import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
from matplotlib import cm
from mpl_toolkits.mplot3d import Axes3D
# Set random seeds for reproducibility
torch.manual_seed(42)
np.random.seed(42)
# Device configuration
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# device = torch.device('mps' if torch.backends.mps.is_available() else device)
print(f"Using device: {device}")Using device: cpu
2. Lotka-volterra¶
class PINN(nn.Module):
"""
Physics-Informed Neural Network for solving PDEs.
The network takes (x, t) as input and outputs u(x, t).
"""
def __init__(self, layers):
super(PINN, self).__init__()
self.layers = nn.ModuleList()
# Build the network
for i in range(len(layers) - 1):
self.layers.append(nn.Linear(layers[i], layers[i + 1]))
# Initialize weights using Xavier initialization
self.init_weights()
def init_weights(self):
for layer in self.layers:
nn.init.xavier_normal_(layer.weight)
nn.init.zeros_(layer.bias)
def forward(self, t):
inputs = t
# Pass through hidden layers with tanh activation
for i in range(len(self.layers) - 1):
inputs = torch.tanh(self.layers[i](inputs))
# Output layer (no activation)
output = self.layers[-1](inputs)
return outputdef compute_pde_residual(model, t, alpha, beta, gamma, delta):
"""
Lotka-Volterra equations residual computation.
Standard form:
dx/dt = alpha*x - beta*x*y (prey: growth - predation)
dy/dt = delta*x*y - gamma*y (predator: growth from eating - death)
"""
t.requires_grad = True
# Forward pass
xy = model(t)
x = xy[:, 0:1] # Keep dimensions for proper broadcasting
y = xy[:, 1:2]
# Compute gradients
x_t = torch.autograd.grad(
x, t, grad_outputs=torch.ones_like(x), create_graph=True, retain_graph=True
)[0]
y_t = torch.autograd.grad(
y, t, grad_outputs=torch.ones_like(y), create_graph=True, retain_graph=True
)[0]
# Residuals for the system of ODEs
residual_x = x_t - (alpha * x - beta * x * y)
residual_y = y_t - (delta * x * y - gamma * y)
return residual_x, residual_y
def initial_condition(device=device):
# Initial conditions at t=0 for (x=prey, y=predator)
# Starting with x=40, y=9 (away from equilibrium to see oscillations)
return torch.tensor([[40.0, 9.0]], device=device)def train_pinn(
model,
n_epochs=5000,
lr=0.001,
n_collocation=1000,
alpha=0.01,
beta=0.01,
gamma=0.01,
delta=0.01,
tMax=50,
ic_weight=10.0,
use_causal=True,
):
"""
Train the Physics-Informed Neural Network.
Args:
model: PINN model
n_epochs: Number of training epochs
lr: Learning rate
n_collocation: Number of collocation points for ODE
alpha, beta, gamma, delta: Lotka-Volterra parameters
tMax: Maximum time for sampling collocation points
ic_weight: Weight for initial condition loss
use_causal: Whether to use causal weighting (prioritize early times)
"""
optimizer = torch.optim.Adam(model.parameters(), lr=lr)
# Use CosineAnnealingWarmRestarts - LR recovers periodically instead of going to zero
scheduler = torch.optim.lr_scheduler.CosineAnnealingWarmRestarts(
optimizer,
T_0=1000, # Restart every 1000 epochs
T_mult=2, # Double the period after each restart
eta_min=1e-6, # Minimum LR
)
losses = []
for epoch in range(n_epochs):
optimizer.zero_grad()
# Generate collocation points for ODE (random points in [0, tMax])
t_pde = (tMax * torch.rand(n_collocation, 1)).to(device)
# Initial condition at t = 0
u_ic = initial_condition() # (1, 2) on device
# 1. PDE residual loss with causal weighting
residual_x, residual_y = compute_pde_residual(
model, t_pde, alpha, beta, gamma, delta
)
if use_causal:
# Causal weighting: earlier times have higher weight
# This helps because solution at later times depends on earlier times
causal_weights = torch.exp(-t_pde / tMax * 2.0) # Exponential decay
loss_pde = torch.mean(causal_weights * residual_x**2) + torch.mean(
causal_weights * residual_y**2
)
else:
loss_pde = torch.mean(residual_x**2) + torch.mean(residual_y**2)
# 2. Initial condition loss (weighted more heavily)
u_pred_ic = model(torch.zeros(1, 1).to(device)) # (1, 2)
loss_ic = torch.mean((u_pred_ic - u_ic) ** 2)
# Total loss with IC weighting
loss = loss_pde + ic_weight * loss_ic
# Backpropagation
loss.backward()
# Gradient clipping for stability
torch.nn.utils.clip_grad_norm_(model.parameters(), max_norm=1.0)
optimizer.step()
scheduler.step()
losses.append(loss.item())
# Print progress
if (epoch + 1) % 500 == 0:
current_lr = optimizer.param_groups[0]["lr"]
print(
f"Epoch [{epoch + 1}/{n_epochs}], Loss: {loss.item():.6f}, "
f"PDE: {loss_pde.item():.6f}, IC: {loss_ic.item():.6f}, LR: {current_lr:.6f}"
)
return losses# Define network architecture - larger network for oscillatory dynamics
layers = [1, 64, 64, 64, 64, 2] # Input: (t), Output: (x, y)
# Create model
model = PINN(layers).to(device)
print(model)PINN(
(layers): ModuleList(
(0): Linear(in_features=1, out_features=64, bias=True)
(1-3): 3 x Linear(in_features=64, out_features=64, bias=True)
(4): Linear(in_features=64, out_features=2, bias=True)
)
)
# Classic Lotka-Volterra parameters for nice oscillations
# Equilibrium point: x* = gamma/delta = 20, y* = alpha/beta = 10
alpha = 1.0 # prey birth rate
beta = 0.1 # predation rate
gamma = 1.5 # predator death rate
delta = 0.075 # predator birth rate (from eating prey)
# Time horizon - period ≈ 2π/√(α*γ) ≈ 5.1, so tMax=30 gives ~6 cycles
tMax = 30
# Train the model
print("\nTraining PINN...")
print(f"Parameters: α={alpha}, β={beta}, γ={gamma}, δ={delta}")
print(f"Equilibrium point: x*={gamma / delta:.1f}, y*={alpha / beta:.1f}")
print(f"Initial conditions: x(0)=40, y(0)=9\n")
losses = train_pinn(
model,
n_epochs=5000,
lr=0.001,
n_collocation=2000,
alpha=alpha,
beta=beta,
gamma=gamma,
delta=delta,
tMax=tMax,
ic_weight=10.0,
use_causal=True,
)
Training PINN...
Parameters: α=1.0, β=0.1, γ=1.5, δ=0.075
Equilibrium point: x*=20.0, y*=10.0
Initial conditions: x(0)=40, y(0)=9
Epoch [500/5000], Loss: 15.467306, PDE: 5.715391, IC: 0.975191, LR: 0.000501
Epoch [1000/5000], Loss: 3.069421, PDE: 3.041733, IC: 0.002769, LR: 0.001000
Epoch [1500/5000], Loss: 0.946446, PDE: 0.946018, IC: 0.000043, LR: 0.000854
Epoch [2000/5000], Loss: 0.487639, PDE: 0.484973, IC: 0.000267, LR: 0.000501
Epoch [2500/5000], Loss: 0.448370, PDE: 0.448359, IC: 0.000001, LR: 0.000147
Epoch [3000/5000], Loss: 0.434111, PDE: 0.434074, IC: 0.000004, LR: 0.001000
Epoch [3500/5000], Loss: 0.419497, PDE: 0.393471, IC: 0.002603, LR: 0.000962
Epoch [4000/5000], Loss: 0.381792, PDE: 0.378299, IC: 0.000349, LR: 0.000854
Epoch [4500/5000], Loss: 0.372748, PDE: 0.371663, IC: 0.000108, LR: 0.000692
Epoch [5000/5000], Loss: 0.387470, PDE: 0.387407, IC: 0.000006, LR: 0.000501
# see solution
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 6))
plt.plot(losses)
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.title("Training Loss over Epochs")
plt.yscale("log")
plt.grid(True)
plt.show()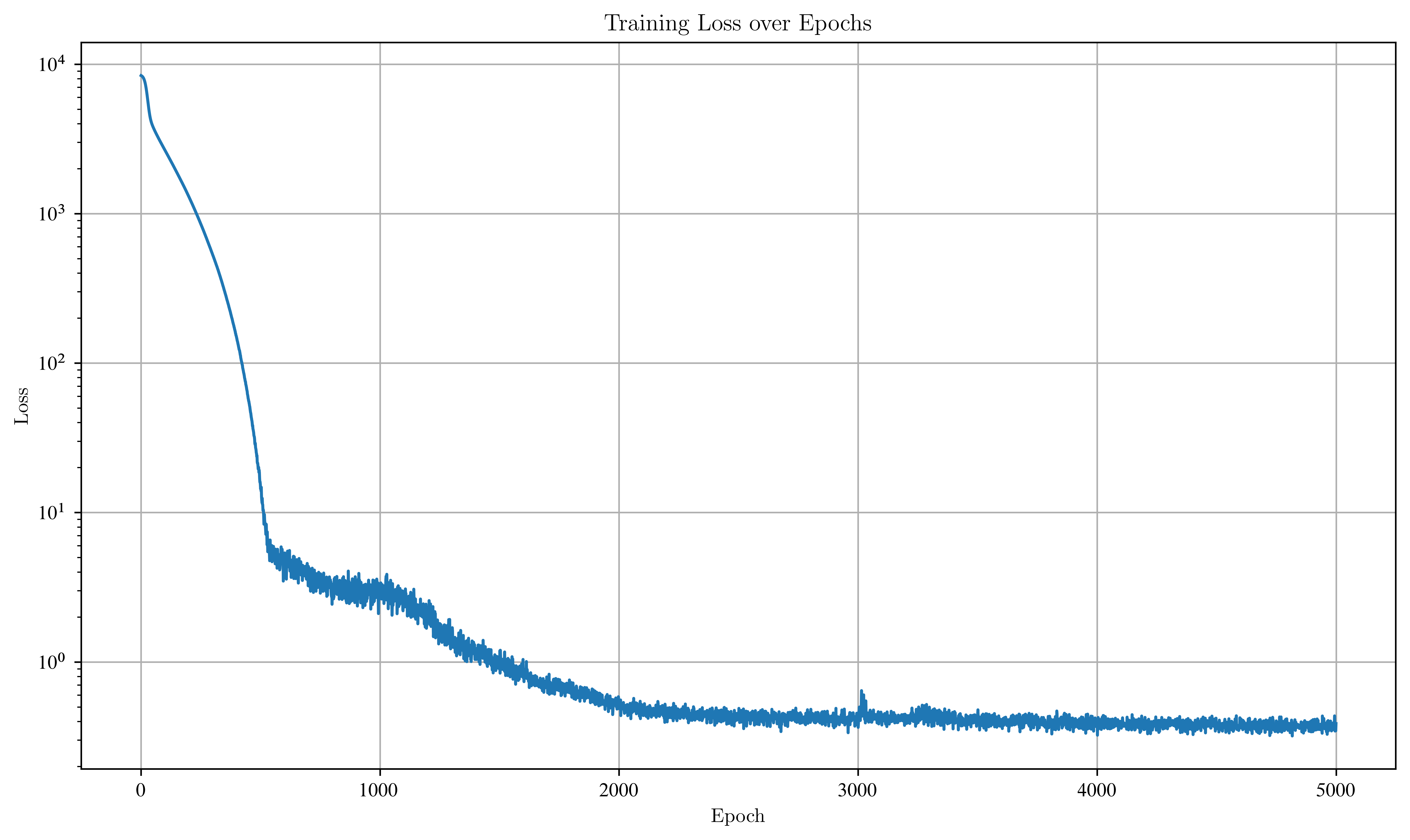
# Plot model solution and compare with numerical solution
from scipy.integrate import solve_ivp
def lotka_volterra(t, y, alpha, beta, gamma, delta):
"""Lotka-Volterra ODE system for solve_ivp."""
x, y_pop = y
dxdt = alpha * x - beta * x * y_pop
dydt = delta * x * y_pop - gamma * y_pop
return [dxdt, dydt]
# Numerical solution using solve_ivp (more modern than odeint)
t_eval = np.linspace(0, tMax, 2000)
y0 = [40.0, 9.0] # Initial conditions matching the PINN
sol = solve_ivp(
lotka_volterra,
[0, tMax],
y0,
args=(alpha, beta, gamma, delta),
t_eval=t_eval,
method="RK45",
)
# PINN prediction
t_tensor = torch.FloatTensor(t_eval[:, None]).to(device)
with torch.no_grad():
xy_pred = model(t_tensor).cpu().numpy()
x_pred = xy_pred[:, 0]
y_pred = xy_pred[:, 1]
# Plot comparison
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
# Time series plot
axes[0].plot(sol.t, sol.y[0], "b-", label="Prey (Numerical)", linewidth=2)
axes[0].plot(sol.t, sol.y[1], "r-", label="Predator (Numerical)", linewidth=2)
axes[0].plot(t_eval, x_pred, "b--", label="Prey (PINN)", linewidth=2, alpha=0.7)
axes[0].plot(t_eval, y_pred, "r--", label="Predator (PINN)", linewidth=2, alpha=0.7)
axes[0].axhline(
y=gamma / delta,
color="b",
linestyle=":",
alpha=0.5,
label=f"x* = {gamma / delta:.1f}",
)
axes[0].axhline(
y=alpha / beta,
color="r",
linestyle=":",
alpha=0.5,
label=f"y* = {alpha / beta:.1f}",
)
axes[0].set_xlabel("Time")
axes[0].set_ylabel("Population")
axes[0].set_title("Lotka-Volterra: PINN vs Numerical Solution")
axes[0].legend(loc="upper right")
axes[0].grid(True)
# Phase space plot
axes[1].plot(sol.y[0], sol.y[1], "g-", label="Numerical", linewidth=2)
axes[1].plot(x_pred, y_pred, "m--", label="PINN", linewidth=2, alpha=0.7)
axes[1].plot(
gamma / delta,
alpha / beta,
"ko",
markersize=10,
label=f"Equilibrium ({gamma / delta:.1f}, {alpha / beta:.1f})",
)
axes[1].plot(y0[0], y0[1], "rs", markersize=10, label=f"Initial ({y0[0]}, {y0[1]})")
axes[1].set_xlabel("Prey (x)")
axes[1].set_ylabel("Predator (y)")
axes[1].set_title("Phase Space")
axes[1].legend()
axes[1].grid(True)
plt.tight_layout()
plt.show()
# Print error metrics
mse_x = np.mean((x_pred - sol.y[0]) ** 2)
mse_y = np.mean((y_pred - sol.y[1]) ** 2)
print(f"\nMSE Prey: {mse_x:.6f}")
print(f"MSE Predator: {mse_y:.6f}")
print(f"Total MSE: {mse_x + mse_y:.6f}")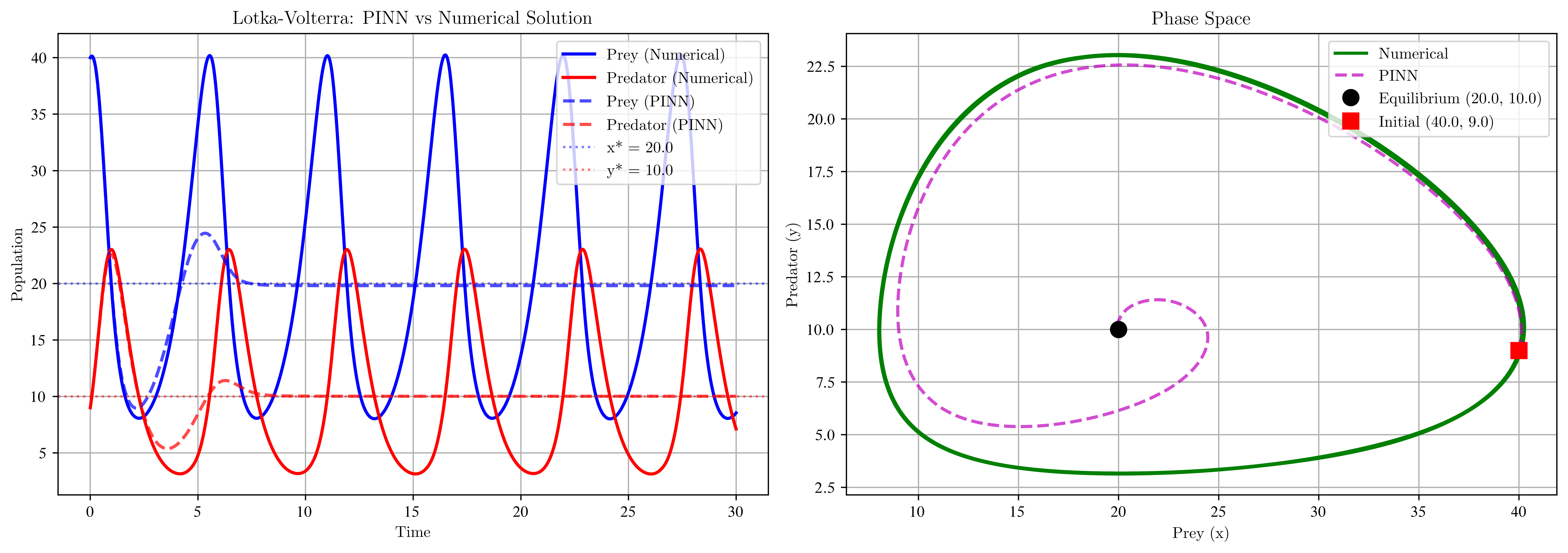
MSE Prey: 102.101862
MSE Predator: 37.963425
Total MSE: 140.065287