_time = time.time()
time_naive = end_time - start_time
# Check the error
error_naive = np.linalg.norm(x_naive - x_true) / np.linalg.norm(x_true)
print(f"Direct solver finished in {time_naive:.4f} seconds.")
print(f"Relative error of the solution: {error_naive:.2e}")Solving Ax = b using the direct method (np.linalg.solve)...
Direct solver finished in 0.0261 seconds.
Relative error of the solution: 3.72e-08
This gives us a baseline for both speed and accuracy. For a 1000x1000 matrix, the direct solver should be quite fast, but as we scale up, its computational cost () and memory requirements () will become significant.
1.4. The Idea behind the Conjugate Gradient Method¶
The CG method combines the concept of steepest descent with Krylow subspaces:
First, we start at some initial point , and take a step in the direction of steepest descent.
This step takes place in the first, one-dimensional Krylow-subspace
Ideally, this step takes us the optimal amount of length along that direction
Then, for each further step, we find the new direction of maximum descent, but we only consider directions that do not undo previous progress
This means that the steps are -conjugate, instead of merely being orthogonal
The dimension of the Krylow-subspace increases with each step, so we require at most steps
This approach gives us the exact solution after at most steps, if we have perfect arithmetic. While we cannot assume this in practice, much fewer steps can give a good approximation.
1.5. The Conjugate Gradient Algorithm (Pseudo-code)¶
Now let’s look at the algorithm we want to implement. The goal is to find that minimizes .
Given the system , an initial guess , a tolerance tol, and a maximum number of iterations max_iter:
Initialize
Iterate while and :
Return
Key variables:
x: The solution vector, which we are improving in each step.r: The residual vector (), which we can use to compute the current error.p: The search direction vector. Note that we always move along the residual, here we choose special “-orthogonal” directions.alpha: The step size.beta: The improvement factor for the next search direction.
2. Exercise: CG Implementation¶
2.1. Implementing the Algorithm¶
Now it’s your turn! Fill in the missing lines in the conjugate_gradient function below. Follow the pseudo-code from the previous section.
Hint: The term appears multiple times. It’s efficient to compute it once per iteration and reuse the value.
def conjugate_gradient(A, b, x0=None, tol=1e-6, max_iter=1000):
"""
Solves the system Ax = b using the Conjugate Gradient method.
Args:
A (np.ndarray): The n x n symmetric positive-definite matrix.
b (np.ndarray): The n x 1 right-hand side vector.
x0 (np.ndarray, optional): Initial guess for the solution. Defaults to a zero vector.
tol (float, optional): The tolerance for convergence.
max_iter (int, optional): The maximum number of iterations.
Returns:
tuple: A tuple containing:
- x (np.ndarray): The solution vector.
- residuals (list): A list of the residual norm at each iteration.
"""
n = len(b)
# Initialize solution vector x
if x0 is None:
x = np.zeros(n)
else:
x = x0
# ---- Initialize r, p, and rs_old ---- #
# r is the residual (b - A*x)
# p is the search direction (initially same as r)
# rs_old is the dot product of the residual with itself (r^T * r)
r = b - (A @ x)
p = r.copy()
rs_old = r.T @ r
# -------------------------------------------- #
residuals = [np.sqrt(rs_old)]
if residuals[0] < tol:
print("Initial guess is already within tolerance.")
return x, residuals
for i in range(max_iter):
# ---- Main CG iteration steps ---- #
# 1. Calculate Ap = A * p
Ap = A @ p
# 2. Calculate step size alpha
alpha = rs_old / (p.T @ Ap)
# 3. Update solution: x_{k+1} = x_k + alpha * p_k
x = x + alpha * p
# 4. Update residual: r_{k+1} = r_k - alpha * Ap
r = r - alpha * Ap
# 5. Calculate new residual norm squared
rs_new = r.T @ r
# ------------------------------------- #
# Check for convergence
residual_norm = np.sqrt(rs_new)
residuals.append(residual_norm)
if residual_norm < tol:
print(f"Converged after {i + 1} iterations.")
break
# ---- Update search direction ---- #
# p_{k+1} = r_{k+1} + (rs_new / rs_old) * p_k
p = r + (rs_new / rs_old) * p
rs_old = rs_new
# ------------------------------------- #
else:
print(f"Did not converge within {max_iter} iterations.")
return x, residuals2.2. Test your Implementation¶
Let’s run your implementation, check that it works, and see how it performs compared to the direct solver.
Don’t worry if the code does not seem to converge, it is quite possible that you will have errors around e-2, even if you implemented everything correctly.
# Play around with n and tol a little and see what happens
n = 600
tol = 1.0e-6
A = generate_spd_matrix(n)
# As a sanity-check, you can also see if your code converges in the trivial case of A = Id
# A = np.eye(n)
x_true = np.random.rand(n)
# Calculate the right-hand side b
b = A @ x_true
print(
f"Generated a {A.shape[0]}x{A.shape[1]} symmetric, positive-definite matrix A and vector b."
)Generated a 600x600 symmetric, positive-definite matrix A and vector b.
# (Optional): Run the naive solver again, for time comparisons
print("Solving Ax = b using the direct method (np.linalg.solve)...")
# Time the direct solver
start_time = time.time()
x_naive = np.linalg.solve(A, b)
end_time = time.time()
time_naive = end_time - start_time
# Check the error
error_naive = np.linalg.norm(x_naive - x_true) / np.linalg.norm(x_true)
print(f"Direct solver finished in {time_naive:.4f} seconds.")
print(f"Relative error of the solution: {error_naive:.2e}")Solving Ax = b using the direct method (np.linalg.solve)...
Direct solver finished in 0.0044 seconds.
Relative error of the solution: 1.03e-08
print("Solving Ax = b using our own CG implementation...")
# Time the CG solver
start_time = time.time()
x_cg, residuals_cg = conjugate_gradient(A, b, tol=tol, max_iter=n)
end_time = time.time()
time_cg = end_time - start_time
# Check the error
error_cg = np.linalg.norm(x_cg - x_true) / np.linalg.norm(x_true)
print(f"\nCG solver finished in {time_cg:.4f} seconds.")
print(f"Relative error of the solution: {error_cg:.2e}")
print(f"Final residual norm: {residuals_cg[-1]:.2e}")
print(f"\n--- Comparison so far ---")
print(f"Direct solver time: {time_naive:.4f} s")
print(f"Simple CG time: {time_cg:.4f} s")Solving Ax = b using our own CG implementation...
Did not converge within 600 iterations.
CG solver finished in 0.0457 seconds.
Relative error of the solution: 1.94e-02
Final residual norm: 2.60e-02
--- Comparison so far ---
Direct solver time: 0.0044 s
Simple CG time: 0.0457 s
What is happening?
Is the result correct? (Compare
error_cgtoerror_naivefrom earlier).How does the computation time compare?
How many iterations did it take to converge? Did it converge at all?
Let’s plot the convergence history to see how the residual norm decreases over time.
plt.figure(figsize=(10, 6))
plt.semilogy(residuals_cg, marker="o", linestyle="-")
plt.title("CG Convergence History")
plt.xlabel("Iteration")
plt.ylabel("Norm of Residual (||r||)")
plt.grid(True, which="both", ls="--")
plt.show()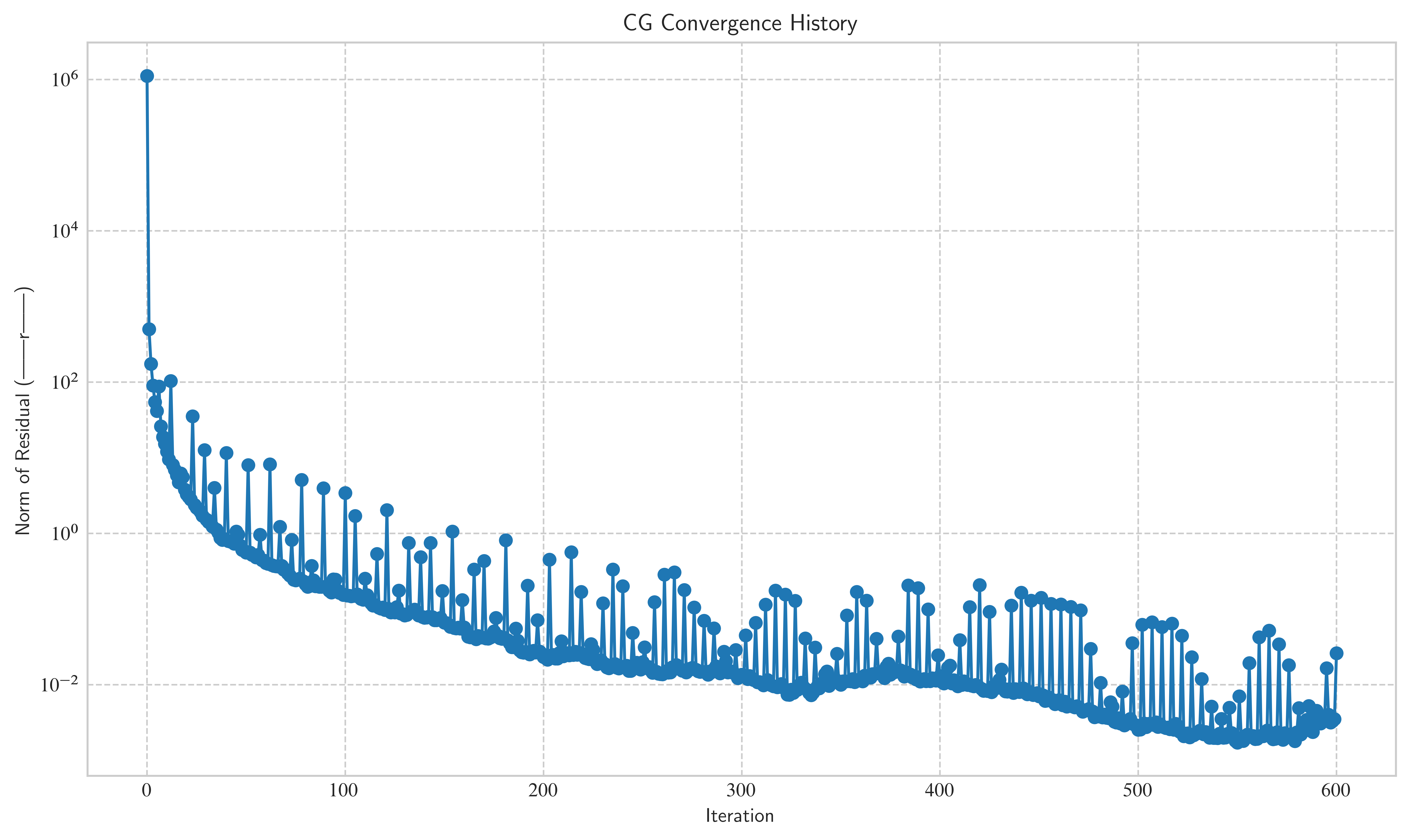
You will probably notice some things with your implementation:
The direct solver is usually way faster:
For one, CG’s strength lies in very large and/or sparse matrices.
Even more importantly, the CG method often converges in much fewer than the steps in practice
As long as our implementation does not converge at most of the time, it is nowhere near as efficient as it could be!
For any non-trivial example () and reasonable error tolerances, the algorithm may not converge: You can see in the plot, that the method has a hard time getting below an error tolerance of 1.e-2.
The algorithm is not guaranteed to converge after steps (you can check this by setting
max-iterto a value higher thann): Again, this is due to the lack of perfectly accurate arithmetic.
As you can see, the method is a lot worse than we would have hoped. In the second part of this exercise, we will take a look at preconditioners to speed up convergence and improve these results.
Bonus exercise¶
If you are finished before the break, have a think about how we could verify our assumption, before implementing preconditioning. Are there some easy ways to modify the construction of (i.e. our generate_spd_matrix method) to make sure it is well-conditioned?
Answer: By increasing the factor in M += np.eye(n) to e.g. M += 100 * np.eye(n), we can make the matrix diagonally dominant. This gives us a well-conditioned matrix, that our simple algorithm should be able to handle well. However, this is only helpful for demonstration purposes, if we really want to solve an ill-conditioned system, we need to go about it in a different way.
3. Preconditioning¶
3.1. The Condition Number¶
The number of iterations CG takes to converge is closely related to the condition number of the matrix , denoted .
(for symmetric positive-definite matrices)
A high condition number means the matrix is “ill-conditioned”. This has two major negative effects:
Error Amplification: Small errors in the input
bcan lead to large errors in the outputx.Slow Convergence: Iterative methods like CG converge very slowly. The number of iterations is roughly proportional to .
Let’s check the condition number of our matrix A.
# Try this with different sizes of A
cond_A = np.linalg.cond(A)
print(f"The condition number of A is: {cond_A:.2e}")The condition number of A is: 8.75e+09
That’s a pretty large number! This explains why our simple CG implementation might have taken many iterations to converge. To fix this, we can use preconditioning.
3.2. Preconditioning Theory¶
The idea is to find a preconditioner matrix that is:
Also symmetric and positive-definite.
A good approximation of .
Easy to invert.
Instead of solving , we solve a modified system. The most common form is left preconditioning, where we solve .
Note: Even if and are symmetric, the new system matrix is generally not symmetric! This means we cannot apply the standard CG algorithm directly to it. The Preconditioned Conjugate Gradient method is a clever variant that is mathematically equivalent to running CG on a transformed, symmetric system, without having to explicitly form .
Our goal is to find a such that .
The simplest (but often effective) preconditioner is the Jacobi Preconditioner (or Diagonal Preconditioner). We choose to be the diagonal of .
Since is symmetric positive-definite, all its diagonal entries are positive, which means is also positive-definite. Inverting this is also trivial, as is just a diagonal matrix with entries .
3.3. Implementing the Jacobi Preconditioner¶
For this part, we will first modify our matrix to generate a new, ill-conditioned matrix that work well with Jacobi preconditioning.
# A = np.array([[-1, 2, 3], [3, -4, 5], [-6, 7, 8]])
# print(A)
# print(np.abs(np.diag(A)))
# print(np.sum(np.abs(A), axis=1))
# print(np.logspace(0, np.log10(1e10), 5))
# ?np.fill_diagonaldef make_diagonally_dominant(A, diag_cond_num=1e15):
n = A.shape[0]
row_sums = np.sum(np.abs(A), axis=1) - np.abs(np.diag(A))
ill_conditioned_part = np.logspace(0, np.log10(diag_cond_num), n)
np.random.shuffle(ill_conditioned_part)
final_diag = ill_conditioned_part + row_sums
np.fill_diagonal(A, final_diag)
returnmake_diagonally_dominant(A)
cond_A = np.linalg.cond(A)
print(f"The condition number of A is now: {cond_A:.2e}")
x_true = np.random.rand(n)
b = A @ x_trueThe condition number of A is now: 1.19e+10
As for the obvious question: Why bother? The Jacobi preconditioner works well for ill-conditioned, but diagonally dominant matrices, where the main source of issues comes from the diagonal. So this modification is only for demonstrative purposes, but we will get back to the original matrix later.
Let’s now create our preconditioner. We will actually compute its inverse, since that’s what the algorithm needs.
# Example of np.diag
matrix = np.array([[1, 2, 3], [3, 4, 5], [5, 6, 8]])
print(1 / np.diag(matrix))
print(np.diag(np.diag(matrix)))
print(np.diag(1 / (np.diag(matrix))))[1. 0.25 0.125]
[[1 0 0]
[0 4 0]
[0 0 8]]
[[1. 0. 0. ]
[0. 0.25 0. ]
[0. 0. 0.125]]
# TODO: Create the inverse of the Jacobi preconditioner, P_inv.
# P is a diagonal matrix containing the diagonal of A.
# P_inv is a diagonal matrix containing the reciprocal of the diagonal of A.
# HINT: There is a function called np.diag()
P_inv = np.diag(1 / (np.diag(A)))
# Let's check the condition number of the preconditioned system
cond_PA = np.linalg.cond(P_inv @ A)
print("--- Condition Numbers ---")
print(f"Original matrix A: {cond_A:.2e}")
print(f"Preconditioned P_inv*A: {cond_PA:.2e}")
print(f"Improvement factor: {cond_A / cond_PA:.2e} times better")--- Condition Numbers ---
Original matrix A: 1.19e+10
Preconditioned P_inv*A: 1.60e+00
Improvement factor: 7.45e+09 times better
As you can see, the condition number has been significantly reduced! This should lead to much faster convergence.
3.4. The Preconditioned Conjugate Gradients Algorithm¶
The algorithm is slightly modified to include the preconditioner. The key step is to solve the system for (which is easy since we have ).
Initialize
Solve (i.e., )
Iterate:
Check for convergence on
Solve
Now, implement the PCG algorithm below.
def preconditioned_conjugate_gradient(A, b, P_inv, x0=None, tol=1e-6, max_iter=1000):
"""
Solves the system Ax = b using the Preconditioned Conjugate Gradient method.
"""
n = len(b)
if x0 is None:
x = np.zeros(n)
else:
x = x0
residuals = []
# ---- TODO: Initialize r, z, p, and rz_old ---- #
r = b - (A @ x)
residuals.append(np.linalg.norm(r))
if residuals[0] < tol:
return x, residuals
# z is the result of applying the preconditioner to r
z = P_inv @ r
# p is the search direction
p = z.copy() # YOUR CODE HERE
# rz_old is the dot product of r and z
rz_old = r.T @ z
# ----------------------------------------------- #
for i in range(max_iter):
# ---- TODO: Main PCG iteration steps ---- #
# 1. Calculate Ap = A * p
Ap = A @ p
# 2. Calculate step size alpha
alpha = rz_old / (p.T @ Ap)
# 3. Update solution: x_{k+1} = x_k + alpha * p_k
x = x + alpha * p
# 4. Update residual: r_{k+1} = r_k - alpha * Ap
r = r - alpha * Ap
# Check for convergence
residual_norm = np.linalg.norm(r)
residuals.append(residual_norm)
if residual_norm < tol:
print(f"Converged after {i + 1} iterations.")
break
# 5. Apply preconditioner for the next step
z = P_inv @ r
# 6. Calculate rz_new
rz_new = r.T @ z
# 7. Update search direction
p = z + (rz_new / rz_old) * p
rz_old = rz_new
# -------------------------------------- #
else:
print(f"Did not converge within {max_iter} iterations.")
return x, residuals3.5. Testing the PCG Algorithm¶
Let’s run our new PCG implementation and see the difference.
# (Optional): Run the naive solver again, for time comparisons
print("Solving Ax = b using the direct method (np.linalg.solve)...")
# Time the direct solver
start_time = time.time()
x_naive = np.linalg.solve(A, b)
end_time = time.time()
time_naive = end_time - start_time
# Check the error
error_naive = np.linalg.norm(x_naive - x_true) / np.linalg.norm(x_true)
print("Solving Ax = b using our own CG implementation...")
# Time the CG solver
start_time = time.time()
x_cg, residuals_cg = conjugate_gradient(A, b, tol=tol, max_iter=n)
end_time = time.time()
time_cg = end_time - start_time
# Check the error
error_cg = np.linalg.norm(x_cg - x_true) / np.linalg.norm(x_true)Solving Ax = b using the direct method (np.linalg.solve)...
Solving Ax = b using our own CG implementation...
Did not converge within 600 iterations.
print("Solving Ax = b using our own PCG implementation...")
# Time the PCG solver
start_time = time.time()
x_pcg, residuals_pcg = preconditioned_conjugate_gradient(A, b, P_inv, max_iter=n)
end_time = time.time()
time_pcg = end_time - start_time
# Check the error
error_pcg = np.linalg.norm(x_pcg - x_true) / np.linalg.norm(x_true)
print(f"\nPCG solver finished in {time_pcg:.4f} seconds.")
print(f"Relative error of the solution: {error_pcg:.2e}")
print(f"Final residual norm: {residuals_pcg[-1]:.2e}")
print(f"\n--- Comparison times ---")
print(f"Direct solver time: {time_naive:.4f} s")
print(f"Simple CG time: {time_cg:.4f} s")
print(f"PCG time: {time_pcg:.4f} s")
print(f"\n--- Comparison errors ---")
print(f"Direct solver error: {error_naive:.2e}")
print(f"Simple CG error: {error_cg:.2e}")
print(f"PCG error: {error_pcg:.2e}")Solving Ax = b using our own PCG implementation...
Converged after 6 iterations.
PCG solver finished in 0.0045 seconds.
Relative error of the solution: 8.33e-16
Final residual norm: 8.63e-10
--- Comparison times ---
Direct solver time: 0.0072 s
Simple CG time: 0.0495 s
PCG time: 0.0045 s
--- Comparison errors ---
Direct solver error: 1.40e-15
Simple CG error: 7.50e-01
PCG error: 8.33e-16
As we can see, the improvement is dramatic and the new algorithm converges after only a handful of steps. Of course, this particular example is designed to show drastic effects, and it will typically not go quite as well in practice, but you get the idea.
Open-ended exercise: Explore the limitations of the methods we implemented¶
Just like the first part, take some time to try different values or matrices, and get a feel for the limitations of the algorithm
What happens, if you go back to the first type of matrices that we used, and apply the Jacobi preconditioner?
Does the result improve?
What may be the reason for its behaviour?
What might a suitable preconditioner for this type of matrix look like?
What happens if you try to apply the CG method to a fully random matrix?
You could also try to come up with other kinds of matrix and see where the Jacobi preconditioner starts to fail
Outlook¶
As you’ve seen in this exercise, the CG method can be a very elegant and efficient approach to solving many issues, if they have the right properties. But just as there were many cases, where it brought about a significant performance benefit, there are also many situations, where it is either unsuitable, or the preconditioner needs to be carefully chosen.
Always keep in mind: Just because you have a hammer, not every problem is a nail.
If you are interested in further reading, you can move on to more advanced topics, such as:
GMRES: The Generalized Minimal Residual applies the ideas of CG in a more general setting. It is on of the standard approaches if you are faced with a non-symmetric matrix . It comes with some drawbacks such as increased memory requirements, though.
Other preconditioners: There are a multitude of preconditioners, and the best choice always depends on the problem you work with. Typical ones include
Gauss-Seidel: It uses the lower triangular part of , and is relatively difficult to parallelize
(Symmetric) Succesive Over-Relaxation (SORR): This method is based on Gauss-Seidel, but slightly more involved.