import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
from matplotlib import cm
from mpl_toolkits.mplot3d import Axes3D
# Set random seeds for reproducibility
torch.manual_seed(42)
np.random.seed(42)
# Device configuration
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# device = torch.device('mps' if torch.backends.mps.is_available() else device)
print(f"Using device: {device}")Using device: cpu
class PINN(nn.Module):
"""
Physics-Informed Neural Network for solving PDEs.
The network takes (x, t) as input and outputs u(x, t).
"""
def __init__(self, layers):
super(PINN, self).__init__()
self.layers = nn.ModuleList()
# Build the network
for i in range(len(layers) - 1):
self.layers.append(nn.Linear(layers[i], layers[i + 1]))
# Initialize weights using Xavier initialization
self.init_weights()
def init_weights(self):
for layer in self.layers:
nn.init.xavier_normal_(layer.weight)
nn.init.zeros_(layer.bias)
def forward(self, x, t):
# Concatenate inputs
inputs = torch.cat([x, t], dim=1)
# Pass through hidden layers with tanh activation
for i in range(len(self.layers) - 1):
inputs = torch.tanh(self.layers[i](inputs))
# Output layer (no activation)
output = self.layers[-1](inputs)
return outputdef compute_pde_residual(model, x, t, nu=0.01):
"""
Compute the PDE residual for the heat equation.
PDE: du/dt - alpha * d2u/dx2 = 0
"""
x.requires_grad = True
t.requires_grad = True
# Forward pass
u = model(x, t)
# Compute gradients
u_x = torch.autograd.grad(
u, x, grad_outputs=torch.ones_like(u), create_graph=True, retain_graph=True
)[0]
u_t = torch.autograd.grad(
u, t, grad_outputs=torch.ones_like(u), create_graph=True, retain_graph=True
)[0]
# Second derivative with respect to x
u_xx = torch.autograd.grad(
u_x, x, grad_outputs=torch.ones_like(u_x), create_graph=True, retain_graph=True
)[0]
# PDE residual
residual = u_t + u * u_x - nu * u_xx
return residual
def initial_condition(x):
"""
Initial condition: u(x, 0) = sin(pi * x)
"""
return torch.sin(np.pi * x)
def boundary_condition(t):
"""
Boundary conditions: u(0, t) = u(1, t) = 0
"""
return torch.zeros_like(t)def train_pinn(
model,
n_epochs=5000,
lr=0.001,
n_collocation=10000,
n_initial=100,
n_boundary=100,
nu=0.01,
):
"""
Train the Physics-Informed Neural Network.
Args:
model: PINN model
n_epochs: Number of training epochs
lr: Learning rate
n_collocation: Number of collocation points for PDE
n_initial: Number of points for initial condition
n_boundary: Number of points for boundary conditions
"""
optimizer = torch.optim.Adam(model.parameters(), lr=lr)
losses = []
for epoch in range(n_epochs):
optimizer.zero_grad()
# Generate collocation points for PDE (random points in domain)
x_pde = torch.rand(n_collocation, 1, requires_grad=True).to(device)
t_pde = torch.rand(n_collocation, 1, requires_grad=True).to(device)
# Initial condition points (t=0)
x_ic = torch.rand(n_initial, 1).to(device)
t_ic = torch.zeros(n_initial, 1).to(device)
u_ic = initial_condition(x_ic)
# Boundary condition points (x=0 and x=1)
t_bc = torch.rand(n_boundary, 1).to(device)
x_bc_left = torch.zeros(n_boundary, 1).to(device)
x_bc_right = torch.ones(n_boundary, 1).to(device)
u_bc = boundary_condition(t_bc)
# Compute losses
# 1. PDE residual loss
residual = compute_pde_residual(model, x_pde, t_pde, alpha)
loss_pde = torch.mean(residual**2)
# 2. Initial condition loss
u_pred_ic = model(x_ic, t_ic)
loss_ic = torch.mean((u_pred_ic - u_ic) ** 2)
# 3. Boundary condition loss
u_pred_bc_left = model(x_bc_left, t_bc)
u_pred_bc_right = model(x_bc_right, t_bc)
loss_bc = torch.mean((u_pred_bc_left - u_bc) ** 2) + torch.mean(
(u_pred_bc_right - u_bc) ** 2
)
# Total loss
loss = loss_pde + loss_ic + loss_bc
# Backpropagation
loss.backward()
optimizer.step()
losses.append(loss.item())
# Print progress
if (epoch + 1) % 500 == 0:
print(
f"Epoch [{epoch + 1}/{n_epochs}], Loss: {loss.item():.6f}, "
f"PDE: {loss_pde.item():.6f}, IC: {loss_ic.item():.6f}, "
f"BC: {loss_bc.item():.6f}"
)
return lossesimport numpy as np
from scipy.special import iv
def analytical_solution(x, t, nu, n_terms=100):
"""
Computes the analytical solution to the 1D viscous Burgers' equation
using the Cole-Hopf transformation.
Parameters:
-----------
x : array_like or float
Spatial coordinate(s) in the domain [0, 1].
t : float
Time point (t > 0).
nu : float
Viscosity coefficient.
n_terms : int, optional
Number of terms to include in the Fourier series summation.
Default is 100. Increase for higher accuracy at small t or small nu.
Returns:
--------
u : ndarray or float
The velocity u(x, t).
"""
# Argument for the Bessel functions
z = 1.0 / (2.0 * np.pi * nu)
# Initialize numerator and denominator sums
# Denominator starts with I_0(z) (the n=0 term)
denominator = iv(0, z)
numerator = 0.0
# Calculate the infinite series (truncated at n_terms)
for n in range(1, n_terms + 1):
# Temporal decay term
decay = np.exp(-(n**2) * np.pi**2 * nu * t)
# Bessel function term I_n(z)
bessel_val = iv(n, z)
# Accumulate sums
numerator += n * bessel_val * np.sin(n * np.pi * x) * decay
denominator += 2 * bessel_val * np.cos(n * np.pi * x) * decay
# Final computation of u(x,t)
u = (4 * np.pi * nu * numerator) / denominator
return u6. Train the Model¶
# Define network architecture
layers = [2, 32, 32, 32, 1] # Input: (x, t), Output: u
# Create model
model = PINN(layers).to(device)
print(model)
# Thermal diffusivity
alpha = 1
nu = 0.01
# Train the model
print("\nTraining PINN...")
losses = train_pinn(model, n_epochs=10000, lr=0.001, nu=nu)PINN(
(layers): ModuleList(
(0): Linear(in_features=2, out_features=32, bias=True)
(1-2): 2 x Linear(in_features=32, out_features=32, bias=True)
(3): Linear(in_features=32, out_features=1, bias=True)
)
)
Training PINN...
Epoch [500/10000], Loss: 0.073245, PDE: 0.009610, IC: 0.043980, BC: 0.019655
Epoch [1000/10000], Loss: 0.011490, PDE: 0.005489, IC: 0.003995, BC: 0.002006
Epoch [1500/10000], Loss: 0.002374, PDE: 0.001594, IC: 0.000334, BC: 0.000446
Epoch [2000/10000], Loss: 0.000737, PDE: 0.000560, IC: 0.000073, BC: 0.000104
Epoch [2500/10000], Loss: 0.000351, PDE: 0.000279, IC: 0.000019, BC: 0.000054
Epoch [3000/10000], Loss: 0.000217, PDE: 0.000176, IC: 0.000013, BC: 0.000028
Epoch [3500/10000], Loss: 0.000387, PDE: 0.000322, IC: 0.000015, BC: 0.000050
Epoch [4000/10000], Loss: 0.000403, PDE: 0.000331, IC: 0.000013, BC: 0.000058
Epoch [4500/10000], Loss: 0.000111, PDE: 0.000083, IC: 0.000011, BC: 0.000017
Epoch [5000/10000], Loss: 0.000098, PDE: 0.000067, IC: 0.000017, BC: 0.000014
Epoch [5500/10000], Loss: 0.000349, PDE: 0.000296, IC: 0.000029, BC: 0.000024
Epoch [6000/10000], Loss: 0.000128, PDE: 0.000099, IC: 0.000008, BC: 0.000021
Epoch [6500/10000], Loss: 0.000408, PDE: 0.000360, IC: 0.000009, BC: 0.000039
Epoch [7000/10000], Loss: 0.000132, PDE: 0.000092, IC: 0.000024, BC: 0.000016
Epoch [7500/10000], Loss: 0.000110, PDE: 0.000072, IC: 0.000027, BC: 0.000010
Epoch [8000/10000], Loss: 0.000131, PDE: 0.000106, IC: 0.000007, BC: 0.000018
Epoch [8500/10000], Loss: 0.000241, PDE: 0.000185, IC: 0.000010, BC: 0.000046
Epoch [9000/10000], Loss: 0.000094, PDE: 0.000077, IC: 0.000007, BC: 0.000010
Epoch [9500/10000], Loss: 0.000051, PDE: 0.000036, IC: 0.000007, BC: 0.000008
Epoch [10000/10000], Loss: 0.000049, PDE: 0.000032, IC: 0.000009, BC: 0.000008
plt.figure(figsize=(10, 6))
plt.plot(losses)
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.title("Training Loss over Epochs")
plt.yscale("log")
plt.grid(True)
plt.show()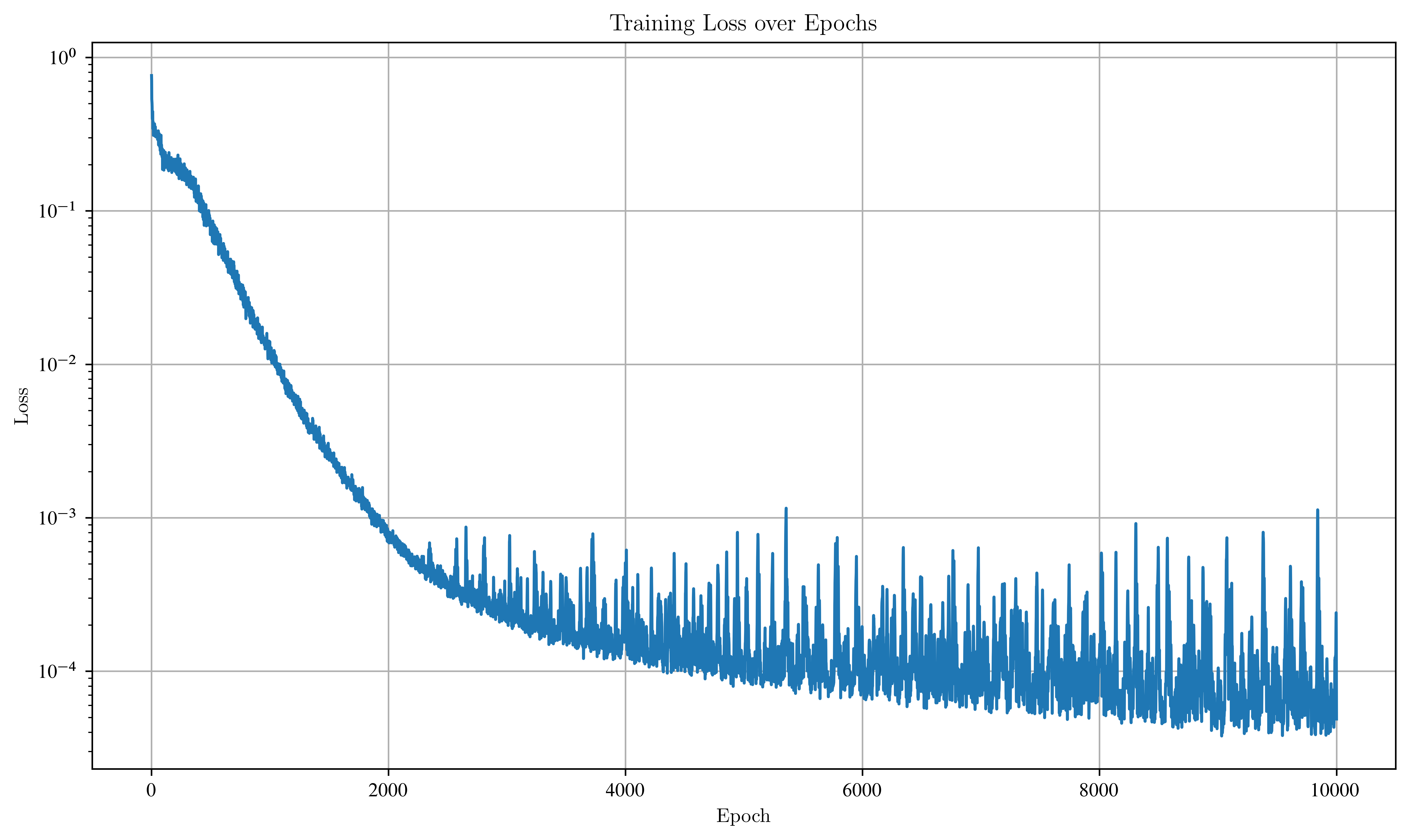
Evaluate and Compare with Analytical Solution¶
# Create a grid for evaluation
x_test = np.linspace(0, 1, 100)
t_test = np.linspace(0, 1, 100)
X, T = np.meshgrid(x_test, t_test)
# Flatten for prediction
x_flat = X.flatten()[:, None]
t_flat = T.flatten()[:, None]
# Convert to tensors
x_tensor = torch.FloatTensor(x_flat).to(device)
t_tensor = torch.FloatTensor(t_flat).to(device)
# Predict with PINN
model.eval()
with torch.no_grad():
u_pred = model(x_tensor, t_tensor).cpu().numpy()
U_pred = u_pred.reshape(X.shape)
# Analytical solution
U_exact = analytical_solution(X, T, alpha)
# Compute error
error = np.abs(U_pred - U_exact)
relative_error = np.linalg.norm(error) / np.linalg.norm(U_exact)
print(f"\nRelative L2 error: {relative_error:.6f}")
Relative L2 error: 0.007149
Visualize Results¶
# 3D surface plots
fig = plt.figure(figsize=(18, 5))
# PINN prediction
ax1 = fig.add_subplot(131, projection="3d")
surf1 = ax1.plot_surface(X, T, U_pred, cmap=cm.viridis, alpha=0.8)
ax1.set_xlabel("x")
ax1.set_ylabel("t")
ax1.set_zlabel("u(x,t)")
ax1.set_title("PINN Prediction")
fig.colorbar(surf1, ax=ax1, shrink=0.5)
# Analytical solution
ax2 = fig.add_subplot(132, projection="3d")
surf2 = ax2.plot_surface(X, T, U_exact, cmap=cm.viridis, alpha=0.8)
ax2.set_xlabel("x")
ax2.set_ylabel("t")
ax2.set_zlabel("u(x,t)")
ax2.set_title("Analytical Solution")
fig.colorbar(surf2, ax=ax2, shrink=0.5)
# Error
ax3 = fig.add_subplot(133, projection="3d")
surf3 = ax3.plot_surface(X, T, error, cmap=cm.hot, alpha=0.8)
ax3.set_xlabel("x")
ax3.set_ylabel("t")
ax3.set_zlabel("|Error|")
ax3.set_title("Absolute Error")
fig.colorbar(surf3, ax=ax3, shrink=0.5)
plt.tight_layout()
plt.show()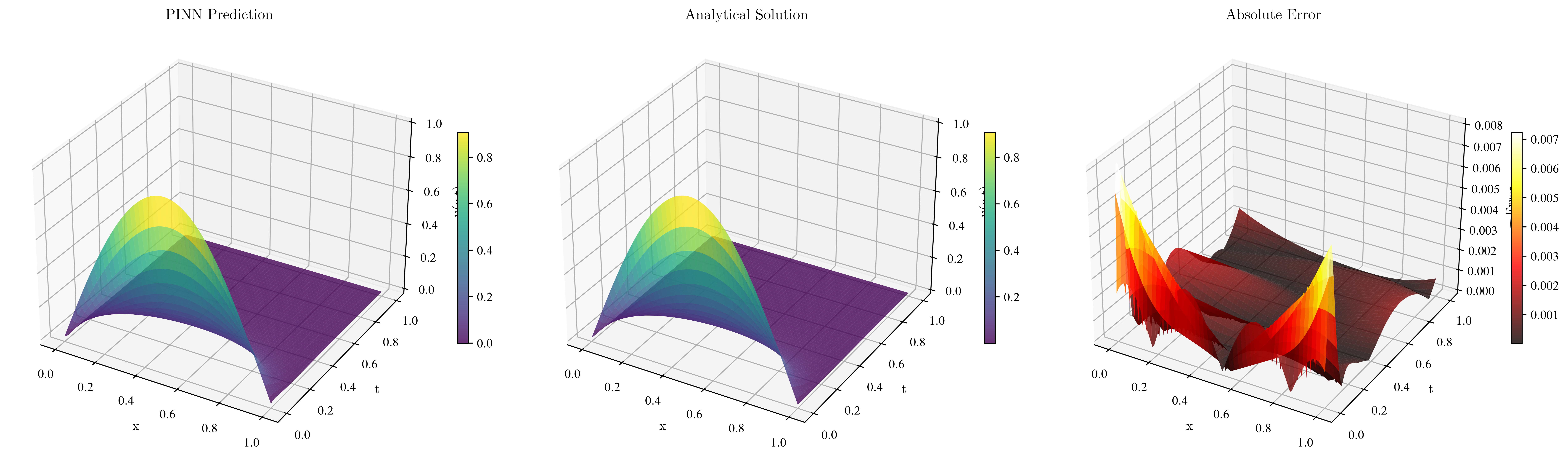
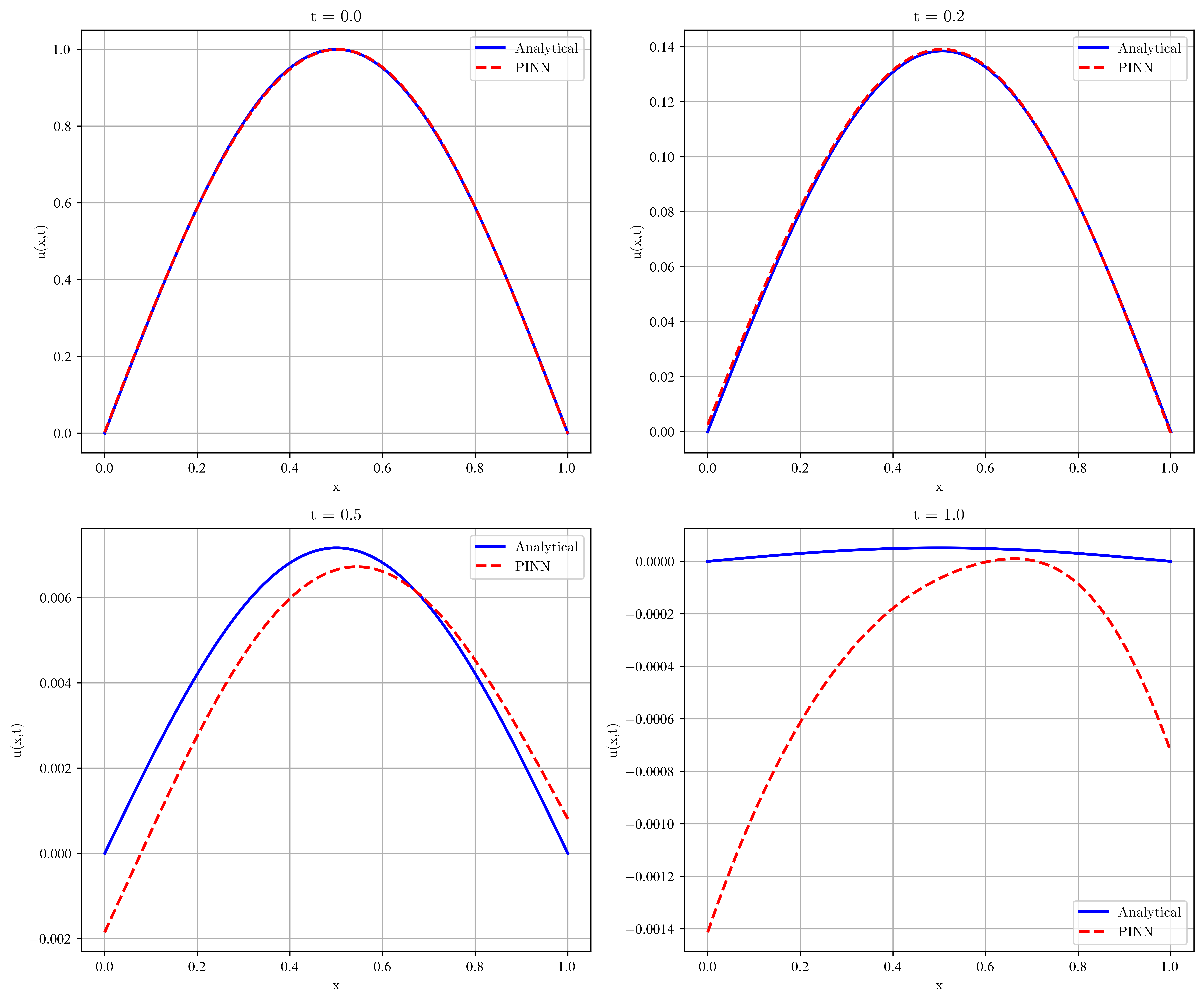
# Compare solutions at different time points
time_points = [0.0, 0.2, 0.5, 1.0]
x_plot = np.linspace(0, 1, 100)
fig, axes = plt.subplots(2, 2, figsize=(12, 10))
axes = axes.flatten()
for idx, t_point in enumerate(time_points):
# Prepare data
t_plot = np.ones_like(x_plot) * t_point
x_tensor = torch.FloatTensor(x_plot[:, None]).to(device)
t_tensor = torch.FloatTensor(t_plot[:, None]).to(device)
# PINN prediction
with torch.no_grad():
u_pred = model(x_tensor, t_tensor).cpu().numpy().flatten()
# Analytical solution
u_exact = analytical_solution(x_plot, t_point, alpha)
# Plot
axes[idx].plot(x_plot, u_exact, "b-", label="Analytical", linewidth=2)
axes[idx].plot(x_plot, u_pred, "r--", label="PINN", linewidth=2)
axes[idx].set_xlabel("x")
axes[idx].set_ylabel("u(x,t)")
axes[idx].set_title(f"t = {t_point}")
axes[idx].legend()
axes[idx].grid(True)
plt.tight_layout()
plt.show()