Physics-Informed Neural Networks (PINNs) with PyTorch¶
This notebook demonstrates the implementation of Physics-Informed Neural Networks for solving partial differential equations (PDEs).
Overview¶
PINNs are neural networks that incorporate physical laws (PDEs) into the training process. They can:
Solve forward problems (finding solutions given a PDE)
Solve inverse problems (discovering parameters in a PDE)
Work with sparse and noisy data
Example Problem: 1D Heat Equation¶
We’ll solve the heat equation:
With boundary conditions:
(left boundary)
(right boundary)
(initial condition)
1. Import Libraries¶
import matplotlib.pyplot as plt
from matplotlib import cm
from torch import (
Tensor,
abs,
cat,
device,
exp,
float32,
linspace,
manual_seed,
meshgrid,
nn,
no_grad,
ones,
ones_like,
pi,
rand,
sin,
tanh,
tensor,
zeros,
zeros_like,
)
from torch.autograd import grad
from torch.linalg import norm
from torch.optim import Adammanual_seed(42)
cpu_device = device("cpu");
# device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# device = torch.device('mps' if torch.backends.mps.is_available() else device)
# print(f"Using device: {device}")2. Define the Neural Network Architecture¶
class PINN(nn.Module):
"""
Physics-Informed Neural Network for solving PDEs.
The network takes (x, t) as input and outputs u(x, t).
"""
def __init__(self, layers):
super().__init__()
self.layers = nn.ModuleList()
# Build the network
for i in range(len(layers) - 1):
self.layers.append(nn.Linear(layers[i], layers[i + 1]))
# Initialize weights using Xavier initialization
self.init_weights()
def init_weights(self):
for layer in self.layers:
nn.init.xavier_normal_(layer.weight)
nn.init.zeros_(layer.bias)
def forward(self, x, t):
# Concatenate inputs
inputs = cat([x, t], dim=1)
# Pass through hidden layers with tanh activation
for i in range(len(self.layers) - 1):
inputs = tanh(self.layers[i](inputs))
# Output layer (no activation)
output = self.layers[-1](inputs)
return output3. Physics-Informed Loss Functions¶
def compute_pde_residual(model, x, t, alpha=0.01):
"""
Compute the PDE residual for the heat equation.
PDE: du/dt - alpha * d2u/dx2 = 0
"""
x.requires_grad = True
t.requires_grad = True
# Forward pass
u = model(x, t)
# Compute gradients
u_x = grad(u, x, grad_outputs=ones_like(u), create_graph=True, retain_graph=True)[0]
u_t = grad(u, t, grad_outputs=ones_like(u), create_graph=True, retain_graph=True)[0]
# Second derivative with respect to x
u_xx = grad(
u_x, x, grad_outputs=ones_like(u_x), create_graph=True, retain_graph=True
)[0]
# PDE residual
residual = u_t - alpha * u_xx
return residual
def initial_condition(x):
"""
Initial condition: u(x, 0) = sin(pi * x)
"""
return sin(pi * x)
def boundary_condition(t):
"""
Boundary conditions: u(0, t) = u(1, t) = 0
"""
return zeros_like(t)4. Training Function¶
def train_pinn(
model,
device,
n_epochs=5000,
lr=0.001,
n_collocation=10000,
n_initial=100,
n_boundary=100,
alpha=0.01,
):
"""
Train the Physics-Informed Neural Network.
Args:
model: PINN model
n_epochs: Number of training epochs
lr: Learning rate
n_collocation: Number of collocation points for PDE
n_initial: Number of points for initial condition
n_boundary: Number of points for boundary conditions
alpha: Thermal diffusivity coefficient
device: device
"""
optimizer = Adam(model.parameters(), lr=lr)
losses = []
for epoch in range(n_epochs):
optimizer.zero_grad()
# Generate collocation points for PDE (random points in domain)
x_pde = rand(n_collocation, 1, requires_grad=True).to(device)
t_pde = rand(n_collocation, 1, requires_grad=True).to(device)
# Initial condition points (t=0)
x_ic = rand(n_initial, 1).to(device)
t_ic = zeros(n_initial, 1).to(device)
u_ic = initial_condition(x_ic)
# Boundary condition points (x=0 and x=1)
t_bc = rand(n_boundary, 1).to(device)
x_bc_left = zeros(n_boundary, 1).to(device)
x_bc_right = ones(n_boundary, 1).to(device)
u_bc = boundary_condition(t_bc)
# Compute losses
# 1. PDE residual loss
residual = compute_pde_residual(model, x_pde, t_pde, alpha)
loss_pde = (residual**2).mean()
# 2. Initial condition loss
u_pred_ic = model(x_ic, t_ic)
loss_ic = ((u_pred_ic - u_ic) ** 2).mean()
# 3. Boundary condition loss
u_pred_bc_left = model(x_bc_left, t_bc)
u_pred_bc_right = model(x_bc_right, t_bc)
loss_bc = ((u_pred_bc_left - u_bc) ** 2).mean() + (
(u_pred_bc_right - u_bc) ** 2
).mean()
# Total loss
loss = loss_pde + loss_ic + loss_bc
# Backpropagation
loss.backward()
optimizer.step()
losses.append(loss.item())
# Print progress
if (epoch + 1) % 500 == 0:
print(
f"Epoch [{epoch + 1}/{n_epochs}], Loss: {loss.item():.6f}, "
f"PDE: {loss_pde.item():.6f}, IC: {loss_ic.item():.6f}, "
f"BC: {loss_bc.item():.6f}"
)
return losses5. Analytical Solution (for comparison)¶
def analytical_solution(x, t: float, alpha: float = 0.01):
"""
Analytical solution for the 1D heat equation with given initial/boundary conditions.
u(x, t) = sin(pi * x) * exp(-alpha * pi^2 * t)
"""
if not isinstance(x, Tensor):
x = tensor(x, dtype=float32)
if not isinstance(t, Tensor):
t = tensor(t, dtype=float32)
return sin(pi * x) * exp(-alpha * pi**2 * t)6. Train the Model¶
# Define network architecture
layers = [2, 32, 32, 32, 1] # Input: (x, t), Output: u
# Create model
model = PINN(layers).to(cpu_device)
print(model)
# Thermal diffusivity
alpha = 0.01
# Train the model
print("\nTraining PINN...")
losses = train_pinn(model, cpu_device, n_epochs=5000, lr=0.001, alpha=alpha)PINN(
(layers): ModuleList(
(0): Linear(in_features=2, out_features=32, bias=True)
(1-2): 2 x Linear(in_features=32, out_features=32, bias=True)
(3): Linear(in_features=32, out_features=1, bias=True)
)
)
Training PINN...
Epoch [500/5000], Loss: 0.004594, PDE: 0.001176, IC: 0.002395, BC: 0.001023
Epoch [1000/5000], Loss: 0.000618, PDE: 0.000302, IC: 0.000254, BC: 0.000062
Epoch [1500/5000], Loss: 0.000163, PDE: 0.000129, IC: 0.000027, BC: 0.000007
Epoch [2000/5000], Loss: 0.000225, PDE: 0.000075, IC: 0.000034, BC: 0.000116
Epoch [2500/5000], Loss: 0.000060, PDE: 0.000047, IC: 0.000010, BC: 0.000003
Epoch [3000/5000], Loss: 0.000042, PDE: 0.000034, IC: 0.000005, BC: 0.000003
Epoch [3500/5000], Loss: 0.000034, PDE: 0.000026, IC: 0.000004, BC: 0.000005
Epoch [4000/5000], Loss: 0.000026, PDE: 0.000020, IC: 0.000003, BC: 0.000003
Epoch [4500/5000], Loss: 0.000022, PDE: 0.000016, IC: 0.000004, BC: 0.000003
Epoch [5000/5000], Loss: 0.000020, PDE: 0.000013, IC: 0.000002, BC: 0.000004
7. Visualize Training Loss¶
plt.plot(losses)
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.title("Training Loss over Epochs")
plt.yscale("log")
plt.grid(True)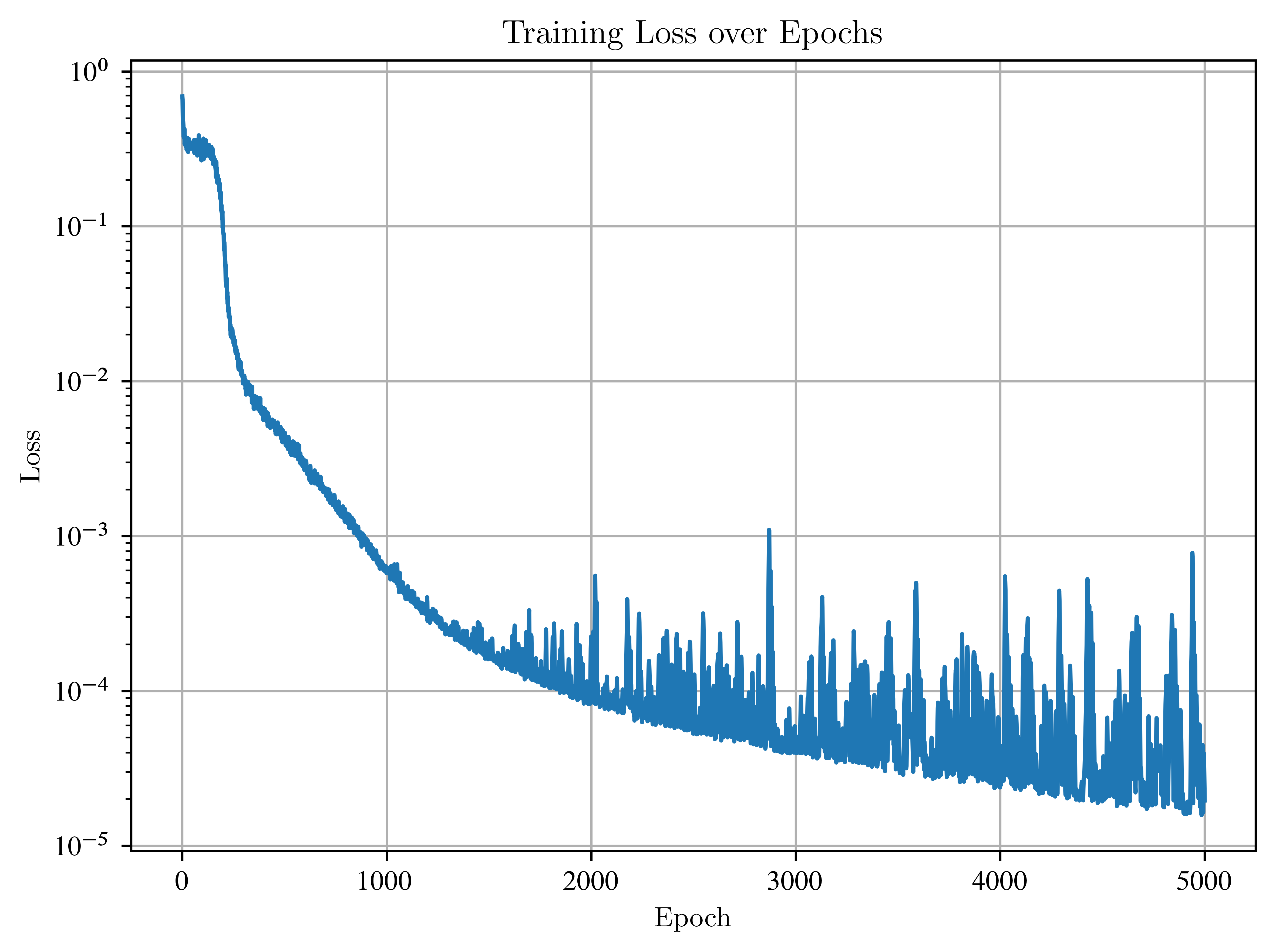
8. Evaluate and Compare with Analytical Solution¶
# Create a grid for evaluation, ensuring tensors are on the correct device from the start.
x_eval = linspace(0, 1, 100, device=cpu_device)
t_eval = linspace(0, 1, 100, device=cpu_device)
# Use 'xy' indexing for meshgrid to ensure correct dimensions for plotting.
X, T = meshgrid(x_eval, t_eval, indexing="xy")
# Predict with PINN on the flattened grid, avoiding intermediate variables.
model.eval()
with no_grad():
u_pred = model(X.flatten().unsqueeze(1), T.flatten().unsqueeze(1))
# Reshape prediction and calculate analytical solution using torch tensors.
U_pred = u_pred.reshape(X.shape)
U_exact = analytical_solution(X, T, alpha)
# Compute and print relative L2 error entirely within PyTorch.
error = abs(U_pred - U_exact)
relative_error = norm(error) / norm(U_exact)
print(f"\nRelative L2 error: {relative_error.item():.6f}")
Relative L2 error: 0.001824
9. Visualize Results¶
# 3D surface plots
fig = plt.figure(figsize=(18, 5))
# PINN prediction
ax1 = fig.add_subplot(131, projection="3d")
surf1 = ax1.plot_surface(X, T, U_pred, cmap=cm.viridis, alpha=0.8)
ax1.set_xlabel(r"$x$")
ax1.set_ylabel(r"$t$")
ax1.set_zlabel(r"$u\left(x,t\right)$")
ax1.set_title("PINN Prediction")
fig.colorbar(surf1, ax=ax1, shrink=0.5)
# Analytical solution
ax2 = fig.add_subplot(132, projection="3d")
surf2 = ax2.plot_surface(X, T, U_exact, cmap=cm.viridis, alpha=0.8)
ax2.set_xlabel(r"$x$")
ax2.set_ylabel(r"$t$")
ax2.set_zlabel(r"$u\left(x,t\right)$")
ax2.set_title("Analytical Solution")
fig.colorbar(surf2, ax=ax2, shrink=0.5)
# Error
ax3 = fig.add_subplot(133, projection="3d")
surf3 = ax3.plot_surface(X, T, error, cmap=cm.hot, alpha=0.8)
ax3.set_xlabel(r"$x$")
ax3.set_ylabel(r"$t$")
ax3.set_zlabel(r"$\left|\text{Error}\right|$")
ax3.set_title("Absolute Error")
fig.colorbar(surf3, ax=ax3, shrink=0.5)
plt.tight_layout()
plt.show()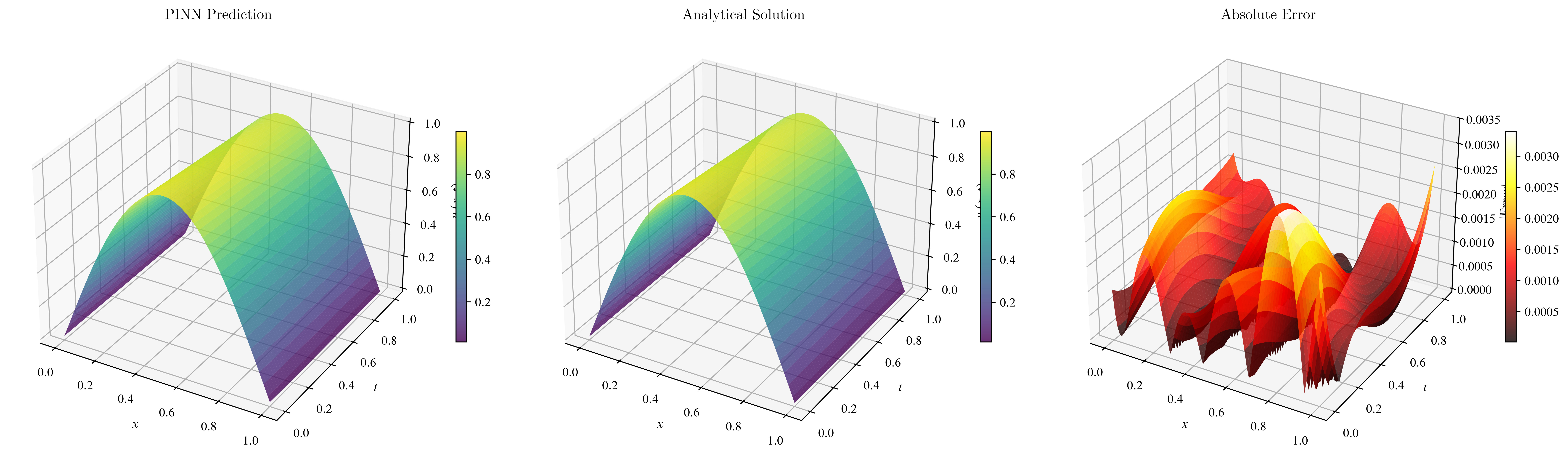
10. Time Snapshots Comparison¶
# Compare solutions at different time points
time_points = [0.0, 0.2, 0.5, 1.0]
# Use torch.linspace for the x-axis points, and place it on the device
x_plot = linspace(0, 1, 100, device=cpu_device)
fig, axes = plt.subplots(2, 2, figsize=(12, 10))
axes = axes.flatten()
model.eval() # Set model to evaluation mode for consistency
for idx, t_point in enumerate(time_points):
# Prepare data as torch tensors on the correct device.
# t_plot is a tensor of the same shape as x_plot, filled with the current time point.
t_plot = ones_like(x_plot) * t_point
# Reshape tensors to be column vectors (N, 1) for the model.
x_tensor = x_plot.unsqueeze(1)
t_tensor = t_plot.unsqueeze(1)
# Get PINN prediction as a torch tensor, no need for .numpy().
with no_grad():
u_pred = model(x_tensor, t_tensor).flatten()
# Calculate analytical solution using torch tensors.
u_exact = analytical_solution(x_plot, t_point, alpha)
# Plotting directly with torch tensors. Matplotlib handles them.
# Moving to .cpu() is good practice if tensors were on GPU.
axes[idx].plot(x_plot.cpu(), u_exact.cpu(), "b-", label="Analytical", linewidth=2)
axes[idx].plot(x_plot.cpu(), u_pred.cpu(), "r--", label="PINN", linewidth=2)
axes[idx].set_xlabel("x")
axes[idx].set_ylabel("u(x,t)")
axes[idx].set_title(f"t = {t_point}")
axes[idx].legend()
axes[idx].grid(True)
plt.tight_layout()
plt.show()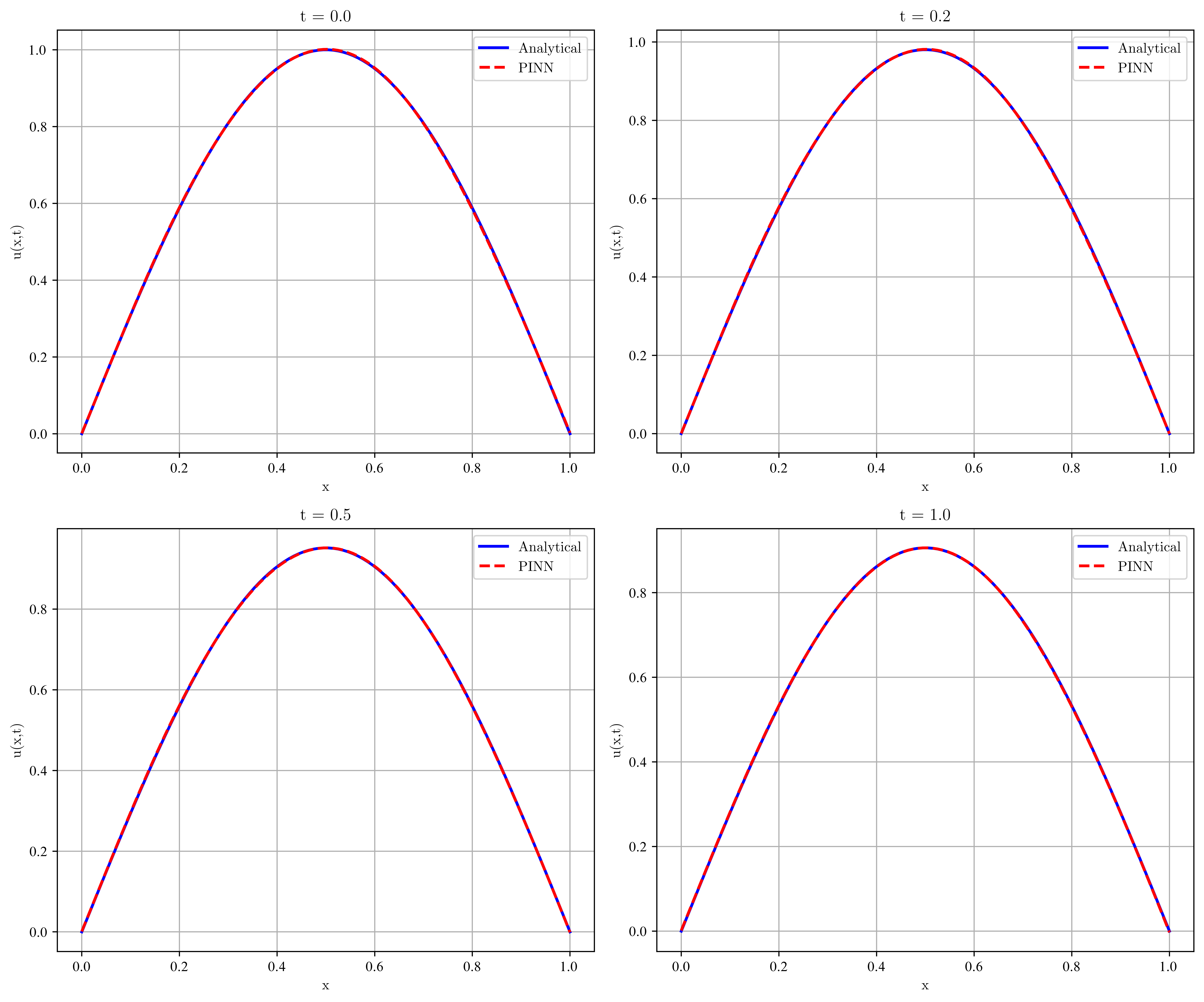
11. Advanced Example: Burgers’ Equation¶
Let’s implement a more complex example with Burgers’ equation:
This is a nonlinear PDE that models fluid dynamics.
# def compute_burgers_residual(model, x, t, nu=0.01):
# """
# Compute the PDE residual for Burgers' equation.
# PDE: du/dt + u * du/dx - nu * d2u/dx2 = 0
# """
# x.requires_grad = True
# t.requires_grad = True
# u = model(x, t)
# u_x = torch.autograd.grad(
# u, x, grad_outputs=torch.ones_like(u), create_graph=True, retain_graph=True
# )[0]
# u_t = torch.autograd.grad(
# u, t, grad_outputs=torch.ones_like(u), create_graph=True, retain_graph=True
# )[0]
# u_xx = torch.autograd.grad(
# u_x, x, grad_outputs=torch.ones_like(u_x), create_graph=True, retain_graph=True
# )[0]
# # Burgers' equation residual (nonlinear term: u * u_x)
# residual = u_t + u * u_x - nu * u_xx
# return residual
# print("Burgers' equation PINN implementation ready!")Summary¶
This notebook demonstrated:
PINN Architecture: A simple feedforward neural network that takes spatial and temporal coordinates as input
Physics-Informed Loss: Combining PDE residuals with initial and boundary conditions
Automatic Differentiation: Using PyTorch’s autograd to compute derivatives needed for the PDE
Training: Optimizing the network to satisfy both data and physics constraints
Validation: Comparing PINN predictions with analytical solutions
Key Advantages of PINNs:¶
Can handle irregular geometries
Work with sparse and noisy data
Can solve inverse problems (parameter estimation)
Mesh-free approach
Can incorporate multiple physical constraints
Further Extensions:¶
Multi-dimensional PDEs
Time-dependent boundary conditions
Inverse problems (parameter discovery)
Complex geometries
Systems of PDEs